Difference between revisions of "Os06g0208800"
(→Function) |
(→Structured Information) |
||
| (19 intermediate revisions by one other user not shown) | |||
| Line 3: | Line 3: | ||
==Annotated Information== | ==Annotated Information== | ||
===Function=== | ===Function=== | ||
| − | *Rice LYP6 Is LysM-Containing Protein Localized at the Plasma Membrane | + | |
| + | |||
| + | *'''Rice LYP6 Is LysM-Containing Protein Localized at the Plasma Membrane''' | ||
The initial goal of this study was to identify the PGN receptors in | The initial goal of this study was to identify the PGN receptors in | ||
| Line 12: | Line 14: | ||
Phylogenetic analysis of the LysM-containing proteins (LYPs) | Phylogenetic analysis of the LysM-containing proteins (LYPs) | ||
from rice and Arabidopsis indicated that these LYPs could be categorized | from rice and Arabidopsis indicated that these LYPs could be categorized | ||
| − | into two subgroups (Figure 1A). Os-LYP4 (Os09g27890), | + | into two subgroups '''(Figure 1A)'''. Os-LYP4 (Os09g27890), |
Os-LYP5 (Os02g53000), and Os-LYP6 (Os06g10660) belong to | Os-LYP5 (Os02g53000), and Os-LYP6 (Os06g10660) belong to | ||
Subgroup I together with At-LYP2 (At1g77630) and At-LYP3 | Subgroup I together with At-LYP2 (At1g77630) and At-LYP3 | ||
| − | (At1g21880), while CEBiP (Os-LYP1 | + | (At1g21880), while CEBiP (Os-LYP1), Os- |
LYP2 (Os11g34570), Os-LYP3 (Os09g37600), and At-LYP1 | LYP2 (Os11g34570), Os-LYP3 (Os09g37600), and At-LYP1 | ||
(At2g17120) are members of Subgroup II. Since CEBiP from | (At2g17120) are members of Subgroup II. Since CEBiP from | ||
| Line 22: | Line 24: | ||
that the rice PGN receptor may be more likely to exist among | that the rice PGN receptor may be more likely to exist among | ||
LYP4, LYP5, and LYP6 from another branch (subgroup I) of the | LYP4, LYP5, and LYP6 from another branch (subgroup I) of the | ||
| − | phylogenetic tree (Figure 1A). We therefore cloned these three | + | phylogenetic tree '''(Figure 1A)'''. We therefore cloned these three |
rice genes, and their protein products consist of 401, 255, and 409 amino acids, respectively. Bioinformatic analysis predicted | rice genes, and their protein products consist of 401, 255, and 409 amino acids, respectively. Bioinformatic analysis predicted | ||
that LYP4 and LYP6 both have an N-terminal signal | that LYP4 and LYP6 both have an N-terminal signal | ||
peptide, two characteristic LysMs, and a putative C-terminal | peptide, two characteristic LysMs, and a putative C-terminal | ||
glycosylphosphatidylinositol (GPI) anchor signal sequence | glycosylphosphatidylinositol (GPI) anchor signal sequence | ||
| − | (v-site) (Figure 1A; see Supplemental Figure 1 online). This domain | + | (v-site) '''(Figure 1A; see Supplemental Figure 1 online)'''. This domain |
structure suggests that these proteins are localized at the | structure suggests that these proteins are localized at the | ||
plasma membrane through a lipid binding GPI anchor. Although | plasma membrane through a lipid binding GPI anchor. Although | ||
| Line 38: | Line 40: | ||
All characterized homologs of LYP4 and LYP6 have been | All characterized homologs of LYP4 and LYP6 have been | ||
verified as plasma membrane proteins. These LYPs include rice | verified as plasma membrane proteins. These LYPs include rice | ||
| − | CEBiP | + | CEBiP , Arabidopsis LYM1 (At-LYP3), LYM2 |
(At-LYP1), and LYM3 (At-LYP2) <ref name="ref6" /><ref name="ref7" />, and Medicago truncatula LYM1 and LYM2 <ref name="ref8" />. To confirm the plasma membrane localization of Os- | (At-LYP1), and LYM3 (At-LYP2) <ref name="ref6" /><ref name="ref7" />, and Medicago truncatula LYM1 and LYM2 <ref name="ref8" />. To confirm the plasma membrane localization of Os- | ||
LYP4 and Os-LYP6, we inserted green fluorescent protein (GFP) | LYP4 and Os-LYP6, we inserted green fluorescent protein (GFP) | ||
| Line 50: | Line 52: | ||
in rice green tissue protoplasts. In both cases, colocalization | in rice green tissue protoplasts. In both cases, colocalization | ||
of GFP signal with the FM4-64–stained plasma membrane was | of GFP signal with the FM4-64–stained plasma membrane was | ||
| − | readily detected under a confocal microscope (Figure 1B). To | + | readily detected under a confocal microscope '''(Figure 1B)'''. To |
provide additional evidence for the plasma membrane localization | provide additional evidence for the plasma membrane localization | ||
of these proteins, we isolated the microsomal fraction | of these proteins, we isolated the microsomal fraction | ||
| Line 60: | Line 62: | ||
verified in immunoblots showing that CEBiP <ref name="ref4" />was highly enriched in the microsomal fraction while GFP and | verified in immunoblots showing that CEBiP <ref name="ref4" />was highly enriched in the microsomal fraction while GFP and | ||
endogenous tubulin proteins, as nonmembrane proteins, were | endogenous tubulin proteins, as nonmembrane proteins, were | ||
| − | exclusively found in the soluble fraction (Figure 1C). As expected, | + | exclusively found in the soluble fraction '''(Figure 1C)'''. As expected, |
LYP4 and LYP6 were both visualized in the microsomal | LYP4 and LYP6 were both visualized in the microsomal | ||
fraction through immunoblotting with anti-GFP antibodies | fraction through immunoblotting with anti-GFP antibodies | ||
| − | (Figure 1C). Taken together, these data confirmed that LYP4 and | + | '''(Figure 1C)'''. Taken together, these data confirmed that LYP4 and |
| − | LYP6 localize at the plasma membrane of rice cells. | + | LYP6 localize at the plasma membrane of rice cells.<ref name="ref1" /> |
| − | *LYP6 Can Physically Bind PGN and Chitin | + | *'''LYP6 Can Physically Bind PGN and Chitin''' |
As we speculated that LYP4 and LYP6 are rice PGN receptors, | As we speculated that LYP4 and LYP6 are rice PGN receptors, | ||
we next addressed the question whether these proteins could | we next addressed the question whether these proteins could | ||
| Line 75: | Line 77: | ||
6His-tagged recombinant LYP4 and LYP6 proteins in solution. | 6His-tagged recombinant LYP4 and LYP6 proteins in solution. | ||
Indeed, both LYP4 and LYP6 could coprecipitate with these | Indeed, both LYP4 and LYP6 could coprecipitate with these | ||
| − | PGNs (Figure 2A). We further found that the HPLC-purified | + | PGNs '''(Figure 2A)'''. We further found that the HPLC-purified |
soluble PGNXoo muropeptides, which are lysostaphin-digested | soluble PGNXoo muropeptides, which are lysostaphin-digested | ||
products of PGNXoo, could compete with the insoluble PGNXoo | products of PGNXoo, could compete with the insoluble PGNXoo | ||
for binding to these proteins, as the increase of muropeptides | for binding to these proteins, as the increase of muropeptides | ||
in solution was coupled with a decrease of PGNXoo-precipitated | in solution was coupled with a decrease of PGNXoo-precipitated | ||
| − | LYP4 and LYP6 (Figure 2B). Moreover, the analysis of PGN | + | LYP4 and LYP6 '''(Figure 2B)'''. Moreover, the analysis of PGN |
binding kinetics suggested that the association of PGNXoo to | binding kinetics suggested that the association of PGNXoo to | ||
these proteins occurred as early as within 1 min and reached | these proteins occurred as early as within 1 min and reached | ||
| Line 88: | Line 90: | ||
domain. In contrast with LYP4 and LYP6, CERK1 extracellular | domain. In contrast with LYP4 and LYP6, CERK1 extracellular | ||
domain could be barely precipitated by any of these | domain could be barely precipitated by any of these | ||
| − | PGNs (Figure 2A). An effort to test CEBiP for PGN binding was | + | PGNs '''(Figure 2A)'''. An effort to test CEBiP for PGN binding was |
hindered by the difficulty in producing 6His-tagged recombinant | hindered by the difficulty in producing 6His-tagged recombinant | ||
CEBiP proteins in Escherichia coli (data not shown). | CEBiP proteins in Escherichia coli (data not shown). | ||
| Line 98: | Line 100: | ||
were used to pull down the purified LYP4 and LYP6 proteins in | were used to pull down the purified LYP4 and LYP6 proteins in | ||
solution. Surprisingly, these proteins were readily precipitated | solution. Surprisingly, these proteins were readily precipitated | ||
| − | by either commercial chitin products (Figure 2A). Furthermore, | + | by either commercial chitin products '''(Figure 2A)'''. Furthermore, |
when highly purified N-acetylchitohexaose (Isosep), a soluble | when highly purified N-acetylchitohexaose (Isosep), a soluble | ||
hexamer of chitin oligosaccharide, was included in the pull-down | hexamer of chitin oligosaccharide, was included in the pull-down | ||
| Line 104: | Line 106: | ||
negative correlation between the amount of N-acetylchitohexaose | negative correlation between the amount of N-acetylchitohexaose | ||
added and the amounts of LYP4 and LYP6 precipitated | added and the amounts of LYP4 and LYP6 precipitated | ||
| − | by chitin beads (Figure 2B). Moreover, the assay of chitin binding | + | by chitin beads '''(Figure 2B)'''. Moreover, the assay of chitin binding |
kinetics suggested that the binding of chitin to these proteins | kinetics suggested that the binding of chitin to these proteins | ||
occurred within 5 min and became saturated in ;20 min | occurred within 5 min and became saturated in ;20 min | ||
| − | (see Supplemental Figure 4 online). In parallel, we also performed | + | '''(see Supplemental Figure 4 online)'''. In parallel, we also performed |
the same chitin pull-down assay for the purified CERK1 extracellular | the same chitin pull-down assay for the purified CERK1 extracellular | ||
domain. In agreement with the previous suggestion<ref name="ref5" />, we did not detect any coprecipitation of CERK1 with | domain. In agreement with the previous suggestion<ref name="ref5" />, we did not detect any coprecipitation of CERK1 with | ||
| − | chitin beads or the crab shell chitin (Figure 2A). Our data suggested | + | chitin beads or the crab shell chitin '''(Figure 2A)'''. Our data suggested |
that LYP4 and LYP6 physically bind chitin in addition to PGN. | that LYP4 and LYP6 physically bind chitin in addition to PGN. | ||
We next introduced cross-competition into the pull-down | We next introduced cross-competition into the pull-down | ||
experiments and found that addition of excess soluble PGNXoo | experiments and found that addition of excess soluble PGNXoo | ||
muropeptides disrupted the precipitation of these proteins by | muropeptides disrupted the precipitation of these proteins by | ||
| − | chitin beads (Figure 2C). Likewise, the presence of excess soluble | + | chitin beads '''(Figure 2C)'''. Likewise, the presence of excess soluble |
N-acetylchitohexaose blocked the coprecipitation of these | N-acetylchitohexaose blocked the coprecipitation of these | ||
| − | proteins with PGNXoo (Figure 2C). By contrast, addition of excess | + | proteins with PGNXoo '''(Figure 2C)'''. By contrast, addition of excess |
soluble LPS (Sigma-Aldrich) failed to affect the amounts of LYP4 | soluble LPS (Sigma-Aldrich) failed to affect the amounts of LYP4 | ||
| − | and LYP6 precipitated by PGNXoo or chitin beads (Figure 2D). These data reinforced that LYP4 and LYP6 selectively bind | + | and LYP6 precipitated by PGNXoo or chitin beads '''(Figure 2D)'''. These data reinforced that LYP4 and LYP6 selectively bind |
PGN and chitin but not LPS. Furthermore, LYP4 and LYP6 expressed in rice protoplasts | PGN and chitin but not LPS. Furthermore, LYP4 and LYP6 expressed in rice protoplasts | ||
also could be precipitated by different bacterial PGNs and | also could be precipitated by different bacterial PGNs and | ||
| − | commercial chitin products (Figure 2E), confirming the physical | + | commercial chitin products '''(Figure 2E)''', confirming the physical |
association of PGN and chitin to LYPs in rice. By contrast, the | association of PGN and chitin to LYPs in rice. By contrast, the | ||
CEBiP proteins successfully expressed in rice protoplasts could | CEBiP proteins successfully expressed in rice protoplasts could | ||
be precipitated only by chitin but not PGN, while the full-length | be precipitated only by chitin but not PGN, while the full-length | ||
rice CERK1 proteins expressed in rice protoplasts could be pulled | rice CERK1 proteins expressed in rice protoplasts could be pulled | ||
| − | down by neither MAMP (Figure 2E). | + | down by neither MAMP '''(Figure 2E)'''.<ref name="ref1" /> |
| + | |||
| + | *'''Silencing of LYP6 Compromises Diverse PGN- and Chitin-Induced Defense Responses in Rice''' | ||
| − | |||
| − | |||
The findings regarding the plasma membrane localization of LYP4 | The findings regarding the plasma membrane localization of LYP4 | ||
and LYP6 and their physical interactions with both PGN and chitin | and LYP6 and their physical interactions with both PGN and chitin | ||
| Line 142: | Line 144: | ||
LYP6 genes share ;80% identity, the LYP4 and LYP6 transcripts | LYP6 genes share ;80% identity, the LYP4 and LYP6 transcripts | ||
were specifically reduced by ;80% by their cognate | were specifically reduced by ;80% by their cognate | ||
| − | RNAi construct in the silencing lines (see Supplemental Figure 5A | + | RNAi construct in the silencing lines '''(see Supplemental Figure 5A |
| − | online). Notably, the expression of CEBiP and CERK1, the two | + | online)'''. Notably, the expression of CEBiP and CERK1, the two |
known genes involved in rice chitin perception, was affected by | known genes involved in rice chitin perception, was affected by | ||
| − | neither RNAi construct (see Supplemental Figure 5B online). On | + | neither RNAi construct '''(see Supplemental Figure 5B online)'''. On |
the other hand, the expression of LYP4 in its OX lines was increased | the other hand, the expression of LYP4 in its OX lines was increased | ||
by 5.2- to 6.8-fold and that of LYP6 in its OX lines was | by 5.2- to 6.8-fold and that of LYP6 in its OX lines was | ||
| − | increased by 14- to 15-fold (see Supplemental Figure 5C online). | + | increased by 14- to 15-fold '''(see Supplemental Figure 5C online)'''. |
Using LYP RNAi or OX transgenic rice, we investigated three | Using LYP RNAi or OX transgenic rice, we investigated three | ||
different cell responses occurring at different defense time points | different cell responses occurring at different defense time points | ||
| Line 158: | Line 160: | ||
its HPLC-purified muropeptides declined by ;40% in either LYP4 | its HPLC-purified muropeptides declined by ;40% in either LYP4 | ||
or LYP6 RNAi transgenic rice when compared with that produced | or LYP6 RNAi transgenic rice when compared with that produced | ||
| − | in the control rice (Figure 3A). Similarly, the amounts of ROS induced | + | in the control rice '''(Figure 3A)'''. Similarly, the amounts of ROS induced |
by chitin or its soluble fragment N-acetylchitohexaose also | by chitin or its soluble fragment N-acetylchitohexaose also | ||
| − | decreased by 37 to 42% in LYP RNAi transgenic rice (Figure 3A). | + | decreased by 37 to 42% in LYP RNAi transgenic rice '''(Figure 3A)'''. |
By contrast, LYP4 and LYP6 OX transgenic rice showed comparable | By contrast, LYP4 and LYP6 OX transgenic rice showed comparable | ||
| − | ROS production after PGN or chitin treatment (Figure 3A). | + | ROS production after PGN or chitin treatment '''(Figure 3A)'''. |
Notably, all transgenic rice lines and the wild-type rice treated with | Notably, all transgenic rice lines and the wild-type rice treated with | ||
LPS exhibited no significant difference in ROS production | LPS exhibited no significant difference in ROS production | ||
| − | (Figure 3A). Since ROS generation is one of the earliest defense | + | '''(Figure 3A)'''. Since ROS generation is one of the earliest defense |
responses<ref name="ref10" />, these results suggested that | responses<ref name="ref10" />, these results suggested that | ||
LYP4 and LYP6 function quite upstream within the PGN- and | LYP4 and LYP6 function quite upstream within the PGN- and | ||
| Line 175: | Line 177: | ||
PGN/muropeptides or chitin/N-acetylchitohexaose treatment | PGN/muropeptides or chitin/N-acetylchitohexaose treatment | ||
in wild-type rice was substantially suppressed in LYP4 or LYP6 | in wild-type rice was substantially suppressed in LYP4 or LYP6 | ||
| − | RNAi transgenic rice (Figure 3B). However, these defense | + | RNAi transgenic rice '''(Figure 3B)'''. However, these defense |
marker genes responded to 30-min LPS treatment equally well | marker genes responded to 30-min LPS treatment equally well | ||
| − | in both wild-type and LYP RNAi transgenic rice (Figure 3B). | + | in both wild-type and LYP RNAi transgenic rice '''(Figure 3B)'''. |
Furthermore, the callose staining spots on rice young leaves | Furthermore, the callose staining spots on rice young leaves | ||
after PGNXoo muropeptides or N-acetylchitohexaose treatment | after PGNXoo muropeptides or N-acetylchitohexaose treatment | ||
were dramatically reduced in LYP4 or LYP6 RNAi transgenic | were dramatically reduced in LYP4 or LYP6 RNAi transgenic | ||
| − | rice when compared with those in the control rice (Figure 3C). By contrast, PGNXoo muropeptides or N-acetylchitohexaose | + | rice when compared with those in the control rice '''(Figure 3C)'''. By contrast, PGNXoo muropeptides or N-acetylchitohexaose |
treatment resulted in more callose deposition in LYP4 or LYP6 | treatment resulted in more callose deposition in LYP4 or LYP6 | ||
| − | OX transgenic rice than in the control rice (Figure 3C). These | + | OX transgenic rice than in the control rice '''(Figure 3C)'''. These |
data further strengthened the notion that LYP4 and LYP6 play | data further strengthened the notion that LYP4 and LYP6 play | ||
| − | crucial roles in PGN- and chitin-induced defense signaling in rice. | + | crucial roles in PGN- and chitin-induced defense signaling in rice.<ref name="ref1" /> |
| + | |||
| + | * '''LYP6 Affects Rice Susceptibility to Both Bacterial and Fungal Pathogens''' | ||
| − | |||
| − | |||
To validate the importance of LYP4 and LYP6 in rice innate | To validate the importance of LYP4 and LYP6 in rice innate | ||
immunity, we conducted pathogen growth assay in LYP RNAi or | immunity, we conducted pathogen growth assay in LYP RNAi or | ||
| Line 196: | Line 198: | ||
compared with the lesion area caused by X. oryzae infection in | compared with the lesion area caused by X. oryzae infection in | ||
wild-type rice, the lesion areas in LYP4- or LYP6-silencing rice | wild-type rice, the lesion areas in LYP4- or LYP6-silencing rice | ||
| − | were significantly enlarged (Figures 4A and 4B). Accordingly, | + | were significantly enlarged '''(Figures 4A and 4B)'''. Accordingly, |
bacterial growth was increased by 25- to 50-fold in LYP silencing | bacterial growth was increased by 25- to 50-fold in LYP silencing | ||
rice (Figure 4C). Similarly, the lesion area due to X. | rice (Figure 4C). Similarly, the lesion area due to X. | ||
oryzicola infection also expanded significantly in LYP4- or LYP6- | oryzicola infection also expanded significantly in LYP4- or LYP6- | ||
| − | silencing rice relative to that in wild-type rice (Figures 4D and 4E). | + | silencing rice relative to that in wild-type rice '''(Figures 4D and 4E)'''. |
Consistent with this, more X. oryzicola growth was detected in | Consistent with this, more X. oryzicola growth was detected in | ||
| − | LYP silencing rice (Figure 4F). In addition, the LYP silencing rice | + | LYP silencing rice '''(Figure 4F)'''. In addition, the LYP silencing rice |
appeared to be more susceptible to fungal M. oryzae infection as | appeared to be more susceptible to fungal M. oryzae infection as | ||
the lesion size per leaf was considerably larger in LYP silencing | the lesion size per leaf was considerably larger in LYP silencing | ||
| − | rice than in wild-type rice (Figures 4G and 4H). Accordingly, the | + | rice than in wild-type rice '''(Figures 4G and 4H)'''. Accordingly, the |
lesion number per leaf was increased by 0.9- to 1.4-fold in LYP | lesion number per leaf was increased by 0.9- to 1.4-fold in LYP | ||
| − | silencing rice compared with that in wild-type rice (Figure 4I). | + | silencing rice compared with that in wild-type rice '''(Figure 4I)'''. |
These data suggested that knockdown of LYP4 and LYP6 | These data suggested that knockdown of LYP4 and LYP6 | ||
expression in rice results in an increased susceptibility to both | expression in rice results in an increased susceptibility to both | ||
bacterial and fungal pathogens. Conversely, the lesion area caused by X. oryzae infection | bacterial and fungal pathogens. Conversely, the lesion area caused by X. oryzae infection | ||
decreased to 41 to 58% in LYP4 or LYP6 OX rice relative to that | decreased to 41 to 58% in LYP4 or LYP6 OX rice relative to that | ||
| − | in wild-type rice (Figure 4B) and the X. oryzae growth in these | + | in wild-type rice '''(Figure 4B)''' and the X. oryzae growth in these |
| − | LYP OX lines was reduced by more than 80% (Figure 4C). More | + | LYP OX lines was reduced by more than 80% '''(Figure 4C)'''. More |
significantly, the lesion size caused by X. oryzicola infection | significantly, the lesion size caused by X. oryzicola infection | ||
decreased to 15 to 38% in LYP OX rice relative to that in wildtype | decreased to 15 to 38% in LYP OX rice relative to that in wildtype | ||
| − | rice (Figure 4E), and the bacterial growth in these LYP OX | + | rice '''(Figure 4E)''', and the bacterial growth in these LYP OX |
| − | rice dropped by more than 90% (Figure 4F). Moreover, the lesion | + | rice dropped by more than 90% '''(Figure 4F)'''. Moreover, the lesion |
area due to M. oryzae infection also shrank to 19 to 40% in LYP | area due to M. oryzae infection also shrank to 19 to 40% in LYP | ||
| − | OX rice (Figure 4H), and the lesion number per leaf was reduced | + | OX rice '''(Figure 4H)''', and the lesion number per leaf was reduced |
| − | to 50 to 63% in comparison with that in wild-type rice (Figure 4I). | + | to 50 to 63% in comparison with that in wild-type rice '''(Figure 4I)'''. |
These results indicated that upregulation of LYP4 and LYP6 | These results indicated that upregulation of LYP4 and LYP6 | ||
expression in rice leads to an enhanced resistance against both | expression in rice leads to an enhanced resistance against both | ||
| − | bacterial and fungal infection. | + | bacterial and fungal infection.<ref name="ref1" /> |
| + | |||
| + | [[File:69F1.jpg|frame|Figure 1. LysM-Containing LYP4 and LYP6 Are Rice Plasma Membrane Proteins. | ||
| + | (A) Phylogenetic tree and domain structure diagram of LYPs in rice and Arabidopsis. The phylogenetic tree was generated using MEGA4. Full-length | ||
| + | amino acids sequences of plant LYPs were selected for generating a bootstrap neighbor-joining phylogenetic tree. Creye2 from Chlamydomonas | ||
| + | reinhardtii was used as an outgroup. Bootstrap probabilities were obtained from 1000 replicates. A scale bar is indicated. A text file of the alignment | ||
| + | used to generate this tree is available as Supplemental Data Set 1 online. The N-terminal signal peptide, LysM, and the C-terminal GPI anchor signal | ||
| + | (v-site) of each LYP are colored in orange, green, and yellow, respectively. | ||
| + | (B) LYP4 and LYP6 localize at rice plasma membrane. Os-LYP4 and Os-LYP6 with GFP inserted behind the N-terminal signal peptide were individually | ||
| + | expressed in rice protoplasts and visualized by confocal microscopy. FM4-64 dye was used to stain the plasma membrane. | ||
| + | (C) LYP4 and LYP6 localize in the microsomal fraction. LYP4-GFP and LYP6-GFP were individually coexpressed with CEBiP-HA and GFP in rice | ||
| + | protoplasts. Microsomal and soluble fractions of protoplast lysates were separated through Suc gradient centrifugation. Distribution of individual | ||
| + | proteins and the endogenous tubulin was analyzed through immunoblotting with anti-GFP, anti-HA, and antitubulin antibodies. | ||
| + | (D) Relative expression levels of LYP4 and LYP6 in different rice tissues and developmental stages. The expression levels of these genes were | ||
| + | determined by qPCR, and the expression level of each gene in rice calli is set as 100%. | ||
| + | (E) Upregulation of LYP4 and LYP6 transcripts in rice seedlings by X. oryzae. Five-day-old rice seedlings were incubated with X. oryzae suspension (105 | ||
| + | cells/mL) or mock treated (sterile water) for the indicated period. The expression levels of these genes were examined by qPCR, and the induction fold | ||
| + | of each gene was calculated by the gene expression level in X. oryzae–treated seedlings relative to that in mock-treated seedlings at the same time | ||
| + | point. hpi, h postinoculation. | ||
| + | (F) Induction of LYP4 and LYP6 expression in mature rice leaf and root by bacterial pathogen X. oryzae. The mature leaf (the fourth leaf) and the primary | ||
| + | root from the indicated Promoter:GUS transgenic rice at the five-leaf stage were immersed into X. oryzae suspension (105 cells/mL) for 2 h before GUS | ||
| + | staining. Bars = 1 mm. | ||
| + | (G) Induction of LYP4 and LYP6 expression by diverse MAMPs. Five-day-old rice seedlings were treated with 100 mg/mL of insoluble PGNXoo, soluble | ||
| + | PGNXoo muropeptides, insoluble crab shell chitin, soluble chitin fragment N-acetylchitohexaose, soluble LPS, or 100 nM flg22 for 1 h, and the induction | ||
| + | of each gene was examined by qPCR. | ||
| + | The experiments in (B) to (G) were repeated three times with similar results. | ||
| + | The data in (D), (E), and (G) represent the mean 6 SD of nine samples from three independent tests.<ref name="ref1" />]] | ||
| + | |||
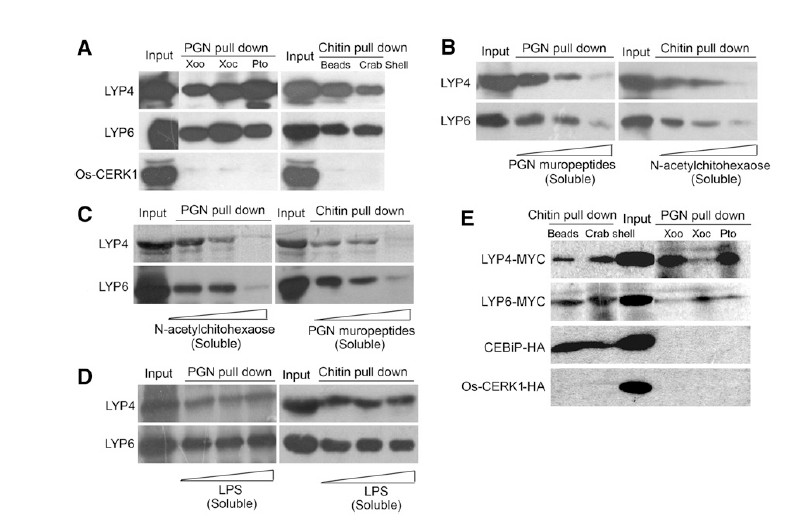
| + | [[File:69F2.jpg|frame|Figure 2. LYP4 and LYP6 Selectively Bind PGN and Chitin but Not LPS. | ||
| + | (A) LYP4 and LYP6 coprecipitate with insoluble PGN or chitin. Sixty micrograms of PGN purified from bacterial pathogens X. oryzae (Xoo), X. oryzicola | ||
| + | (Xoc), or P. syringae pv tomato DC3000 (Pto), commercial chitin beads (NEB), and crab shell chitin (Sigma-Aldrich) were individually used to pull down 1 | ||
| + | mg purified 6His-tagged recombinant Os-LYP4, Os-LYP6, or Os-CERK1 in solution. | ||
| + | (B) Soluble PGN muropeptides and N-acetylchitohexaose compete with insoluble PGN and chitin, respectively, for LYP4 and LYP6 binding. Fifty | ||
| + | micrograms of PGNXoo (left panel) or chitin beads (right panel) were used to pull down 1 mg of purified recombinant Os-LYP4 or Os-LYP6 in the presence | ||
| + | of increasing amounts (0, 40, and 80 mg) of PGNXoo muropeptides (left panel) or N-acetylchitohexaose (right panel). | ||
| + | (C) Soluble PGN muropeptides and N-acetylchitohexaose cross-compete with insoluble chitin and PGN, respectively, for LYP4 and LYP6 binding. Fifty | ||
| + | micrograms of PGNXoo (left panel) or chitin beads (right panel) were used to pull down 1 mg of purified recombinant Os-LYP4 or Os-LYP6 in the presence | ||
| + | of increasing amounts (0, 40, and 80 mg) of N-acetylchitohexaose (left panel) or PGNXoo muropeptides (right panel). | ||
| + | (D) Soluble LPS shows no binding to LYP4 and LYP6. Fifty micrograms of PGNXoo (left panel) or chitin beads (right panel) was used to pull down 1 mg of | ||
| + | purified recombinant Os-LYP4 or Os-LYP6 in the presence of increasing amounts (0, 40, and 80 mg) of LPS. | ||
| + | (E) LYP4 and LYP6 produced in planta coprecipitate with insoluble PGN or chitin. One hundred micrograms of PGN or chitin was individually used to | ||
| + | pull down MYC-tagged LYP4 and LYP6 or HA-tagged CEBiP and CERK1, which was individually expressed in 2 mL of rice protoplasts. Triton X-100 | ||
| + | (0.5%) was used to lyse protoplasts and solubilize membrane proteins. | ||
| + | One of the three biological repeats with similar results is shown, and results were obtained by immunoblotting using anti-His, anti-MYC, or anti-HA | ||
| + | antibodies.<ref name="ref1" />]] | ||
| + | |||
| + | [[File:69F3.jpg|frame|Figure 3. RNAi Silencing or Overexpressing of LYP4 and LYP6 Specifically Modulate PGN- and Chitin- but Not LPS-Induced Defense Responses in Rice. | ||
| + | (A) ROS generation induced by PGN or chitin is significantly suppressed in LYP4 or LYP6 RNAi transgenic rice lines. Roots from 4-d-old wild-type or | ||
| + | transgenic rice seedlings were incubated with 100 mg/mL PGNXoo, PGNXoo muropeptides, crab shell chitin, N-acetylchitohexaose, LPS, or sterile water (mock) | ||
| + | for 30 min before ROS measurement. The data represent the mean 6 SD of nine samples from three independent tests. CK, control (empty vector transgenic | ||
| + | rice); RLU, relative light units; WT, the wild type. | ||
| + | (B) Defense gene activation induced by PGN or chitin is compromised in LYP4 or LYP6 RNAi transgenic rice lines. Callus cells from wild-type or RNAi | ||
| + | transgenic rice lines were incubated with 100 mg/mL PGNXoo, PGNXoo muropeptides, crab shell chitin, N-acetylchitohexaose, LPS, or sterile water | ||
| + | (mock) for 30 min, and the induction of four representative defense marker genes Beta-Glu, MLO, WRKY13, and PAL was determined by qPCR. The | ||
| + | data represent the mean 6 SD of nine samples from three independent tests. | ||
| + | (C) Callose deposition induced by PGN or chitin is substantially impaired in LYP4 and LYP6 RNAi transgenic rice but enhanced in LYP OX rice. The first | ||
| + | leaf of 5-d-old wild-type, LYP4-RNAi-1, LYP6-RNAi-2, LYP4-OX-2, or LYP6-OX-3 rice seedling was immersed into solution containing 10 mg/mL | ||
| + | PGNXoo muropeptides or N-acetylchitohexaose and vacuumed for 30 min. Callose staining was conducted 18 h later. | ||
| + | At least three biological repeats were conducted for individual experiments.<ref name="ref1" />]] | ||
===Expression=== | ===Expression=== | ||
| − | * LYP6 Expression Can Be Induced Quickly by | + | * '''LYP6 Expression Can Be Induced Quickly by Bacterial Pathogen Infection or Diverse MAMPs''' |
| − | Bacterial Pathogen Infection or Diverse MAMPs | ||
To understand better the function of LYP4 and LYP6, we | To understand better the function of LYP4 and LYP6, we | ||
checked the expression profiles of these genes in different | checked the expression profiles of these genes in different | ||
| Line 233: | Line 292: | ||
PCR (qPCR). LYP4 and LYP6 were most abundantly | PCR (qPCR). LYP4 and LYP6 were most abundantly | ||
expressed in rice callus cells, and both transcripts progressively | expressed in rice callus cells, and both transcripts progressively | ||
| − | decreased during maturation (Figure 1D). Furthermore, analysis of | + | decreased during maturation '''(Figure 1D)'''. Furthermore, analysis of |
LYP4 and LYP6 expression patterns in Promoter:GUS (for | LYP4 and LYP6 expression patterns in Promoter:GUS (for | ||
b-glucuronidase) transgenic rice demonstrated strong GUS | b-glucuronidase) transgenic rice demonstrated strong GUS | ||
staining in young seedlings, particularly in the root meristem region | staining in young seedlings, particularly in the root meristem region | ||
| − | and the lateral root primordium (see Supplemental Figure 2 | + | and the lateral root primordium '''(see Supplemental Figure 2 |
| − | online), resembling the expression patterns of their ortholog LYM1 | + | online)''', resembling the expression patterns of their ortholog LYM1 |
in M. truncatula <ref name="ref8" />. Interestingly, expression | in M. truncatula <ref name="ref8" />. Interestingly, expression | ||
of LYP4 and LYP6 in rice seedlings, mature leaves, and roots | of LYP4 and LYP6 in rice seedlings, mature leaves, and roots | ||
| Line 244: | Line 303: | ||
pathogen Xanthomonas oryzae pv oryzae. In 5-d-old rice seedlings, | pathogen Xanthomonas oryzae pv oryzae. In 5-d-old rice seedlings, | ||
1 h X. oryzae treatment induced a 24- and 26-fold increase | 1 h X. oryzae treatment induced a 24- and 26-fold increase | ||
| − | of endogenous LYP4 and LYP6 transcripts, respectively (Figure | + | of endogenous LYP4 and LYP6 transcripts, respectively '''(Figure |
| − | 1E). Moreover, incubation with X. oryzae suspension for 2 h | + | 1E)'''. Moreover, incubation with X. oryzae suspension for 2 h |
rendered a strong GUS activity in mature leaves and roots of | rendered a strong GUS activity in mature leaves and roots of | ||
the Promoter:GUS transgenic rice, while incubation with sterile | the Promoter:GUS transgenic rice, while incubation with sterile | ||
| − | water had no effect on the GUS activity (Figure 1F). The possibility | + | water had no effect on the GUS activity '''(Figure 1F)'''. The possibility |
of false positive GUS staining due to pathogen contamination | of false positive GUS staining due to pathogen contamination | ||
could be excluded as no GUS activity could be detected in the | could be excluded as no GUS activity could be detected in the | ||
| Line 254: | Line 313: | ||
===Evolution=== | ===Evolution=== | ||
| − | The promiscuity of Os-LYP4 and Os-LYP6 in binding carbohydrate | + | *The promiscuity of Os-LYP4 and Os-LYP6 in binding carbohydrate MAMPs PGN and chitin is reminiscent of At-FLS2 binding three different ligands, including flagellin and Ax21 as well as the endogenous CLV3 peptide <ref name="ref11" /> |
| − | MAMPs PGN and chitin is reminiscent of At-FLS2 | ||
| − | binding three different ligands, including flagellin and Ax21 as | ||
| − | well as the endogenous CLV3 peptide <ref name="ref11" /> | ||
<ref name="ref12" /> The promiscuity of PRRs in sensing multiple | <ref name="ref12" /> The promiscuity of PRRs in sensing multiple | ||
MAMPs provides a distinct physiological advantage to the | MAMPs provides a distinct physiological advantage to the | ||
| Line 268: | Line 324: | ||
groups, bacteria and fungi. As the expression of LYP4 and | groups, bacteria and fungi. As the expression of LYP4 and | ||
LYP6 genes could be rapidly upregulated upon recognition of | LYP6 genes could be rapidly upregulated upon recognition of | ||
| − | either MAMP (Figure 1G), it seems that either type of microbial | + | either MAMP'''(Figure 1G)''', it seems that either type of microbial |
infection would quickly sensitize rice for further infection by | infection would quickly sensitize rice for further infection by | ||
both groups of microbes. Interestingly, although the transgenic | both groups of microbes. Interestingly, although the transgenic | ||
rice overexpressing LYP4 or LYP6 indeed demonstrated | rice overexpressing LYP4 or LYP6 indeed demonstrated | ||
| − | an enhanced pathogen resistance (Figure 4), the PGN- or chitininduced | + | an enhanced pathogen resistance '''(Figure 4)''', the PGN- or chitininduced |
ROS production in these rice plants did not show significant | ROS production in these rice plants did not show significant | ||
| − | difference compared that in wild-type rice (Figure 3A). | + | difference compared that in wild-type rice '''(Figure 3A)'''. |
Pathogen resistance is a complicated consequence of innate | Pathogen resistance is a complicated consequence of innate | ||
immunity, whereas ROS production is just one of the very early | immunity, whereas ROS production is just one of the very early | ||
| Line 286: | Line 342: | ||
(LYP2, LYP3, and LYP5) still remain enigmatic. LYP2 and LYP5 | (LYP2, LYP3, and LYP5) still remain enigmatic. LYP2 and LYP5 | ||
are presumably located in the apoplastic space rather than the | are presumably located in the apoplastic space rather than the | ||
| − | plasma membrane due to lack of the GPI anchor (Figure 1A), | + | plasma membrane due to lack of the GPI anchor '''(Figure 1A)''', |
whereas LYP3 likely resides at the plasma membrane like LYP4 | whereas LYP3 likely resides at the plasma membrane like LYP4 | ||
and LYP6. LYP2, LYP3, and LYP5 all contain two intact LysMs | and LYP6. LYP2, LYP3, and LYP5 all contain two intact LysMs | ||
| − | (Figure 1A; see Supplemental Figure 1 online); thus, their binding | + | '''(Figure 1A; see Supplemental Figure 1 online)'''; thus, their binding |
capacity to PGN or chitin cannot be excluded at this moment. | capacity to PGN or chitin cannot be excluded at this moment. | ||
By inference, they may serve certain regulatory functions in PGN | By inference, they may serve certain regulatory functions in PGN | ||
| Line 299: | Line 355: | ||
when expressed in rice cells as the GFP hybrids of both proteins | when expressed in rice cells as the GFP hybrids of both proteins | ||
showed an actual molecular mass around 100 kD instead of the | showed an actual molecular mass around 100 kD instead of the | ||
| − | predicted molecular mass of 65 kD (Figure 1C), reminiscent of | + | predicted molecular mass of 65 kD '''(Figure 1C)''', reminiscent of |
other LYP proteins expressed in planta <ref name="ref4" /> | other LYP proteins expressed in planta <ref name="ref4" /> | ||
<ref name="ref8" /><ref name="ref7" /> Intriguingly, the | <ref name="ref8" /><ref name="ref7" /> Intriguingly, the | ||
| Line 315: | Line 371: | ||
redundant. This was because knockdown of single LYP gene | redundant. This was because knockdown of single LYP gene | ||
expression in rice was sufficient to impair both PGN- and | expression in rice was sufficient to impair both PGN- and | ||
| − | chitin-induced defense responses significantly (Figure 3) and | + | chitin-induced defense responses significantly '''(Figure 3)''' and |
| − | to cause severe bacterial or fungal infection phenotypes (Figure 4). | + | to cause severe bacterial or fungal infection phenotypes '''(Figure 4)'''. |
Similar observations recently have been made for Arabidopsis | Similar observations recently have been made for Arabidopsis | ||
PGN receptors LYM1 and LYM3, where lym1 lym3 double mutant | PGN receptors LYM1 and LYM3, where lym1 lym3 double mutant | ||
| Line 329: | Line 385: | ||
role in rice chitin perception. In line with this speculation, | role in rice chitin perception. In line with this speculation, | ||
RNAi silencing of CEBiP diminished the chitin-induced ROS | RNAi silencing of CEBiP diminished the chitin-induced ROS | ||
| − | generation by 85% | + | generation by 85% , while silencing of LYP4/6 |
| − | only reduced the chitin-induced ROS generation by ;40% (Figure | + | only reduced the chitin-induced ROS generation by ;40% '''(Figure |
| − | 3A). However, as 30% of the upregulated genes and 20% of the | + | 3A)'''. However, as 30% of the upregulated genes and 20% of the |
downregulated genes could respond to chitin equally well in wildtype | downregulated genes could respond to chitin equally well in wildtype | ||
and in CEBiP-RNAi rice cells, CEBiP and LYP4/6 proteins are | and in CEBiP-RNAi rice cells, CEBiP and LYP4/6 proteins are | ||
| Line 359: | Line 415: | ||
and chitin perception machineries between rice and Arabidopsis | and chitin perception machineries between rice and Arabidopsis | ||
will provide valuable evolutionary insights for understanding critical | will provide valuable evolutionary insights for understanding critical | ||
| − | mechanisms underlying innate immunity signaling in plants. | + | mechanisms underlying innate immunity signaling in plants.<ref name="ref1" /> |
| + | |||
| + | [[File:69F4.jpg|frame|Figure 4. RNAi Silencing or Overexpressing of LYP4 and LYP6 Affects Rice Innate Immunity against Bacterial and Fungal Pathogens.<ref name="ref1" />]] | ||
| + | |||
| + | [[File:69F5.jpg|frame|Figure 5. PGN and Chitin Engage Overlapping Receptor Components in | ||
| + | Rice. | ||
| + | (A) Rice cells pretreated with excess PGN have a dramatically attenuated | ||
| + | alkalinization response to subsequent chitin treatment. Rice callus cells | ||
| + | were first treated with 600 mg/mL insoluble PGNXoo for 40 min (endpoint | ||
| + | marked by the arrow) and then treated with 100 mg/mL PGNXoo muropeptides | ||
| + | (green curve), 100 mg/mL N-acetylchitohexaose (blue curve), | ||
| + | or 10 nM flg22 (purple curve). | ||
| + | (B) Rice cells pretreated with excess chitin have a dramatically attenuated | ||
| + | alkalinization response to subsequent PGN treatment. Rice callus | ||
| + | cells were first treated with 600 mg/mL crab shell chitin for 40 min | ||
| + | (endpoint marked by the arrow) and then treated with 100 mg/mL PGNXoo | ||
| + | muropeptides (green curve), 100 mg/mL N-acetylchitohexaose (blue | ||
| + | curve), or 10 nM flg22 (purple curve).(C) PGNXoo muropeptides and N-acetylchitohexaose used in (A) and (B) | ||
| + | are active. Rice callus cells were first treated with 1 mM flg22 for 40 min | ||
| + | (endpoint marked by the arrow) and then treated with 100 mg/mL PGNXoo | ||
| + | muropeptides (green curve), 100 mg/mL N-acetylchitohexaose (blue | ||
| + | curve), or 10 nM flg22 (purple curve). | ||
| + | Three biological replicates were conducted for (A) to (C), and similar | ||
| + | results were obtained.<ref name="ref1" />]] | ||
==Others about this gene== | ==Others about this gene== | ||
| − | PGN and Chitin Engage Overlapping Perception | + | *'''PGN and Chitin Engage Overlapping Perception Components in Rice''' |
| − | Components in Rice | ||
The aforementioned data suggested that LYP4 and LYP6 may | The aforementioned data suggested that LYP4 and LYP6 may | ||
be shared by both PGN and chitin perception systems in rice. If | be shared by both PGN and chitin perception systems in rice. If | ||
| Line 380: | Line 458: | ||
the saturating PGN treatment induced a second spike of medium | the saturating PGN treatment induced a second spike of medium | ||
alkalinization, while the N-acetylchitohexaose treatment | alkalinization, while the N-acetylchitohexaose treatment | ||
| − | led to only a slight increase of medium pH (Figure 5A). The | + | led to only a slight increase of medium pH '''(Figure 5A)'''. The |
alkalinization response to flg22 excluded the possibility that | alkalinization response to flg22 excluded the possibility that | ||
disappearance of the alkalinization response to chitin was due | disappearance of the alkalinization response to chitin was due | ||
| Line 392: | Line 470: | ||
10 nM flg22. Likewise, although the flg22 treatment following the | 10 nM flg22. Likewise, although the flg22 treatment following the | ||
saturating chitin treatment could induce another dramatic elevation of medium pH, the PGNXoo muropeptides gave rise to | saturating chitin treatment could induce another dramatic elevation of medium pH, the PGNXoo muropeptides gave rise to | ||
| − | only a negligible alkalinization response (Figure 5B). These | + | only a negligible alkalinization response '''(Figure 5B)'''. These |
results repeatedly suggested that chitin and PGN are sharing | results repeatedly suggested that chitin and PGN are sharing | ||
overlapping perception systems. In the third set of experiments, | overlapping perception systems. In the third set of experiments, | ||
| Line 399: | Line 477: | ||
N-acetylchitohexaose, and 10 nM flg22. In contrast with flg22, | N-acetylchitohexaose, and 10 nM flg22. In contrast with flg22, | ||
both PGNXoo muropeptides and N-acetylchitohexaose could | both PGNXoo muropeptides and N-acetylchitohexaose could | ||
| − | provoke a further medium alkalinization (Figure 5C), verifying | + | provoke a further medium alkalinization '''(Figure 5C)''', verifying |
that PGNXoo muropeptides and N-acetylchitohexaose used in | that PGNXoo muropeptides and N-acetylchitohexaose used in | ||
| − | these experiments (Figures 5A to 5C) were active elicitors. | + | these experiments '''(Figures 5A to 5C)''' were active elicitors. |
Taken together, these data indirectly support the notion that LYP4 and LYP6 are dual function receptors for PGN and chitin | Taken together, these data indirectly support the notion that LYP4 and LYP6 are dual function receptors for PGN and chitin | ||
| − | in rice. | + | in rice.<ref name="ref1" /> |
==Labs working on this gene== | ==Labs working on this gene== | ||
| − | + | *State Key Laboratory of Biocontrol, Key Laboratory of Gene Engineering of the Ministry of Education and Guangdong Key Laboratory of Plant Resources, School of Life Sciences, Sun Yat-sen University, 510275 Guangzhou, People’s Republic of China | |
==References== | ==References== | ||
| Line 468: | Line 546: | ||
==Structured Information== | ==Structured Information== | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
[[Category:Genes]] | [[Category:Genes]] | ||
[[Category:Japonica mRNA]] | [[Category:Japonica mRNA]] | ||
Latest revision as of 08:09, 12 June 2015
The rice gene Os06g020880,namely LYP6,is a Lysin Motif-Containing Protain and regarded as dual functional PRRs sensing bacterial peptidoglycan (PGN) and fungal chitin. [1]
Contents
Annotated Information
Function
- Rice LYP6 Is LysM-Containing Protein Localized at the Plasma Membrane
The initial goal of this study was to identify the PGN receptors in rice. As LysM was known as the binding motif for PGN in prokaryotes and the PGN-related chitin or Nod factor in plants [2][3] , we postulated that the potential rice PGN receptor highly likely contains LysM. Phylogenetic analysis of the LysM-containing proteins (LYPs) from rice and Arabidopsis indicated that these LYPs could be categorized into two subgroups (Figure 1A). Os-LYP4 (Os09g27890), Os-LYP5 (Os02g53000), and Os-LYP6 (Os06g10660) belong to Subgroup I together with At-LYP2 (At1g77630) and At-LYP3 (At1g21880), while CEBiP (Os-LYP1), Os- LYP2 (Os11g34570), Os-LYP3 (Os09g37600), and At-LYP1 (At2g17120) are members of Subgroup II. Since CEBiP from subgroup II was previously characterized as the chitin receptor in rice[4][5], we reasoned that the rice PGN receptor may be more likely to exist among LYP4, LYP5, and LYP6 from another branch (subgroup I) of the phylogenetic tree (Figure 1A). We therefore cloned these three rice genes, and their protein products consist of 401, 255, and 409 amino acids, respectively. Bioinformatic analysis predicted that LYP4 and LYP6 both have an N-terminal signal peptide, two characteristic LysMs, and a putative C-terminal glycosylphosphatidylinositol (GPI) anchor signal sequence (v-site) (Figure 1A; see Supplemental Figure 1 online). This domain structure suggests that these proteins are localized at the plasma membrane through a lipid binding GPI anchor. Although LYP5 shows a substantial sequence similarity to the N terminus of LYP6, a long C-terminal portion (;150 amino acids) including the GPI anchor signal sequence is absent from this protein (Figure 1A). Therefore, we focused only on LYP4 and LYP6, which have the GPI anchor for membrane attachment, in the subsequent study. All characterized homologs of LYP4 and LYP6 have been verified as plasma membrane proteins. These LYPs include rice CEBiP , Arabidopsis LYM1 (At-LYP3), LYM2 (At-LYP1), and LYM3 (At-LYP2) [6][7], and Medicago truncatula LYM1 and LYM2 [8]. To confirm the plasma membrane localization of Os- LYP4 and Os-LYP6, we inserted green fluorescent protein (GFP) behind the N-terminal signal peptide in both proteins. This fusion strategy was based on the fact that both the N-terminal signal peptide and the C-terminal GPI anchor signal sequence eventually would be removed from a mature GPI-anchored protein. Since plant protoplast systems have been used successfully to detect the cell surface localization of GPI-anchored proteins [9], we transiently expressed LYP4-GFP or LYP6-GFP using a monocot-specific constitutive Act1 promoter in rice green tissue protoplasts. In both cases, colocalization of GFP signal with the FM4-64–stained plasma membrane was readily detected under a confocal microscope (Figure 1B). To provide additional evidence for the plasma membrane localization of these proteins, we isolated the microsomal fraction from rice cells in which LYP4-GFP or LYP6-GFP was coexpressed with GFP and CEBiP-HA (HA tag behind the N-terminal signal peptide of CEBiP). These constructs were all transiently expressed in rice protoplasts under the control of the constitutive Act1 promoter. Successful preparation of rice microsomal fractions was verified in immunoblots showing that CEBiP [4]was highly enriched in the microsomal fraction while GFP and endogenous tubulin proteins, as nonmembrane proteins, were exclusively found in the soluble fraction (Figure 1C). As expected, LYP4 and LYP6 were both visualized in the microsomal fraction through immunoblotting with anti-GFP antibodies (Figure 1C). Taken together, these data confirmed that LYP4 and LYP6 localize at the plasma membrane of rice cells.[1]
- LYP6 Can Physically Bind PGN and Chitin
As we speculated that LYP4 and LYP6 are rice PGN receptors, we next addressed the question whether these proteins could physically bind PGN. Insoluble PGN purified from three different bacterial pathogens, X. oryzae pv oryzae (PGNXoo), X. oryzae pv oryzicola (PGNXoc), and Pseudomonas syringae pv tomato (Pto) DC3000 (PGNPto), was individually used to pull down the purified 6His-tagged recombinant LYP4 and LYP6 proteins in solution. Indeed, both LYP4 and LYP6 could coprecipitate with these PGNs (Figure 2A). We further found that the HPLC-purified soluble PGNXoo muropeptides, which are lysostaphin-digested products of PGNXoo, could compete with the insoluble PGNXoo for binding to these proteins, as the increase of muropeptides in solution was coupled with a decrease of PGNXoo-precipitated LYP4 and LYP6 (Figure 2B). Moreover, the analysis of PGN binding kinetics suggested that the association of PGNXoo to these proteins occurred as early as within 1 min and reached saturation in ;30 min (see Supplemental Figure 4 online). In parallel, we conducted a PGN pull-down assay for the purified 6His-tagged recombinant Os-CERK1 (Os-LysM-RLK9) extracellular domain. In contrast with LYP4 and LYP6, CERK1 extracellular domain could be barely precipitated by any of these PGNs (Figure 2A). An effort to test CEBiP for PGN binding was hindered by the difficulty in producing 6His-tagged recombinant CEBiP proteins in Escherichia coli (data not shown). These data revealed a specific physical interaction between bacterial PGN and the two rice LYPs. As PGN is structurally related to chitin[3], we tested whether LYP4 and LYP6 could also physically bind to chitin. Chitin beads (NEB) and insoluble crab shell chitin (Sigma-Aldrich) were used to pull down the purified LYP4 and LYP6 proteins in solution. Surprisingly, these proteins were readily precipitated by either commercial chitin products (Figure 2A). Furthermore, when highly purified N-acetylchitohexaose (Isosep), a soluble hexamer of chitin oligosaccharide, was included in the pull-down assay to compete with the chitin beads, we observed a clear negative correlation between the amount of N-acetylchitohexaose added and the amounts of LYP4 and LYP6 precipitated by chitin beads (Figure 2B). Moreover, the assay of chitin binding kinetics suggested that the binding of chitin to these proteins occurred within 5 min and became saturated in ;20 min (see Supplemental Figure 4 online). In parallel, we also performed the same chitin pull-down assay for the purified CERK1 extracellular domain. In agreement with the previous suggestion[5], we did not detect any coprecipitation of CERK1 with chitin beads or the crab shell chitin (Figure 2A). Our data suggested that LYP4 and LYP6 physically bind chitin in addition to PGN. We next introduced cross-competition into the pull-down experiments and found that addition of excess soluble PGNXoo muropeptides disrupted the precipitation of these proteins by chitin beads (Figure 2C). Likewise, the presence of excess soluble N-acetylchitohexaose blocked the coprecipitation of these proteins with PGNXoo (Figure 2C). By contrast, addition of excess soluble LPS (Sigma-Aldrich) failed to affect the amounts of LYP4 and LYP6 precipitated by PGNXoo or chitin beads (Figure 2D). These data reinforced that LYP4 and LYP6 selectively bind PGN and chitin but not LPS. Furthermore, LYP4 and LYP6 expressed in rice protoplasts also could be precipitated by different bacterial PGNs and commercial chitin products (Figure 2E), confirming the physical association of PGN and chitin to LYPs in rice. By contrast, the CEBiP proteins successfully expressed in rice protoplasts could be precipitated only by chitin but not PGN, while the full-length rice CERK1 proteins expressed in rice protoplasts could be pulled down by neither MAMP (Figure 2E).[1]
- Silencing of LYP6 Compromises Diverse PGN- and Chitin-Induced Defense Responses in Rice
The findings regarding the plasma membrane localization of LYP4 and LYP6 and their physical interactions with both PGN and chitin pointed to a more exciting possibility that these proteins may be not only the PGN receptors but also previously unknown chitin receptors in rice. This motivated us to evaluate both PGN- and chitin-induced defense responses in LYP4 or LYP6 RNAi or overexpressing (OX) transgenic rice. Two representative lines of RNAi or OX transgenic rice were used for each gene in the subsequent analysis. The empty vector transgenic rice and the wild-type rice were used as controls. Although LYP4 and LYP6 genes share ;80% identity, the LYP4 and LYP6 transcripts were specifically reduced by ;80% by their cognate RNAi construct in the silencing lines (see Supplemental Figure 5A online). Notably, the expression of CEBiP and CERK1, the two known genes involved in rice chitin perception, was affected by neither RNAi construct (see Supplemental Figure 5B online). On the other hand, the expression of LYP4 in its OX lines was increased by 5.2- to 6.8-fold and that of LYP6 in its OX lines was increased by 14- to 15-fold (see Supplemental Figure 5C online). Using LYP RNAi or OX transgenic rice, we investigated three different cell responses occurring at different defense time points after PGN or chitin exposure. These defense responses included ROS generation (a very early response), defense gene activation (an early response), and callose deposition (a late response). LPS, as another glycoconjugate elicitor, was used as control MAMP during the examination of the two earlier defense responses. Indeed, the amounts of ROS triggered by the purified PGNXoo or its HPLC-purified muropeptides declined by ;40% in either LYP4 or LYP6 RNAi transgenic rice when compared with that produced in the control rice (Figure 3A). Similarly, the amounts of ROS induced by chitin or its soluble fragment N-acetylchitohexaose also decreased by 37 to 42% in LYP RNAi transgenic rice (Figure 3A). By contrast, LYP4 and LYP6 OX transgenic rice showed comparable ROS production after PGN or chitin treatment (Figure 3A). Notably, all transgenic rice lines and the wild-type rice treated with LPS exhibited no significant difference in ROS production (Figure 3A). Since ROS generation is one of the earliest defense responses[10], these results suggested that LYP4 and LYP6 function quite upstream within the PGN- and chitin-induced defense signaling pathways and strongly corroborated the notion that these proteins are potential receptors for the two MAMPs. Moreover, the activation of four representative defense marker genes, namely, Beta-Glu, MLO, WRKY13, and PAL, by 30 min PGN/muropeptides or chitin/N-acetylchitohexaose treatment in wild-type rice was substantially suppressed in LYP4 or LYP6 RNAi transgenic rice (Figure 3B). However, these defense marker genes responded to 30-min LPS treatment equally well in both wild-type and LYP RNAi transgenic rice (Figure 3B). Furthermore, the callose staining spots on rice young leaves after PGNXoo muropeptides or N-acetylchitohexaose treatment were dramatically reduced in LYP4 or LYP6 RNAi transgenic rice when compared with those in the control rice (Figure 3C). By contrast, PGNXoo muropeptides or N-acetylchitohexaose treatment resulted in more callose deposition in LYP4 or LYP6 OX transgenic rice than in the control rice (Figure 3C). These data further strengthened the notion that LYP4 and LYP6 play crucial roles in PGN- and chitin-induced defense signaling in rice.[1]
- LYP6 Affects Rice Susceptibility to Both Bacterial and Fungal Pathogens
To validate the importance of LYP4 and LYP6 in rice innate immunity, we conducted pathogen growth assay in LYP RNAi or OX transgenic rice using the bacterial blight pathogen X. oryzae, the bacterial streak pathogen Xanthomonas oryzicola, and the fungal blast pathogen Magnaporthe oryzae. As expected, compared with the lesion area caused by X. oryzae infection in wild-type rice, the lesion areas in LYP4- or LYP6-silencing rice were significantly enlarged (Figures 4A and 4B). Accordingly, bacterial growth was increased by 25- to 50-fold in LYP silencing rice (Figure 4C). Similarly, the lesion area due to X. oryzicola infection also expanded significantly in LYP4- or LYP6- silencing rice relative to that in wild-type rice (Figures 4D and 4E). Consistent with this, more X. oryzicola growth was detected in LYP silencing rice (Figure 4F). In addition, the LYP silencing rice appeared to be more susceptible to fungal M. oryzae infection as the lesion size per leaf was considerably larger in LYP silencing rice than in wild-type rice (Figures 4G and 4H). Accordingly, the lesion number per leaf was increased by 0.9- to 1.4-fold in LYP silencing rice compared with that in wild-type rice (Figure 4I). These data suggested that knockdown of LYP4 and LYP6 expression in rice results in an increased susceptibility to both bacterial and fungal pathogens. Conversely, the lesion area caused by X. oryzae infection decreased to 41 to 58% in LYP4 or LYP6 OX rice relative to that in wild-type rice (Figure 4B) and the X. oryzae growth in these LYP OX lines was reduced by more than 80% (Figure 4C). More significantly, the lesion size caused by X. oryzicola infection decreased to 15 to 38% in LYP OX rice relative to that in wildtype rice (Figure 4E), and the bacterial growth in these LYP OX rice dropped by more than 90% (Figure 4F). Moreover, the lesion area due to M. oryzae infection also shrank to 19 to 40% in LYP OX rice (Figure 4H), and the lesion number per leaf was reduced to 50 to 63% in comparison with that in wild-type rice (Figure 4I). These results indicated that upregulation of LYP4 and LYP6 expression in rice leads to an enhanced resistance against both bacterial and fungal infection.[1]


Expression
- LYP6 Expression Can Be Induced Quickly by Bacterial Pathogen Infection or Diverse MAMPs
To understand better the function of LYP4 and LYP6, we checked the expression profiles of these genes in different rice tissues and developmental stages by quantitative realtime PCR (qPCR). LYP4 and LYP6 were most abundantly expressed in rice callus cells, and both transcripts progressively decreased during maturation (Figure 1D). Furthermore, analysis of LYP4 and LYP6 expression patterns in Promoter:GUS (for b-glucuronidase) transgenic rice demonstrated strong GUS staining in young seedlings, particularly in the root meristem region and the lateral root primordium (see Supplemental Figure 2 online), resembling the expression patterns of their ortholog LYM1 in M. truncatula [8]. Interestingly, expression of LYP4 and LYP6 in rice seedlings, mature leaves, and roots could be quickly induced upon exposure to the rice bacterial pathogen Xanthomonas oryzae pv oryzae. In 5-d-old rice seedlings, 1 h X. oryzae treatment induced a 24- and 26-fold increase of endogenous LYP4 and LYP6 transcripts, respectively (Figure 1E). Moreover, incubation with X. oryzae suspension for 2 h rendered a strong GUS activity in mature leaves and roots of the Promoter:GUS transgenic rice, while incubation with sterile water had no effect on the GUS activity (Figure 1F). The possibility of false positive GUS staining due to pathogen contamination could be excluded as no GUS activity could be detected in the empty vector (pCAMBIA-1391Z containing an intact GUS gene.
Evolution
- The promiscuity of Os-LYP4 and Os-LYP6 in binding carbohydrate MAMPs PGN and chitin is reminiscent of At-FLS2 binding three different ligands, including flagellin and Ax21 as well as the endogenous CLV3 peptide [11]
[12] The promiscuity of PRRs in sensing multiple MAMPs provides a distinct physiological advantage to the host so that a limited number of PRRs would be able to perceive a maximum number of MAMPs. Considering plants in nature are often exposed concurrently to several groups of microbial pathogens, the advantage brought by LYP4 and LYP6 in rice is particularly spectacular in that these PRRs could detect PGN and chitin derived individually from the two major microbial groups, bacteria and fungi. As the expression of LYP4 and LYP6 genes could be rapidly upregulated upon recognition of either MAMP(Figure 1G), it seems that either type of microbial infection would quickly sensitize rice for further infection by both groups of microbes. Interestingly, although the transgenic rice overexpressing LYP4 or LYP6 indeed demonstrated an enhanced pathogen resistance (Figure 4), the PGN- or chitininduced ROS production in these rice plants did not show significant difference compared that in wild-type rice (Figure 3A). Pathogen resistance is a complicated consequence of innate immunity, whereas ROS production is just one of the very early defense responses in plant innate immunity. The biological significance of ROS production in plant defense is not fully understood. It is very likely that the intensity of ROS generation is not linearly correlated with the magnitude of pathogen resistance, as the latter is shaped by complex interplay between multiple layers of defense responses that are induced by a cocktail of MAMPs from pathogens and occur with distinct dynamics. The functions of other three evolutionarily related Os-LYPs (LYP2, LYP3, and LYP5) still remain enigmatic. LYP2 and LYP5 are presumably located in the apoplastic space rather than the plasma membrane due to lack of the GPI anchor (Figure 1A), whereas LYP3 likely resides at the plasma membrane like LYP4 and LYP6. LYP2, LYP3, and LYP5 all contain two intact LysMs (Figure 1A; see Supplemental Figure 1 online); thus, their binding capacity to PGN or chitin cannot be excluded at this moment. By inference, they may serve certain regulatory functions in PGN or chitin signaling in rice, similar to the case of Eix1 and Eix2 proteins in tomato. Both Sl-Eix1 and Sl-Eix2 were found to bind the fungal elicitor xylanase, where only the Eix2 receptor mediated defense signaling while Eix1 acted as a decoy receptor to attenuate the xylanase-induced defense signaling [13][14].Both Os-LYP4 and Os-LYP6 are likely to be N-glycosylated when expressed in rice cells as the GFP hybrids of both proteins showed an actual molecular mass around 100 kD instead of the predicted molecular mass of 65 kD (Figure 1C), reminiscent of other LYP proteins expressed in planta [4] [8][7] Intriguingly, the N-glycosylation of these receptors appeared to be dispensable for ligand binding since the recombinant receptors expressed in E. coli were competent in PGN and chitin binding (Figure 2A). Similar findings were also obtained for their orthologs in Arabidopsis [7]. Recently, it has been revealed that the Nglycosylation of NFP, the LysM-RLK for Nod factor perception, was not essential for its biological activity including the ligand binding [15]. Our work and others thus suggest a role of N-glycosylation in regulating the protein trafficking of these receptors. Although Os-LYP4 or Os-LYP6 are each able to bind PGN and chitin, our data suggest that they are not functionally redundant. This was because knockdown of single LYP gene expression in rice was sufficient to impair both PGN- and chitin-induced defense responses significantly (Figure 3) and to cause severe bacterial or fungal infection phenotypes (Figure 4). Similar observations recently have been made for Arabidopsis PGN receptors LYM1 and LYM3, where lym1 lym3 double mutant did not show increased susceptibility to bacterial pathogens when compared with lym single mutants [7]. Both studies favor a cooperative relationship between the pair of PRRs, suggesting that they may work in the same receptor complex. The identification of LYP4 and LYP6 as additional chitin receptors in rice raises the question regarding their relationship with the previously identified rice chitin receptor CEBiP[4]. Kaku and coworkers found that 70% of the genes activated by chitin in wild-type rice cells lost responsiveness in CEBiP-RNAi rice cells, suggesting that CEBiP might play a major role in rice chitin perception. In line with this speculation, RNAi silencing of CEBiP diminished the chitin-induced ROS generation by 85% , while silencing of LYP4/6 only reduced the chitin-induced ROS generation by ;40% (Figure 3A). However, as 30% of the upregulated genes and 20% of the downregulated genes could respond to chitin equally well in wildtype and in CEBiP-RNAi rice cells, CEBiP and LYP4/6 proteins are very likely to work in different chitin receptor complexes. In support of this speculation, a major portion of CEBiP proteins were visualized as homodimers in blue native PAGE analysis [5]. Nevertheless, a small fraction of CEBiP proteins did exist as larger-size oligomers[5], which makes it ambiguous whether some CEBiP proteins can be in the same complexes with LYP4/6. Further investigation of the composition and stoichiometry of rice chitin receptor complexes as well as chitin responses in CEBiP and LYP4/6 triple knockdown rice will be necessary to dissect fully their contributions in rice chitin perception and signaling. In sum, two LysM-containing proteins, LYP4 and LYP6, are dual function receptors for bacterial PGN and fungal chitin in rice. We provided three key lines of evidence pertaining to the function of these proteins. First, they are localized at plant cell surface. Second, they can specifically bind PGN and chitin. Third, knockdown of their expression perturbs the PGN- and chitin-induced defense responses in rice. LYP4 and LYP6 are unique among known PRRs in that they can recognize MAMPs across microbial groups. Future investigation of the LysMs in these PRRs will be meaningful not only for understanding the biochemical basis of LysM-PGN and LysM-chitin interactions, but also for guiding the engineering of promiscuous PRRs into other crop species to improve disease resistance. Moreover, comparison of the similarities and differences in PGN and chitin perception machineries between rice and Arabidopsis will provide valuable evolutionary insights for understanding critical mechanisms underlying innate immunity signaling in plants.[1]