Difference between revisions of "Os03g0111300"
(→Function) |
(→Structure information=) |
||
| Line 15: | Line 15: | ||
Transfer lipids across membranes. May play a role in plant defense or in the biosynthesis of cuticle layers. | Transfer lipids across membranes. May play a role in plant defense or in the biosynthesis of cuticle layers. | ||
=='''Structure information'''=== | =='''Structure information'''=== | ||
| + | The structure of nsLTP2 was obtained using | ||
| + | 813 distance constraints, 30 hydrogen bond constraints, | ||
| + | and 19 dihedral angle constraints. Fifteen of the 50 random | ||
| + | simulated annealing structures satisfied all of the | ||
| + | constraints and possessed good nonbonded contacts. | ||
| + | The novel three-dimensional fold of rice nsLTP2 contains | ||
| + | a triangular hydrophobic cavity formed by three | ||
| + | prominent helices. The four disulfide bonds required for | ||
| + | stabilization of the nsLTP2 structure show a different | ||
| + | pattern of cysteine pairing compared with nsLTP1. The | ||
| + | C terminus of the protein is very flexible and forms a cap | ||
| + | over the hydrophobic cavity. Molecular modeling studies | ||
| + | suggested that the hydrophobic cavity could accommodate | ||
| + | large molecules with rigid structures, such as | ||
| + | sterols. The positively charged residues on the molecular | ||
| + | surface of nsLTP2 are structurally similar to other | ||
| + | plant defense proteins. | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
[[File:110.gif]] | [[File:110.gif]] | ||
Revision as of 13:54, 8 June 2014
Please input one-sentence summary here.
Contents
Annotated Information
NsLTPs bind to a variety of lipid molecules and catalyze their transfer across membranes in vitro [2]. NsLTPs also have additional biological functions, including biosynthesis of cutin, involvement in defense against pathogens, and managing abiotic stress conditions imparted by temperature or drought [2] and [3]. The nsLTP superfamily possesses eight highly conserved cysteine residues forming four disulfide bonds [1] and [4]. NsLTPs are subdivided into two subfamilies that differ in molecular mass, nsLTP1 (9 kDa), and nsLTP2 (7 kDa) [1].
Sequence similarities
Belongs to the plant LTP family. B11E subfamily.
Mass spectrometry
Molecular mass is 7001.8 Da from positions 1 - 69. Determined by ESI.[5]
You can also add sub-section(s) at will.
Function
Transfer lipids across membranes. May play a role in plant defense or in the biosynthesis of cuticle layers.
Structure information=
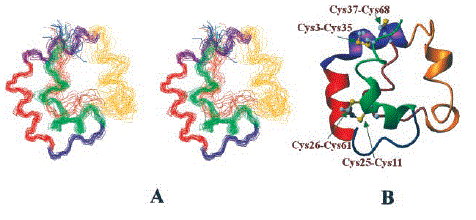
The structure of nsLTP2 was obtained using 813 distance constraints, 30 hydrogen bond constraints, and 19 dihedral angle constraints. Fifteen of the 50 random simulated annealing structures satisfied all of the constraints and possessed good nonbonded contacts. The novel three-dimensional fold of rice nsLTP2 contains a triangular hydrophobic cavity formed by three prominent helices. The four disulfide bonds required for stabilization of the nsLTP2 structure show a different pattern of cysteine pairing compared with nsLTP1. The C terminus of the protein is very flexible and forms a cap over the hydrophobic cavity. Molecular modeling studies suggested that the hydrophobic cavity could accommodate large molecules with rigid structures, such as sterols. The positively charged residues on the molecular surface of nsLTP2 are structurally similar to other plant defense proteins.
Reference
[1]J.P Douliez, T Michon, K Elmorjani, D Marion Structure, biological and technological functions of lipid transfer proteins and indolines, the major lipid binding proteins from cereal kernels J. Cereal Sci., 32 (2000), pp. 1–20
[2]J.C Kader Lipid transfer proteins in plants Annu. Rev. Plant Physiol. Plant Mol. Biol., 47 (1996), pp. 627–654
[3]K.L Larsen, J.R Winther Surprisingly high stability of barley lipid transfer protein, LTP1, towards denaturant, heat, and proteases FEBS Lett., 488 (2001), pp. 145–148
[4]J.P Douliez, C Pato, H Rabesona, D Molle, D Marion Disulfide bond assignment, lipid transfer activity and secondary structure of a 7-kDa plant lipid transfer protein, LTP2 Eur. J. Biochem., 268 (2001), pp. 1400–1403
[5]"Purification and characterization of a novel 7-kDa non-specific lipid transfer protein-2 from rice (Oryza sativa)." Liu Y.-J., Samuel D., Lin C.H., Lyu P.-C.
Structured Information
| Gene Name |
Os03g0111300 |
|---|---|
| Description |
Nonspecific lipid-transfer protein 2 (nsLTP2) (7 kDa lipid transfer protein) |
| Version |
NM_001055258.1 GI:115450244 GeneID:4331362 |
| Length |
476 bp |
| Definition |
Oryza sativa Japonica Group Os03g0111300, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 3:630670..631145 |
| Sequence Coding Region |
630809..631099 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008396:630670..631145 source=RiceChromosome03 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008396:630670..631145 source=RiceChromosome03 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>atgatgaggaagttggcggtgttggtgttggcggtggcgatggtggcggcgtgcggcggcggcgtcgtgggtgtagcgggggccggttgcaacgctgggcagctgacggtgtgcacgggggcgatcgcgggcggggcgcggccgacggcggcgtgctgctccagcctgcgggcgcagcagggctgcttctgccagttcgccaaggacccgcgctacgggcgctacgtcaacagccccaacgcccgcaaggccgtctcctcctgcggcatcgccctccccacctgccactga</cdnaseq> |
| Protein Sequence |
<aaseq>MMRKLAVLVLAVAMVAACGGGVVGVAGAGCNAGQLTVCTGAIAG GARPTAACCSSLRAQQGCFCQFAKDPRYGRYVNSPNARKAVSSCGIALPTCH</aaseq> |
| Gene Sequence |
<dnaseqindica>47..337#gtacctcgcagcaccaagctagctagcttcgatcagtagctggaggatgatgaggaagttggcggtgttggtgttggcggtggcgatggtggcggcgtgcggcggcggcgtcgtgggtgtagcgggggccggttgcaacgctgggcagctgacggtgtgcacgggggcgatcgcgggcggggcgcggccgacggcggcgtgctgctccagcctgcgggcgcagcagggctgcttctgccagttcgccaaggacccgcgctacgggcgctacgtcaacagccccaacgcccgcaaggccgtctcctcctgcggcatcgccctccccacctgccactgatccatccatccatcgtctcctactcttttattttgtgatgtggtgtacgtgttcgtagtattgtcttggcgtcatctcgtcacgacgcgtatatgcatgcgcagtgcggcttttgaataaaaggggatcgaccgatgtt</dnaseqindica> |
| External Link(s) |