Difference between revisions of "Os01g0771200"
(→References) |
(→Character) |
||
| Line 8: | Line 8: | ||
*Cell-surface pattern recognition receptors (PRRs) are key components of the innate immune response in animals and plants. These receptors typically carry or associate with non-RD kinases to control early events of innate immunity signaling. Despite their importance, the mode of regulation of PRRs is largely unknown. Here we show that the rice PRR, XA21, interacts with XA21 binding protein 24 (XB24), a previously undescribed ATPase. | *Cell-surface pattern recognition receptors (PRRs) are key components of the innate immune response in animals and plants. These receptors typically carry or associate with non-RD kinases to control early events of innate immunity signaling. Despite their importance, the mode of regulation of PRRs is largely unknown. Here we show that the rice PRR, XA21, interacts with XA21 binding protein 24 (XB24), a previously undescribed ATPase. | ||
| − | *The rice PRR, XA21, recognizes the PAMP, Ax21 (Activator of XA21-mediated immunity), which is highly conserved in all sequenced genomes of Xanthomonas and in Xylella | + | *The rice PRR, XA21, recognizes the PAMP, Ax21 (Activator of XA21-mediated immunity), which is highly conserved in all sequenced genomes of Xanthomonas and in Xylella . Previous studies have shown that the intracellular non-RD cytoplasmic kinase domain of XA21 contains intrinsic kinase activity. Phosphorylation of amino acids Ser-686, Thr-688, and Ser-689 of XA21 is required to stabilize the XA21 protein . To date, three XA21 binding (XB) proteins—XB3 (an E3 ubiquitinligase), XB10 (OsWRKY62), and XB15 (a PP2C phosphatase)—have been shown to regulate XA21-mediated immunity .XB24 associates with XA21 in vivo and modulates XA21 function. XB24 belongs to a large class of broadly conserved ATPases of unknown function. The association between XB24 and XA21 is compromised upon inoculation of the Xanthomonas oryzae pv. Oryzae (Xoo) strain PXO99, which secretes the Ax21 PAMP . XB24 promotes autophosphorylation of XA21 through its ATPase activity. Rice plants silenced for Xb24 display enhanced XA21-mediated immunity, whereas rice plants overexpressing XB24 are compromised for immunity. XA21 is degraded in the presence of Ax21 when XB24 is overexpressed. These findings reveal that XB24 negatively regulates XA21 PRR function. |
===Function=== | ===Function=== | ||
Revision as of 01:29, 9 June 2014
Gene Os01g0771200,namely XB24, is a XA21 binding protein gene .XB24 is a unique ATPase from a previously unclassified subclass. It does not belong to any of these previously described superfamilies of ATPases or HSPs.
Contents
Annotated Information
Character
(Don't Delete)
- Cell-surface pattern recognition receptors (PRRs) are key components of the innate immune response in animals and plants. These receptors typically carry or associate with non-RD kinases to control early events of innate immunity signaling. Despite their importance, the mode of regulation of PRRs is largely unknown. Here we show that the rice PRR, XA21, interacts with XA21 binding protein 24 (XB24), a previously undescribed ATPase.
- The rice PRR, XA21, recognizes the PAMP, Ax21 (Activator of XA21-mediated immunity), which is highly conserved in all sequenced genomes of Xanthomonas and in Xylella . Previous studies have shown that the intracellular non-RD cytoplasmic kinase domain of XA21 contains intrinsic kinase activity. Phosphorylation of amino acids Ser-686, Thr-688, and Ser-689 of XA21 is required to stabilize the XA21 protein . To date, three XA21 binding (XB) proteins—XB3 (an E3 ubiquitinligase), XB10 (OsWRKY62), and XB15 (a PP2C phosphatase)—have been shown to regulate XA21-mediated immunity .XB24 associates with XA21 in vivo and modulates XA21 function. XB24 belongs to a large class of broadly conserved ATPases of unknown function. The association between XB24 and XA21 is compromised upon inoculation of the Xanthomonas oryzae pv. Oryzae (Xoo) strain PXO99, which secretes the Ax21 PAMP . XB24 promotes autophosphorylation of XA21 through its ATPase activity. Rice plants silenced for Xb24 display enhanced XA21-mediated immunity, whereas rice plants overexpressing XB24 are compromised for immunity. XA21 is degraded in the presence of Ax21 when XB24 is overexpressed. These findings reveal that XB24 negatively regulates XA21 PRR function.
Function
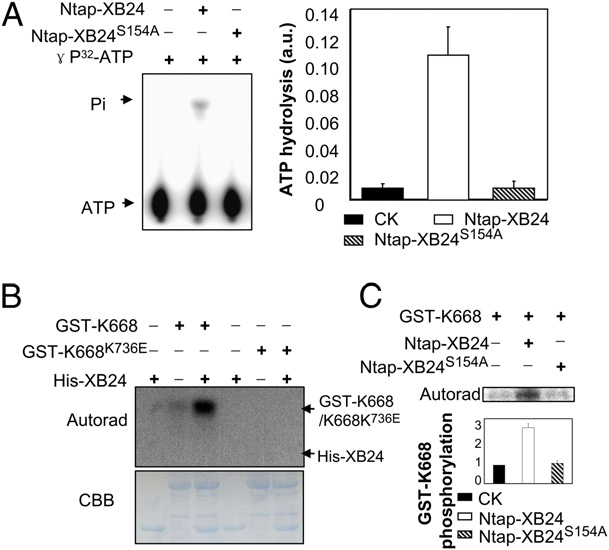
- XB24 ATPase Enhances Autophosphorylation of XA21K668. The author tested whether XB24 is a substrate of XA21 or affects XA21 kinase autophosphorylation. Purified His-XB24 and GST-XA21K668 were co-incubated in the presence of [32P]ATP for kinase analysis. For a control, the purified His-XB24 was co-incubated with GST-XA21K668K736E, a catalytically inactive mutant[1]. As ( Fig. 2B ) shows, the GST-XA21K668 autophosphorylates as expected, whereas His-XB24 does not autophosphorylate or become transphosphorylated by GST-XA21K668. The phosphorylation of GSTXA21K668 is highly enhanced in the presence ofHis-XB24 protein.No phosphorylation of GST-XA21K668K736E can be detected in reactions carried out in the presence of absence of His-XB24. These results demonstrate that XB24 promotes XA21K668 autophosphorylation.To test whether XB24 promotes autophosphorylation of intact, native XA21 protein, the immunoprecipitated ProAXA21 protein from rice tissue described above (0, 1, or 2 days post-PXO99 inoculation) was co-incubated with the purified His-XB24 for kinase autophosphorylation analyses.These results demonstrate that XB24 promotes autophosphorylation of the native XA21 protein. Furthermore, XB24 is not transphosphorylated by the XA21 protein with or without PXO99 inoculation. To test whether the ATPase activity of XB24 is required for promoting XA21K668 autophosphorylation, the purified Ntap-XB24 and NtapXB24S154A were incubatedwithGST-taggedXA21K668 in the presence of [32P]ATP for kinase analyses. Autophosphorylation of GST-XA21K668 is enhanced in the presence of rice-expressed Ntap-XB24 but not Ntap-XB24S154A (Fig. 2C). Autophosphorylation of the GST-XA21K668 fusion protein is also enhanced in the presence of the His-XB24 protein but not His-XB24S154A. These results demonstrate that XB24 enhances XA21 autophosphorylation and that itsATPase activity is required for this function[2].
- XB24 Possesses ATPase Activity. Because XB24 contains aC-terminal ATPasemotif and the residue serine (Ser) 154 is a predicted key site for this motif, we tested whether it indeed possesses intrinsic ATPase activity.Wepurified Ntap-XB24 andNtap-XB24S154A(containing a single amino acid change of Serine 154 by Alanine) from Ntap-Xb24 and Ntap-XB24S154A transgenic plants, respectively, and performed the ATP hydrolysis assay. As shown in Fig. 2A, Ntap-XB24 displayed significant ATP hydrolysis activity, whereas Ntap-XB24S154A had only negligible ATPase activity.We also found that E. coli-produced recombinant protein His-XB24 possesses ATPase activity and that the S154A mutant completely abolished the ATPase activity of XB24 (Fig. S4). Taken together, these results show that the XB24 protein possesses an ATPase activity and that amino acid S154 is essential for its ATPase activity.
- Silencing of Xb24 Enhances Xa21-Mediated Resistance. To investigate the biological function of XB24, we used the RNA interference(RNAi) approach[3][4] to silence the Xb24 gene and monitored its effects on disease resistance. They developed two independent lines, Xb24RNAi-3 and Xb24RNAi-9, each containing a single-locus insertion, using the rice cultivar Kitaake as the transgene recipient.RT-PCR analysis revealed that Xb24 transcript levels were significantly reduced in these two lines. Both lines show similar disease lesion lengths compared to the control line Kitaake after challenge with PXO99, indicating that silencing of Xb24 does not affect the susceptibility of Kitaake to Xoo. To explore the role of XB24 in XA21-mediated signaling, they crossed Xb24RNAi-3 and Xb24RNAi-9 with Xa21 lines and obtained one progeny form the Xa21/Xb24RNAi-3 cross and three from the Xa21/Xb24RNAi-9 cross. Our initial results indicated that silencing of Xb24 enhanced resistance. To confirm these results, they developed an F4 line (A176) from one of the F1 plants.TheA176 line carries homozygous Xa21 and homozygous Xb24RNAi-9. They then inoculated 3-week-old A176 plants.As shown in (Fig. 3A), these plants developed much shorter lesion lengths (3 ± 0.9 cm) than the wildtype Xa21 plants (6.8±1.2 cm), which show only partial resistance at the 3-weeks-old (tilling) stage[5]. At test gave a P value of 8.62 ×10−13, showing a highly significant difference. Rice line Xb24RNAi-9 showed similar disease lesion lengths (16 ± 2.5 cm) as Kitaake (P =0.56). Bacterial growth curve analysis revealed that Xa21/Xb24RNAi-9 lines harbor 3.2-fold less Xoo bacteria (1.48 × 107 ±1.2 × 106) in their leaves than the Xa21 lines (4.8 × 107±4.4 × 106) at 12 days postinoculation (Fig. 3B), consistent with the leaf lesion length measurements described above. This experiment was repeated three times, and similar results were obtained each time. These results demonstrate that silencing of Xb24 expression enhances XA21-mediated disease resistance[2].
- Overexpression of XB24 Compromises XA21-Mediated Resistance. To investigate the involvement of XB24 in the XA21-mediated signaling, the author created construct Ubi-Xb24 to overexpress XB24 using the maize Ubi-1 promoter. They introduced the Ubi-Xb24 construct directly into an Xa21 (in the TP309 genetic background)line by Agrobacterium-mediated transformation using mannose selection[6] and generated five independent T0 plants. After PCR-based genotyping and RT-PCR-based transcripts expression analyses to confirm that Xb24 is overexpressed, we challenged 6-week-old Xa21 lines with PXO99.They found that all of the five lines have longer disease lesion lengths compared with the wild-type Xa21 plants. Two homozygous lines (Xa21/Xb24ox-1 and -2) fromtwo of these five independent lines were then developed. Overexpression of XB24 (XB24ox) in the progeny from these homozygous lines was confirmed by protein gel blotting analysis(Fig. 4A). Six-week-old plants were challenged with PXO99. Disease lesion lengths on both the Xa21/Xb24ox-1 and -2 lines (7.3 ± 0.5 cmfor line 1 and 6.0 ± 0.5 cmfor line 2) were longer than those observed on Xa21 lines (1.3 ±0.4 cm) (Fig. 4 A and B). The low P values (5.02 × 10−21 for Xa21/Xb24ox-1 and 2.06 × 10−23 for Xa21/Xb24ox-2) indicate that these differences are statistically significant. At 12 days postinoculation,the accumulation of bacterial populations, as measured by bacterial growth curve analysis, in the two Xa21/Xb24ox lines (1.23 × 108 ±1.88 × 107 for Xa21/Xb24ox-1 and 1.08 × 108 ± 1.97 × 107 for Xa21/Xb24ox-2)was clearly higher (>2-fold) than in theXa21 lines (5.20×107 ± 8.9 × 105) (Fig. 4C). Again, the low P values (8.27 × 10−4 forXa21/Xb24ox-1 and 2.72 × 10−3 for Xa21/Xb24ox-2) of bacterial accumulation at 12 days postinoculation indicate that these differences are statistically significant. Rice lines overexpressing Xb24 display similar levels of susceptibility as control lines lacking overexpressed Xb24 in three independent biological replicates.These results demonstrate that overexpression of XB24 compromises XA21-mediated resistance[2].
- Overexpression of Xb24 Causes XA21 Instability Following Ax21 Recognition. To gain insight into the mechanism of XB24-mediated regulation of XA21 function, they tested whether XB24 affects the amount of the XA21 protein after Xoo inoculation. As shown in Fig. 6 A and B, without Xoo inoculation (Mock treatment). Overexpression of XB24(Xa21/Ntap-Xb24) caused no significant decrease in the ProA-XA21 protein level compared to overexpression of Ntap (Xa21/Ntap) alone. In contrast, after inoculation with PXO99, the Xa21/Xb24ox line showed a sharp decrease in the ProA-XA21 protein level. The Xa21/Ntap control line showed amarked increase. When inoculated with the Xoo strain PXO99ΔraxST, the Xa21/Xb24ox sample showed an increase in the ProA-XA21 level similar to that of the Xa21/Ntap control. Similar results were obtained from three biological repeats of this experiment.These results indicate that the sharp decrease in the XA21 protein level is Ax21-specific[2].



Expression
- The XB24 cDNAis expressed from a unique rice gene, Os01g56470, and encodes a 198-aa protein. The predicted secondarystructure has no significant motifs except for a C-terminal ATP synthase α-and β-subunits signature (ATPase) motif with the sequence PSINERESSS.Although 38 human proteins, 43 Arabidopsis proteins, and 67 additional rice proteins are annotated to contain a conserved ATPase motif, none share similarity beyond the ATPase motif with XB24 and most are not functionally characterized. Thus, XB24 belongs to a previously uncharacterized class of ATPases.The only conserved structure in XB24 is the region composed of 10 amino acids PSINERES154SS that is predicted as the ATPase motif, (P-[SAP]-[LIV]-[DNH]-{LKGN}-{F}-{S}-S-{DCPH}-S).
- Silencing of Xb24 Enhances Xa21-Mediated Resistance.
- overexpression of XB24 Compromises XA21-Mediated Resistance.
Evolution
XB24 ATPase enzyme activity is required for XB24 function. XA21 is degraded in the presence of the pathogen-associated molecular pattern Ax21 when XB24 is overexpressed. These results demonstrate a function for this large class of broadly conserved ATPases in PRR-mediated immunity.XB24 promotes autophosphorylation of XA21 through its ATPase activity. Rice lines silenced for Xb24 display enhanced XA21-mediated immunity, whereas rice lines overexpressing XB24 are compromised for immunity.
Other associations with XA21
The association between XB24 and XA21 is compromised upon inoculation of the Xanthomonas oryzae pv. oryzae (Xoo) strain PXO99, which secretes the Ax21 PAMP [7]. XB24 promotes autophosphorylation of XA21 through its ATPase activity. Rice plants silenced for Xb24 display enhanced XA21-mediated immunity, whereas rice plants overexpressing XB24 are compromised for immunity. XA21 is degraded in the presence of Ax21 when XB24 is overexpressed. These findings reveal that XB24 negatively regulates XA21 PRR function.
XB24 Physically Associates with XA21 in Vivo. The author isolated XB24 as an XA21 interacting protein through yeast two-hybrid screening. The XB24 cDNAis expressed froma unique rice gene,Os01g56470, and encodes a 198-aa protein. The predicted secondary structure has no significant motifs except for a C-terminal ATP synthase α- and β-subunits signature (ATPase) motif with the sequence PSINERESSS. Although 38 human proteins, 43 Arabidopsis proteins, and 67 additional rice proteins are annotated to contain a conserved ATPase motif, none share similarity beyond the ATPase motif with XB24 and most are not functionally characterized. Thus, XB24 belongs to a previously uncharacterized class of ATPases.
XB24 Dissociates from XA21 in Response to PXO99 Inoculation.To determine whether XB24 is degraded in response to Xoo strain PXO99 inoculation, they performed a Western blot analysis to detect the XB24 protein before and after inoculation. They found that a similar amount of XB24 protein was detected in Xa21 and Kitaake plants before inoculation and 1 day or 2 days after inoculation. This result shows that XB24 is not degraded in response to Ax21. Wenext investigated whether Ax21 recognition affects the interaction of XA21 and XB24. We performed coimmunoprecipitation experiments withPAP(targeting ProA-XA21)using rice leaf tissues fromtheXa21line inoculated with Xoo strain PXO99 or Xoo strain PXO99ΔraxST, which lacks Ax21 activity due to a knockout of the raxST gene. They then carried out immunoblotting to detect XB24. A similar coimmunoprecipitation was performed using Kitaake rice leaves as a control.
ATPase Activity Is Essential for XB24-Mediated Regulation of XA21 Function. The author tested whether XB24 ATPase activity was required for XB24 to regulate XA21 function. They developed Xa21/Xb24ox and Xa21/Xb24S154Aox plants using NtapXb24ox and NtapXb24S154Aox plants, respectively, to cross with ProAXa21 plants, and inoculated these plantswith PXO99.
Labs working on this gene
- College of Life Science, Zhejiang Sci-Tech University, Hangzhou 310018, China.
- Department of Plant Pathology, University of California, Davis, CA 95616,USA.
References
- ↑ Liu GZ, Pi LY, Walker JC, Ronald PC, Song WY (2002) Biochemical characterization of the kinase domain of the rice disease resistance receptor-like kinase XA21. J Biol Chem 277:20264–20269.
- ↑ 2.0 2.1 2.2 2.3 Xuewei Chen, Mawsheng Chern, Patrick E. Canlas, Deling Ruan, Caiying Jiang, and Pamela C. Ronald(2010) An ATPase promotes autophosphorylation of the pattern recognition receptor XA21 and inhibits XA21-mediated immunity. Proc Natl Acad Sci USA 107: 8029–8034.
- ↑ Fire A, et al. (1998) Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391:806–811.
- ↑ Mourrain P, et al. (2000) Arabidopsis SGS2 and SGS3 genes are required for posttranscriptional gene silencing and natural virus resistance. Cell 101:533–542.
- ↑ Century KS, et al. (1999) Developmental control of Xa21-mediated disease resistance in rice. Plant J 20:231–236.
- ↑ Lucca P, Ye X, Potrykus I (2001) Effective selection and regeneration of transgenic rice plants with mannose as selective agent. Mol Breed 7:43–49.
- ↑ Lee SW, Han SW, Bartley LE, Ronald PC (2006) Unique characteristics of Xanthomonas oryzae pv. oryzae AvrXa21 and implications for plant innate immunity. Proc Natl Acad Sci USA 103:18395–18400.
Structured Information
| Gene Name |
Os01g0771200 |
|---|---|
| Description |
Similar to Mal d 1-associated protein |
| Version |
NM_001050918.1 GI:115440206 GeneID:4324614 |
| Length |
1888 bp |
| Definition |
Oryza sativa Japonica Group Os01g0771200, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 1:34299150..34301037 |
| Sequence Coding Region |
34299283..34299494,34300420..34300804 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008394:34299150..34301037 source=RiceChromosome01 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008394:34299150..34301037 source=RiceChromosome01 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>atgggttggcgttggcacgacgacggcgatgacggcggccgcggcctgggcgacatccccgacctcgccggcggcggcggaggcggagatggggagcgctgcgccacgcgccgggtggtgcagtctcggtgccacacggaggaggtggagcccggccgcttcgtccgcaagtgcgagaagaccgagcagctcctccgcgactgcgtcggcaggccctctgaactggtggaatcaaaaactgaaaatactgaagaagacgtcacagatgaaatgaaaagcgggtcactatctcttggttttccgaccaatgagccctttgcatttcctggacttcgcagtgacatagaagctcttgagaaaggccttttcgggagcattggtagctttctggatgatgctgagaggatgaccaatgatttcttgaagtcttttggtgtcccttccatcaatgaaagggagtcgagctcatttgatggacaacctacaggcaggcacattggtggacaacctgcaggcaggcacattgaggaaggtactgcaaaggacactaaacagaacgactacgcagaattcagcagcaagattacagatgtgtaa</cdnaseq> |
| Protein Sequence |
<aaseq>MGWRWHDDGDDGGRGLGDIPDLAGGGGGGDGERCATRRVVQSRC HTEEVEPGRFVRKCEKTEQLLRDCVGRPSELVESKTENTEEDVTDEMKSGSLSLGFPT NEPFAFPGLRSDIEALEKGLFGSIGSFLDDAERMTNDFLKSFGVPSINERESSSFDGQ PTGRHIGGQPAGRHIEEGTAKDTKQNDYAEFSSKITDV</aaseq> |
| Gene Sequence |
<dnaseqindica>134..345#1271..1655#agaaacgagccggccggattgctctccaagccaaacacggcccagagaggcgagagcccccacaccgccaaacccgacccggaaatcaactgcacacgctccgatcccctctctccagatcgattcggaccccatgggttggcgttggcacgacgacggcgatgacggcggccgcggcctgggcgacatccccgacctcgccggcggcggcggaggcggagatggggagcgctgcgccacgcgccgggtggtgcagtctcggtgccacacggaggaggtggagcccggccgcttcgtccgcaagtgcgagaagaccgagcagctcctccgcgactgcgtcggcaggtatatacatatactccatggcctcgcgcttttccctgatctgttccttcttttcctgcgaggatcccctctttgcatattactgttcgttgttttagtgatgaattggtgataacatgtcatgcgcagttgtgttactattagtcccgtgtatgattgtgagctgtgtatagccgcctctctgtggacgattttcgtgagtctgtcctcgctgatttgatacaaacacgaattcattaagaatggggaatgtgccgtgatgatcgagacgtttgagttctgtggtgaatagtatgatcaaatgtgtgtaataacttgggcttttgaattttggtagctaattagatatggaatttctatccagttttaggtgcccgtgccatcgacatgggttaaaagttattgtgggctattttgcgctcaacttgttaatcaacaaattttggacggaggcagtactttgtaggcatggggtaaatttgtatgatacacttgttgcttgaagatgttcgtgaactcatgggatttctagacacctacaaccaattacattacagtgttattatattctaacgatatgtaaagagcagctgtgttcgtgttgtctacactaaagattcaacaactagtggtcatcatcaaaccgatagcatatcctcattttgcaggattttgaaaactcacatctcgttagcatccttccttttatttcttcacatgcctcatgttaaacttttaagtgcatgaaacccaaatttctgtatttcgtagctccttgtaaccaatgtacaacactatgattagtattctggatagattttgtacttcccaaagtattttggttgttgacaagctattacaataatctgatggataacctaccatacattatttctcctaaccgatcacaataatcttcttctaggccctctgaactggtggaatcaaaaactgaaaatactgaagaagacgtcacagatgaaatgaaaagcgggtcactatctcttggttttccgaccaatgagccctttgcatttcctggacttcgcagtgacatagaagctcttgagaaaggccttttcgggagcattggtagctttctggatgatgctgagaggatgaccaatgatttcttgaagtcttttggtgtcccttccatcaatgaaagggagtcgagctcatttgatggacaacctacaggcaggcacattggtggacaacctgcaggcaggcacattgaggaaggtactgcaaaggacactaaacagaacgactacgcagaattcagcagcaagattacagatgtgtaaggatctacagttagctgacgcacctttgggagcagcttgccaattttgtattttgaacatctccatggttgtaattggaaggggaaggatcagtttgactgttttataagcagagtcgtctgaagtctgaagtgttgcgcttataagaacaattgtgtatattactgttttagataagcctgtttgtgttcctcaacaatgagatcaattatggatggtttttttctctgctt</dnaseqindica> |
| External Link(s) |