Difference between revisions of "Os05g0128400"
(→Function) |
(→Function) |
||
| Line 58: | Line 58: | ||
GFP:mtp1-1 line 2. (ii) Bar chart of average total chlorophyll content (μg) per seedling; significance calculated using paired t-test. See | GFP:mtp1-1 line 2. (ii) Bar chart of average total chlorophyll content (μg) per seedling; significance calculated using paired t-test. See | ||
part (i) for explanation of keys.]] | part (i) for explanation of keys.]] | ||
| + | |||
| + | [[File:HF4.jpg|frame|Fig. 4. OsMTP1 localizes to the vacuole when expressed in | ||
| + | Arabidopsis and yeast. Upper: (A) OsMTP1 vacuolar membrane | ||
| + | localization in the A. thaliana mtp1-1 mutant expressing OsMTP1– | ||
| + | GFP. The red fluorescence is caused by cell walls stained with | ||
| + | propidium iodide (A–D). GFP vacuolar fluorescence (green) in cells | ||
| + | of the root tip region, containing small immature vacuoles (A) and | ||
| + | in elongated root cells with a large central vacuole (B–D). Scale | ||
| + | bar=10 μm. GFP vacuolar fluorescence in cotyledon cells containing | ||
| + | chloroplasts (arrow) (E). Scale bar=25 μm. Lower: OsMTP1 localizes | ||
| + | to the vacuoles of zrc1 cot1 yeast cells expressing OsMTP1–EGFP, | ||
| + | as shown via confocal microscopy. The OsMTP1–EGFP (green)]] | ||
===Expression=== | ===Expression=== | ||
Revision as of 07:27, 3 June 2014
Gene Os05g0128400,namely OsMTP1 and OZT1, is a rice metal tolerance protein and a vacuolar zinc transporter.[1][2]
Contents
Annotated Information
Function
- Rice (Oryza sativa L. ‘Nipponbare’) cDNA subtractive suppression hybridization (SSH) libraries constructed using cadmium (Cd)-treated seedling roots were screened to isolate Cd-responsive genes. A cDNA clone, encoding the rice homolog of Metal Tolerance Protein (OsMTP1), was induced by Cd treatment. Plant MTPs belong to cation diffusion facilitator (CDF) protein family, which are widespread in bacteria, fungi, plants, and animals.OsMTP1 fused to green fluorescent protein was localized in onion epidermal cell plasma membranes, consistent with an OsMTP1 function in heavy metal transporting. OsMTP1 dsRNAi mediated by transgenic assay in rice seedlings resulted in heavy metal sensitivity and changed the heavy metal accumulation in different organs of mature rice under low-concentration heavy metal stress. [1]
- Heavy metal homeostasis is maintained in plant cells by specialized transporters which compartmentalize or efflux metal ions, maintaining cytosolic concentrations within a narrow range. OsMTP1 is a member of the cation diffusion facilitator (CDF)/metal tolerance protein (MTP) family of metal cation transporters in Oryza sativa, which is closely related to Arabidopsis thaliana MTP1. Functional complementation of the Arabidopsis T-DNA insertion mutant mtp1-1 demonstrates that OsMTP1 transports Zn in planta and localizes at the tonoplast. When heterologously expressed in the yeast mutant zrc1 cot1, OsMTP1 complemented its Zn hypersensitivity and was also localized to the vacuole.OsMTP1 alleviated, to some extent, the Co sensitivity of this mutant, rescued the Fe hypersensitivity of the ccc1 mutant at low Fe concentrations, and restored growth of the Cd-hypersensitive mutant ycf1 at low Cd concentrations.These results suggest that OsMTP1 transports Zn but also Co, Fe, and Cd, possibly with lower affinity. Site-directed mutagenesis studies revealed two substitutions in OsMTP1 that alter the transport function of this protein. OsMTP1 harbouring a substitution of Leu82 to a phenylalanine can still transport low levels of Zn, with an enhanced affinity for Fe and Co, and a gain of function for Mn. A substitution of His90 with an aspartic acid completely abolishes Zn transport but improves Fe transport in OsMTP1. These amino acid residues are important in determining substrate specificity and may be a starting point for refining transporter activity in possible biotechnological applications, such as biofortification and phytoremediation.[3]
- In plants, CDFs are known as metal tolerance proteins(MTPs)[4]. MTP genes have been cloned from a number of plant species and are shown to have a role in heavy metal transport [5][6][7][8][9][10][11]. Twelve MTP genes have been classified in Arabidopsis and 10 in rice [12]. The first, identified as ZAT or Zinc Transporter of Arabidopsis [13], was renamed AtMTP1. Overexpression in Arabidopsis enhances Zn resistance[13] while T-DNA insertion [7] or RNA interference (RNAi)-mediated silencing[14] increases Zn sensitivity.The Zn hyperaccumulator plant Arabidopsis halleri had a pentaplication of the MTP1 gene during the evolutionary process,which is believed to have a role in Zn hypertolerance [15]. AtMTP3 is also thought to be involved in Zn transport [16], while AtMTP11 transports Mn (8; 9).The rice orthologue OsMTP1 was recently characterized by Yuan et al. [1] and Lan et al. [2]Located on chromosome 5, it is most highly expressed in mature leaves and stem [1]. Both overexpression and RNAimediated silencing suggest a role for the transporter in Zn,Cd, and nickel (Ni) movement, a hypothesis strengthened by functional complementation of yeast mutants [1]. However, there is controversy over OsMTP1 localization,reported at the plasma membrane when expressed in onion epidermal cells[1] or at the vacuole when expressed in Saccharomyces cerevisiae[1].This study aims to clarify the membrane localization of OsMTP1 and investigate the importance of key residues in transport ability and specificity. It is shown here that OsMTP1 is localized to the vacuole when stably expressed in planta. It is further demonstrated that OsMTP1 is a Zn transporter that can also transport cobalt (Co), Fe, and Cd. Moreover, it is shown that single residue substitutions can alter substrate specificity.
- The CDF family is a ubiquitous family that has been identified in prokaryotes, eukaryotes, and archaea. Members of this family are important heavy metal transporters that transport metal ions out of the cytoplasm. In this research, a full length cDNA named Oryza sativa Zn Transporter 1 (OZT1) that closely related to rat ZnT-2 (Zn Transporter 2) gene was isolated from rice. The OZT1 encoding a CDF family protein shares 28.2 % ~ 84.3 % of identities and 49.3 % ~ 90.9 % of similarities with other zinc transporters such as RnZnT-2, HsZnT-8, RnZnT-8 and AtMTP1.[2]
- OsMTP1 complements the Zn sensitivity of the A. thaliana mtp1-1 mutant.The T-DNA insertion mutant of A. thaliana (Wassilewskija)mtp1-1 is sensitive to elevated Zn, suffering stunted growth and chlorosis compared with the wild type (Fig. 3A).These differences are significant at upward of 100 μM Zn,as indicated by fresh weight and chlorophyll determinations(Fig. 3B). Resistance to Zn is restored when mtp1-1 is transformed with OsMTP1–GFP under the control of the constitutive 35S promoter. Lines 1 and 2 are homozygous T3 lines expressing a C-terminally GFP-tagged version(35S:OsMTP1-GFP:mtp1-1). Similar results were obtained when OsMTP1 without a GFP tag was expressed in the mtp1-1 mutant (results not shown), indicating that GFP tagging does not interfere with the rescue. This also indicates that the localization of OsMTP1 in planta can be investigated using the GFP-tagged construct.[3]
- OsMTP1 localizes to the vacuolar membrane when stably expressed in A. thaliana.The OsMTP1–GFP-expressing lines that showed rescue of the Arabidopsis mtp1-1 mutant phenotype were used to determine the membrane localization of OsMTP1. Confocal images show that OsMTP1 is localized in the vacuolar membrane of plant cells (Fig. 4, upper). Different cells were analysed,including cells in the root tip region containing small immature vacuoles (Fig. 4, upper A), elongated root cells with a large central vacuole (Fig. 4, upper B–D), and cotyledon cells containing chloroplasts (Fig. 4, upper E). All show the same vacuolar localization pattern.[3]
- OsMTP1 localizes to the vacuole of S. cerevisiae.In order to study the subcellular localization of OsMTP1, it was cloned into the pAG426-GFP vector, which fuses a GFP tag to the C-terminus of the gene to enable fluorescence localization studies. zrc1 cot1 yeast cells expressing the OsMTP1–GFP construct were analysed via confocal microscopy which showed that OsMTP1–GFP co-localizes with the vacuoles, which appear as a depression in the differential interference contrast (DIC) image (Fig. 4, lower). The empty vector gives a GFP signal throughout the cytoplasm (Fig. 4, lower B).[3]
- Functional analysis of OsMTP1 in yeast: metal specificity and site-directed mutations.Wild-type and mutant OsMTP1 cDNAs were cloned into the pAG426-GFP vector. OsMTP1 constructs with and without a stop codon were used so that the activity of OsMTP1 with and without a C-terminal GFP tag could be compared when expressed in yeast using the same expression vector. Only the OsMTP1 without a stop codon (indicated by an asterisk in the figures) should produce the OsMTP1–GFP fusion protein.Having such a tag facilitates localization studies and therefore allowed the evaluation of whether particular mutations had an effect on subcellular localization. Although reducing the transport function slightly, GFP tagging did not change the specificity observed in the metal tolerance tests when compared with the non-tagged OsMTP1 protein. These results suggest that OsMTP1 is functionally active with the GFP tag at the C-terminal region. Mutated versions of OsMTP1 with a C-terminal GFP tag also localized to the vacuole, indicating that any differences observed in transport function are not due to altered localization (Supplementary Fig. S1 at JXB online).The transmembrane data predicted by Phobius and TMHMM were used to construct the hypothetical membrane topology of OsMTP1 in Fig. 5A. Seven mutations were tested for their influence on the transport function of OsMTP1(Fig. 5B). The residues were selected based on conservation between CDF family members and also because mutation of some may confer a gain of function. Figure 5A also highlights the substitution sites generated during site-directed mutagenesis,to demonstrate their positions within the protein.[3]
- Zn and Co assays in the zrc1 cot1 yeast mutant.The zrc1 cot1 yeast mutant shows both Zn and Co susceptibility because of lack of the ZRC1 and COT1 genes (Figs6, 7). Heterologous expression of OsMTP1 fully rescues the mutant phenotype at 10 mM Zn (Fig. 6). OsMTP1–GFP also shows rescue of the Zn-sensitive phenotype, with slightly decreased growth at 5 mM and 10 mM Zn compared with the non-tagged OsMTP1, but was as competent at lower concentrations.Compared with OsMTP1–GFP, all of the mutations reduced the rescue of the Zn-sensitive phenotype to some extent, except R149G. This is obvious in L82S and L82F, but most apparent in H90D, which fails to rescue the Zn sensitivity of zrc1 cot1 on 0.25 mM Zn (Fig. 6).In contrast to the results from Yuan et al.[1], it is shown here that OsMTP1 does have Co transport function.OsMTP1 shows partial rescue of Co sensitivity at all concentrations tested (Fig.7), but wild-type growth is far from being restored. The GFP tag again impacts the rescuing ability of OsMTP1 (Fig. 7). The rescue of the Co-sensitive phenotype is less obvious than that of Zn. L82S again reduces the rescuing ability of OsMTP1, whereas L82F, E145G, R149G, and L317A improve Co transport relative to the non-mutated OsMTP1–GFP construct.[3]
- Fe assays in the ccc1 yeast mutant.The Fe transport-deficient ccc1 mutant lacks a vacuolar Fe sequestration function, demonstrating an Fe-sensitive phenotype at high Fe. Growth of ccc1 transformed with an empty pAG426-GFP vector is inhibited on 5 mM Fe compared with the wild type (Fig. 8). OsMTP1 shows rescue of ccc1 on 3.5 mM (data not shown) and 5 mM Fe (Fig. 8), indicating that OsMTP1 can transport iron in yeast. GFP tagging slightly reduces the rescuing ability of OsMTP1 when compared with the non-tagged version of OsMTP1 (Fig. 8).L82F, H90D, and R149G mutations improve Fe transport in ccc1. L82S slightly enhances the sensitive phenotype of ccc1,with poorer growth observed compared with OsMTP1–GFP.[3]
- Cd assay in the ycf1 yeast mutant.The Cd-hypersensitive ycf1 mutant transformed with OsMTP1 clearly confers tolerance to low levels of Cd (Fig. 9),indicating that OsMTP1 can transport Cd in yeast. For all metals tested, expressing OsMTP1–GFP consistently reduces transport when compared with the non-tagged construct, and on Cd this reduction is very obvious (Fig. 9). As the mutations were performed in the OsMTP1–GFP construct, their effects on Cd transport are not conclusive, but it seems that none of the mutations have an effect (Fig. 9). To confirm this,L82F and H90D mutations were tested without GFP tagging,but no difference was noticed in ycf1 complementation compared with the non-mutated OsMTP1 (data not shown).[3]
- Mn assay in the pmr1 yeast mutant.OsMTP1 does not confer Mn tolerance to the Mn-hypersensitive pmr1 mutant (Fig. 10). Of all mutations tested, it was observed that L82F results in a gain of function for Mn complementation on low Mn concentrations(0.25 mM) (Fig. 10).[3]
- Summary of the impact of mutations on substrate.specificity of OsMTP1 Table 2 summarizes the impact of the mutations on the functional complementation of metal-sensitive yeast mutants by OsMTP1. This includes the positioning of residues within the protein predicted by Phobius and TMHMM, also highlighted in Fig. 2.All mutations negatively impact Zn transport ability when compared with the non-mutated OsMTP1–GFP construct,except for R149G; this mutation maintains Zn transport levels and enhances Co and Fe transport, but has no effect on Cd and Mn complementation. L82S shows considerable reduction in Zn, Fe, and Co transport ability. A similar reduction in growth was seen for G127S on Zn-containing media, although no further effect was seen on Co, Fe, Cd,and Mn. All other substitutions have no effect on Cd, but enhance transport of Co or Fe: E145G and L317A both show increased survival on Co. The most notable mutations are L82F and H90D. L82F caused considerable reduction in Zn transport,failing to rescue zrc1cot1 above 1 mM Zn(Fig. 6) but resulting in a considerable increase in both Co and Fe transport and a gain of function for Mn. H90D appears to have abolished Zn transport, with certainly no rescue of zrc1cot1 on or above 0.25 mM Zn; the substitution shows enhanced survival on Fe.[3]




Expression
- OsMTP1 heterologous expression in yeast mutants showed that OsMTP1 was able to complement the mutant strains’ hypersensitivity to Ni, Cd, and Zn, but not other metals including Co and Mn. OsMTP1 expression increased tolerance to Zn, Cd, and Ni in wild-type yeast BY4741 during the exponential growth phase. [1]
- OZT1 was constitutively expressed in various rice tissues. The OZT1 expression was significantly induced both in the seedlings of japonica rice Nipponbare and indica rice IR26 in response to Zn2+ and Cd2+ treatments. Besides, OZT1 expression was also increased when exposed to other excess metals, such as Cu2+, Fe2+ and Mg2+. Subcellular localization analysis indicated that OZT1 localized to vacuole. Heterologous expression of OZT1 in yeast increased tolerance to Zn2+ and Cd2+ stress but not the Mg2+ stress.[2]
Evolution
- OsMTP1 is a bivalent cation transporter localized in the cell membrane, which is necessary for efficient translocation of Zn, Cd and other heavy metals, and maintain ion homeostasis in plant.[1]
- OZT1 is a CDF family vacuolar zinc transporter conferring tolerance to Zn2+ and Cd2+ stress, which is important to transporting and homeostasis of Zn, Cd or other heavy metals in plants.[2]
Labs working on this gene
- 1. Key Laboratory of Plant Resources Conservation and Sustainable Utilization, South China Botanical Garden, Chinese Academy of Sciences
- 2. Graduate School of the Chinese Academy of Sciences
- 3. State Key Laboratory of Crop Genetics and Germplasm Enhancement, Nanjing Agricultural University
- 4.Centre for Biological Science, University of Southampton, Life Sciences Building 85, Highfield Campus, Southampton SO17 1B, UK
- 5.Departamento de Botânica, Instituto de Biociências, Universidade Federal do Rio Grande do Sul, Av. Bento Gonçalves 9500, Porto
Alegre, 91501–970, Brazil
- 6.Centro de Biotecnologia, Universidade Federal do Rio Grande do Sul, Av. Bento Gonçalves 9500, PO Box 15005, Porto Alegre,
91501–970, Brazil
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 1.7 1.8 1.9 Lianyu Yuan,Songguang Yang,Baoxiu Liu,et al.Molecular characterization of a rice metal tolerance protein, OsMTP1.Plant Cell Reports, 2012, 31(1): 67-79 .
- ↑ 2.0 2.1 2.2 2.3 2.4 Hong-Xia Lan, Zhou-Fei Wang, Qi-Hong Wang, et al.Characterization of a vacuolar zinc transporter OZT1 in rice (Oryza sativa L.).Molecular Biology Reports, 2013, 40(2): 1201-1210.
- ↑ 3.0 3.1 3.2 3.3 3.4 3.5 3.6 3.7 3.8 3.9 Paloma K. Menguer, Emily Farthing, Kerry A. Peaston,et al.Functional analysis of the rice vacuolar zinc transporter OsMTP.Journal of Experimental Botany,2013,64(10):2871–2883.
- ↑ Montanini B, Blaudez D, Jeandroz S, Sanders D, Chalot M. 2007. Phylogenetic and functional analysis of the Cation Diffusion Facilitator (CDF) family: improved signature and prediction of substrate specificity. BMC Genomics 8, 107.
- ↑ Blaudez D, Kohler A, Martin F, Sanders D, Chalot M. 2003.Poplar metal tolerance protein 1 confers zinc tolerance and is an oligomeric vacuolar zinc transporter with an essential leucine zipper motif. The Plant Cell 15, 2911–2928.
- ↑ Kim D, Gustin JL, Lahner B, Persans MW, Baek D, Yun DJ, Salt DE. 2004. The plant CDF family member TgMTP1 from the Ni/Zn hyperaccumulator Thlaspi goesingense acts to enhance efflux of Zn at the plasma membrane when expressed in Saccharomyces cerevisiae. The Plant Journal 39, 237–251.
- ↑ 7.0 7.1 Kobae Y, Uemura T, Sato MH, Ohnishi M, Mimura T, Nakagawa T, Maeshima M. 2004. Zinc transporter of Arabidopsis thaliana AtMTP1 is localized to vacuolar membranes and implicated in zinc homeostasis. Plant and Cell Physiology 45, 1749–1758.
- ↑ Delhaize E, Gruber BD, Pittman JK, White RG, Leung H, Miao Y, Jiang L, Ryan PR, Richardson AE. 2007. A role for the AtMTP11 gene of Arabidopsis in manganese transport and tolerance. The Plant Journal 51, 198–210.
- ↑ Peiter E, Montanini B, Gobert A, Pedas P, Husted S, Maathuis FJ, Blaudez D, Chalot M, Sanders D. 2007. A secretory pathwaylocalized cation diffusion facilitator confers plant manganese tolerance. Proceedings of the National Academy of Sciences, USA 104,8532–8537.
- ↑ Kawachi M, Kobae Y, Kogawa S, Mimura T, Kramer U,Maeshima M. 2012. Amino acid screening based on structural modeling identifies critical residues for function, ion selectivity and structure of Arabidopsis MTP1. FEBS Journal 279, 2339–2356.
- ↑ Podar D, Scherer J, Noordally Z, Herzyk P, Nies D, Sanders D. 2012. Metal selectivity determinants in a family of transition metal transporters. Journal of Biological Chemistry 287, 3185–3196.
- ↑ Gustin JL, Zanis MJ, Salt DE. 2011. Structure and evolution of the plant cation diffusion facilitator family of ion transporters. BMC Evolutionary Biology 11, 76.
- ↑ 13.0 13.1 van der Zaal BJ, Neuteboom LW, Pinas JE, Chardonnens AN, Schat H, Verkleij JA, Hooykaas PJ. 1999. Overexpression of a novel Arabidopsis gene related to putative zinc-transporter genes from animals can lead to enhanced zinc resistance and accumulation. Plant Physiology 119, 1047–1055.
- ↑ Desbrosses-Fonrouge AG, Voigt K, Schroder A, Arrivault S, Thomine S, Kramer U. 2005. Arabidopsis thaliana MTP1 is a Zn transporter in the vacuolar membrane which mediates Zn detoxification and drives leaf Zn accumulation. FEBS Letters 579,4165–4174.
- ↑ Shahzad Z, Gosti F, Frerot H, Lacombe E, Roosens N, Saumitou-Laprade P, Berthomieu P. 2010. The five AhMTP1 zinc transporters undergo different evolutionary fates towards adaptive evolution to zinc tolerance in Arabidopsis halleri. PLoS Genetics 6, e1000911.
- ↑ Arrivault S, Senger T, Kramer U. 2006. The Arabidopsis metal tolerance protein AtMTP3 maintains metal homeostasis by mediatingZn exclusion from the shoot under Fe deficiency and Zn oversupply. The Plant Journal 46, 861–879.
Structured Information
- Sequence analysis of OsMTP1
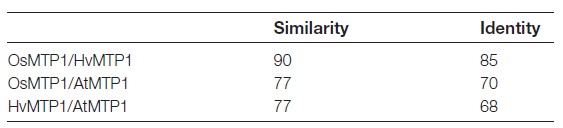
The full-length cDNA fragment of OsMTP1 (accession no. AY266290 /LOC_Os05g03780) was amplified by high-fidelity RT–PCR based on GenBank™ and the Rice Annotation Project information from NCBI (http://www.ncbi.nlm.nih. gov/nuccore). It encodes a protein with 418 amino acids. Percentage identities and similarities between OsMTP1, AtMTP1, and HvMTP1 are shown in Table 1. The highest identity (85%) and similarity (90%) scores at the amino acid level were observed between OsMTP1 and HvMTP1. MTP amino acid sequences were searched for in the genomes of six monocotyledons, Sorghum bicolor, Zea mays, Setaria italica, Hordeum vulgare, O. sativa, and Brachypodium distachyon, and a phylogenetic tree including A. thaliana MTP1 and MTP3 sequences was generated (Fig.1). In the search, it is clear that O. sativa has only one MTP1 sequence, while Z. mays and B. distachyon have two, indicating that the appearance of new MTP1 sequences occurred after the speciation of each monocot. The multiple sequence alignment of AtMTP1, OsMTP1, and HvMTP1 proteins is shown in Fig. 2; positions with a single, fully conserved residue are highlighted, as are those with conserved similar properties. TMDs predicted by Phobius (http://phobius.sbc.su.se/) are also marked in Fig. 2. The proteins are predicted to possess cytoplasmic N- and C-termini and six TMDs, with a small cytoplasmic loop between TMDs II and III, and a large cytoplasmic, histidine-rich loop between TMDs IV and V. OsMTP1 possesses the loop with the most histidine residues, followed by HvMTP1. The 17 residues of the CDF signature sequence are identical for each sequence, and are highlighted in Fig. 2.
| Gene Name |
Os05g0128400 |
|---|---|
| Description |
Similar to Metal tolerance protein 1 (AtMTP1) (Zinc transporter ZAT-1) (ZAT1p) [Contains: Metal tolerance protein 1 short form] |
| Version |
NM_001061074.1 GI:115461878 GeneID:4337694 |
| Length |
3560 bp |
| Definition |
Oryza sativa Japonica Group Os05g0128400, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 5:1653539..1657098 |
| Sequence Coding Region |
1653962..1655218 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008398:1653539..1657098 source=RiceChromosome05 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008398:1653539..1657098 source=RiceChromosome05 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>atggacagccataactcagcacctccccagattgctgaagtgagaatggacatctcatcatctacttctgtagcagctgggaacaaagtttgcagaggtgctgcttgtgacttttctgattccagtaatagctcaaaagatgcaagggagagaatggcgtcaatgaggaagctcattattgctgtgatcctttgcatcatattcatggcggtcgaagtggttggaggtatcaaagcaaacagtttggcaatcttgactgatgcagcccatctcctttcggatgttgcggcctttgccatatctttgttctctctttgggcagctggatgggaagctacaccacagcagtcatatgggtttttccgtatagaaattcttggtgccctggtttctattcagctcatatggctccttgctggtattcttgtctatgaagctattgtaaggctcattaatgaaagtggtgaggtccagggctccctcatgtttgctgtctcagcatttggcttatttgttaacatcataatggctgtcttgcttggtcatgaccatggacatggacacggacatggtcatgggcatggacattcccatgaccatgatcatggtggttctgaccatgaccatcaccaccatgaagatcaagagcatggccatgtacatcaccacgaagatggccatggtaattcaattaccgtcaatctccatcaccatccaggcactggacaccaccaccatgatgctgaggaaccattgctcaagagtgatgctggttgtgacagcacccaatctggtgccaaggatgccaagaaggctcgtcgtaatatcaatgtacacagtgcttatctgcatgtgcttggggattcaatccagagcatcggtgtgatgattggaggggctatcatctggtacaagcccgagtggaagattattgatctcatctgcaccctcatcttctccgtgatcgtactcttcaccacaatcaagatgctgcgcaacatccttgaggtcctgatggagagcacgccccgcgagatcgatgccaccagccttgagaatggcctccgcgacatggacggtgtggttgcagtacatgagctgcacatctgggccataacggtggggaaggttctcctggcgtgccatgtgacaatcactcaggacgcagacgctgatcaaatgctggacaaggtgattgggtacatcaagtctgagtacaacatcagccatgtgaccattcagattgagcgcgagtag</cdnaseq> |
| Protein Sequence |
<aaseq>MDSHNSAPPQIAEVRMDISSSTSVAAGNKVCRGAACDFSDSSNS SKDARERMASMRKLIIAVILCIIFMAVEVVGGIKANSLAILTDAAHLLSDVAAFAISL FSLWAAGWEATPQQSYGFFRIEILGALVSIQLIWLLAGILVYEAIVRLINESGEVQGS LMFAVSAFGLFVNIIMAVLLGHDHGHGHGHGHGHGHSHDHDHGGSDHDHHHHEDQEHG HVHHHEDGHGNSITVNLHHHPGTGHHHHDAEEPLLKSDAGCDSTQSGAKDAKKARRNI NVHSAYLHVLGDSIQSIGVMIGGAIIWYKPEWKIIDLICTLIFSVIVLFTTIKMLRNI LEVLMESTPREIDATSLENGLRDMDGVVAVHELHIWAITVGKVLLACHVTITQDADAD QMLDKVIGYIKSEYNISHVTIQIERE</aaseq> |
| Gene Sequence |
<dnaseqindica>1881..3137#atcctaccccccgttctctcctccccatctccaacgcacgcacgcgccatcaccaccacctcgacgccggcggcgacgaccacggcgacggcaacggcggcggcggcagagaagccctcctcatccccaaggttagcgcctcatcgccgcatccccatcccctttcccaccacctcttcctacgatggattcgtcgtccccaccgagagaccagatcaacctctgctcctctctcgttcgttccacgaatctgctcttcgctcgccgcgggtttccctggggtcattccctgttgctgttattgtttcggattttggagcgctagtttttttttttttgtttaggatggaatattgtttcatggaggaggaggagaaatcaggggagcggcggttacgggtcacgggattgggttgttcgactcgaggttacgggttgctactgcgggattgggttgttcgagtggaggtgtggggtctcgcttcgattcgcttctccgtccatttgatcgggggagagaaaaatttttcaaaatcggtcgaatcacgagtccctgtctccaattgcggtgttacggggtgatgatagggatcgatttttggtcgtttcctccttgggttcttcctaacactacagattcgattggtctggagtaggtttcaactttcgtcagccacaaactgtttgtccgcgagcacggcgtcattgaattggcgaaataaatcgatcaatcaatcaatcagcgttggtccctgaccagtccgtcctcgtgtccaatgaaaagcttccgatggatggatggattggttggaaaatctcattacgatagctctggagtgcttcattagtttcattatattgtccttttggatttgtggacatacactctaatgtaggttatcgtaaacatgaatgaatgccgattagcttgctcccttggtggtgttcccctcttgtgctttcgctgtaaccagcgattgccggttgccttcggttaggtagagagaacttgatctgtttgagggtattttgatttacgaagtacctacttcaatcttgtgacgtttgtttgcaatcatgtgctgaatgtctagttatgtttagggggaccttcttgttgttggattagtgcgacttaattaatcttccctttcgtgatactgtcacttttagttggaaaagactgccaggtgccagctaactgttcttgtactgccagcacgacaattctagtgctctccttctgcattcagttttatggtttttttatttcctgttttgttgtgcttttttaatacatcttgaaagctttagcctgactaagaaaatggtttcattatttgcaatgcttatattatatttgcactagcaagaactatgcacttgctatatttgttgagcaaagagacattggcaccttctaaaaacgaacatatttacttctattcattatatcctttggccaacctttttgtacctgctactgttgtctcctggttaggaaggtggtactgatgcacatttttttaatgtgccgcttaacaaaaggctactcattgttacctggcataagtaccataaaagtacgctctgccatggcacggagattttttacttgtaaacagtgttgcctatgttatatatgtttgatgttttaacttcgttattcattcatgttaagaatgttcatgccaccttttactttatgaagtgctgttcagtttttttcctcccccagtgacacaagaaaggattcttatcactagcttttcgcactatttctaccaggaaccttatgttaacatgttttctcagagcatgagaagtaaatatagcatcatctttgtacttgtgtattgacatgattcttttccccctagatggacagccataactcagcacctccccagattgctgaagtgagaatggacatctcatcatctacttctgtagcagctgggaacaaagtttgcagaggtgctgcttgtgacttttctgattccagtaatagctcaaaagatgcaagggagagaatggcgtcaatgaggaagctcattattgctgtgatcctttgcatcatattcatggcggtcgaagtggttggaggtatcaaagcaaacagtttggcaatcttgactgatgcagcccatctcctttcggatgttgcggcctttgccatatctttgttctctctttgggcagctggatgggaagctacaccacagcagtcatatgggtttttccgtatagaaattcttggtgccctggtttctattcagctcatatggctccttgctggtattcttgtctatgaagctattgtaaggctcattaatgaaagtggtgaggtccagggctccctcatgtttgctgtctcagcatttggcttatttgttaacatcataatggctgtcttgcttggtcatgaccatggacatggacacggacatggtcatgggcatggacattcccatgaccatgatcatggtggttctgaccatgaccatcaccaccatgaagatcaagagcatggccatgtacatcaccacgaagatggccatggtaattcaattaccgtcaatctccatcaccatccaggcactggacaccaccaccatgatgctgaggaaccattgctcaagagtgatgctggttgtgacagcacccaatctggtgccaaggatgccaagaaggctcgtcgtaatatcaatgtacacagtgcttatctgcatgtgcttggggattcaatccagagcatcggtgtgatgattggaggggctatcatctggtacaagcccgagtggaagattattgatctcatctgcaccctcatcttctccgtgatcgtactcttcaccacaatcaagatgctgcgcaacatccttgaggtcctgatggagagcacgccccgcgagatcgatgccaccagccttgagaatggcctccgcgacatggacggtgtggttgcagtacatgagctgcacatctgggccataacggtggggaaggttctcctggcgtgccatgtgacaatcactcaggacgcagacgctgatcaaatgctggacaaggtgattgggtacatcaagtctgagtacaacatcagccatgtgaccattcagattgagcgcgagtaggcttgaccgcttgagacaggtaggtagtgttgatgatagaacggatcatcttcatctcatctcatgggaccaacttatcaggaacctgtttcactggttttatgcagttactgttagctctgcagtgcaaatcagatcagcagagtccaacaattcctgcaggttcatctgaatgttagcttctgatgctgtttattactataagtacgtgttattagtactatcttagtcaattaggtgagttgatgttaatcagtttcgtagtcggttgtattcacatgggtgcagttttcagacaagttttcaggcacctgtgagtacgtcaacctgccttctgcgtctgtagtctagcgcccagccgtactagtttttatgtaaatctgcactggcatgggaaataataaccaagtttcgtttggcctt</dnaseqindica> |
| External Link(s) |