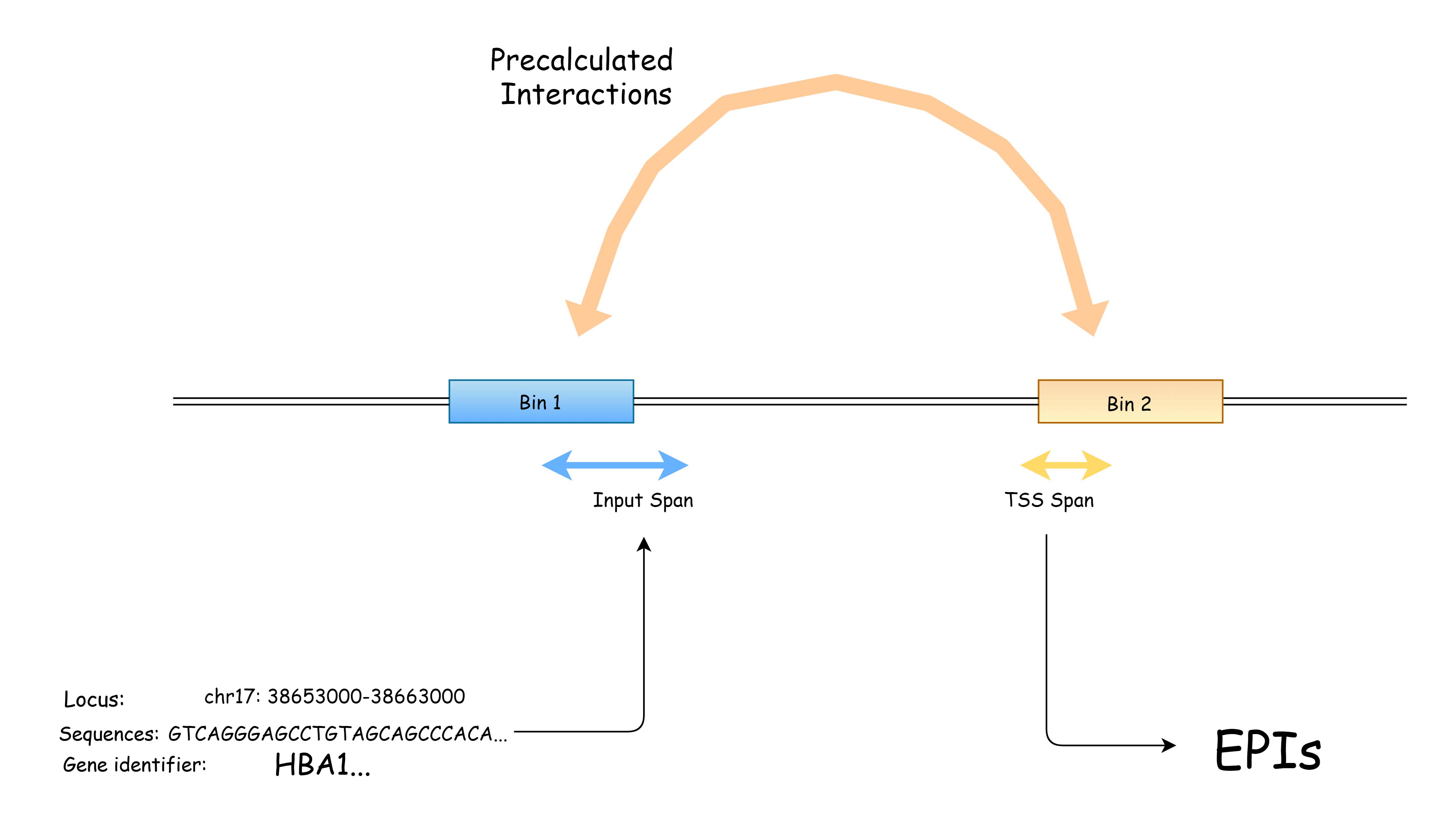
The server is primarily designed for searching distal regulatory interaction. Using the quicksearch box above, the user can search for
a locus, a DNA sequence or an accession number from mainstream databases. The input will be mapped to reference genome and any potential interacting TSS regions will be listed in result page.
Currently, the server has included results of more than 30 cell lines from 3 species. You could narrow down your search scope in Custom Search Page,
then you can browse the research samples that meet your requirment in result page, and reanalyse the result with a new chosen sample.
You can access more info about our searching algorithms in help page.
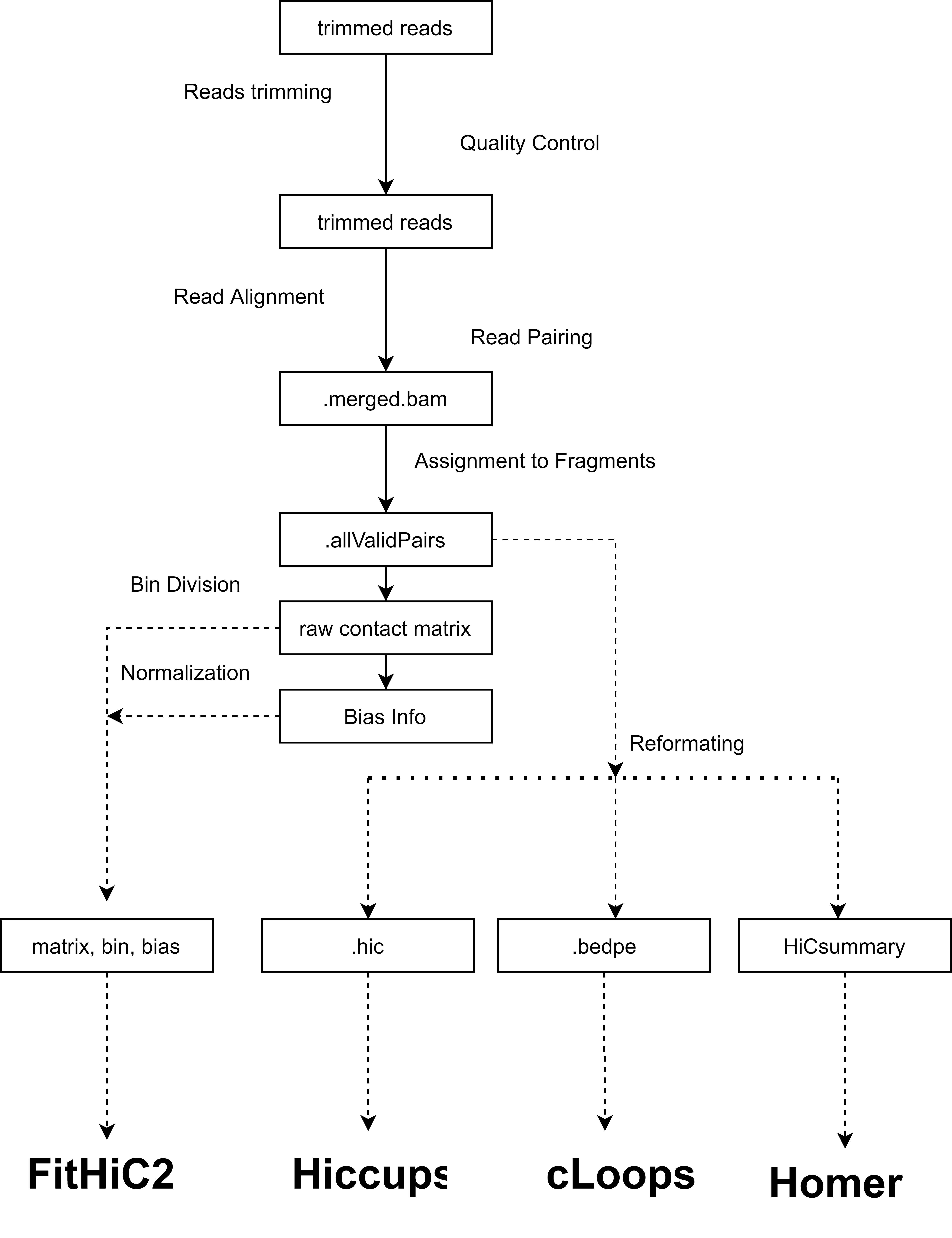
All data are pre-analysed using the same standard pipeline from raw fastq file to obtain unbiased and comparable results. Briefly, read trimming and quality control for raw fastq reads through trim_galore. Then HiC-Pro is used for Hi-C data processing workflow. Valid contact pairs from different samples are pooled in accordance with the source paper to generate .allValidPairs file and contact matrix. Finally, four stand-alone methods are used to call distal interactions separately at 5k,10k and 25k resolution. Detailed explanation about analysing process and parameter setting could be found at Help Page.