Difference between revisions of "Os09g0449000"
(→Mutation) |
(→Microarray analysis) |
||
| Line 29: | Line 29: | ||
===Microarray analysis=== | ===Microarray analysis=== | ||
Please input Microarray analysis here. | Please input Microarray analysis here. | ||
| + | Microarray analysis revealed 2,943 genes with a 2-fold or greater expression change over the two developmental stages in ptc1 anthers compared with the wild type (570 down-regulated and 278 up-regulated at stage 8; 1,252 down-regulated and 843 up-regulated at stage 9. | ||
| + | The 2,417 genes can be group into four COG (for Cluster of Orthologous Groups of proteins) categories: I, information storage and processing (144 genes); II, cellular processes and signaling (363 genes); III, metabolism (551 genes); and IV, poorly characterized genes (1,359 genes) | ||
| + | |||
| + | |||
| + | [[File:p5.jpg]] | ||
===Evolution=== | ===Evolution=== | ||
Revision as of 00:42, 6 May 2014
Please input one-sentence summary here.
Contents
Annotated Information
Function
Please input function information here. PERSISTANT TAPETAL CELL1 (PTC1), which controls programmed tapetal development and pollen exine formation in rice. PTC1 encodes a PHD-finger (for plant homeodomain) protein, which is expressed specifically in tapetal cells and microspores during anther development in stages 8 and 9, when the wild-type tapetal cells initiate a typical apoptosis-like cell death[1]. PTC1 may functions downstream of GAMYB[2], and in parallel with TDR in regulating programmed anther development and pollen formation[3].
Mutation
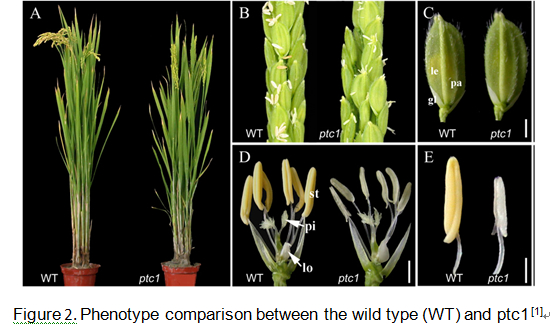
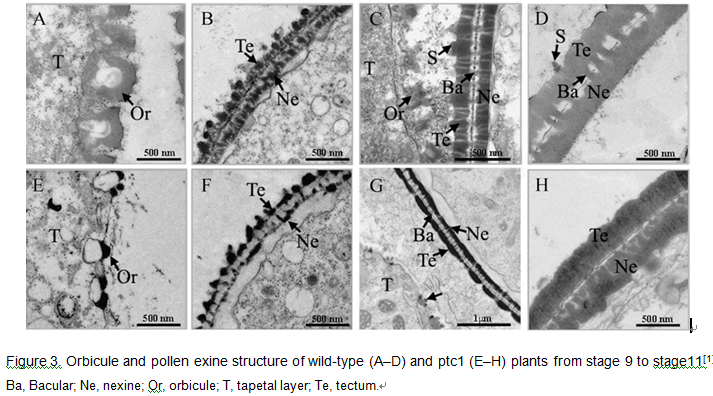
Please input mutation information Compared with wild-type plants, ptc1 exhibited normal vegetative development (Fig. 2A). During the reproductive stage, ptc1 plants developed normal panicles and floral organs (Fig. 2, B–D), but ptc1 anthers were smaller, white, and lacked viable pollen grains (Fig. 2, D and E).
From stage 9 (free microspore released from the tetrad) to early stage 10 (vacuolated microspore), ptc1 tapetal cells were more vacuolated than wild-type. From late stage 10 to stage 11 (mitosis I), ptc1 tapetum became abnormally enlarged, and cytosolic constituents seemed to be secreted into the anther locule. At stage 13, in the ptc1 anther, only cell debris of both tapetal cells and pollen grains remained. The ptc1 tapetum also appeared to have less accumulation of pollen wall precursor materials
Expression
Please input expression information
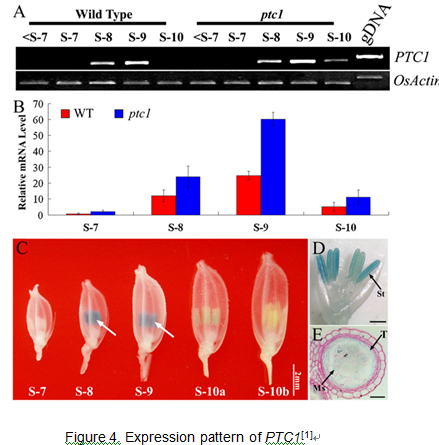
According to the RiceGE (Gene Expression Atlas Data Sources), PTC1 expression is highest in the 10- to 15-cm inflorescence (stages 7–10), gradually decreasing in the 15- to 20-cm inflorescence (stages 10–12). The data from laser microdissection coupled with microarray analysis also suggest that PTC1 is expressed in both the tapetum and microspores at stage 8. Reverse transcription (RT)-PCR and quantitative RT-PCR (RT-qPCR) results indicated that PTC1 was preferentially expressed in the anther from stage 8 to stage 9. In transgenic plants expressing the PTC1pro:GUS fusion, GUS staining was specifically detected in the anther from stages 8 to 9 and at a low level in stage 10. No GUS expression was seen in the palea, lemma, pistil or other organs, but GUS was also apparent within the microspores,
Microarray analysis
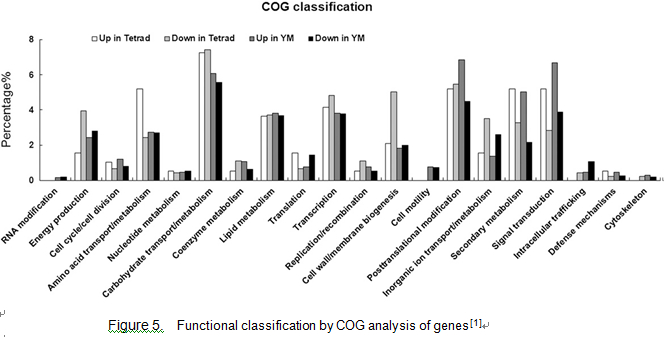
Please input Microarray analysis here. Microarray analysis revealed 2,943 genes with a 2-fold or greater expression change over the two developmental stages in ptc1 anthers compared with the wild type (570 down-regulated and 278 up-regulated at stage 8; 1,252 down-regulated and 843 up-regulated at stage 9. The 2,417 genes can be group into four COG (for Cluster of Orthologous Groups of proteins) categories: I, information storage and processing (144 genes); II, cellular processes and signaling (363 genes); III, metabolism (551 genes); and IV, poorly characterized genes (1,359 genes)
Evolution
Please input evolution information here.
You can also add sub-section(s) at will.
Labs working on this gene
Please input related labs here.
References
Please input cited references here.
Structured Information
| Gene Name |
Os09g0449000 |
|---|---|
| Description |
Protein of unknown function UPF0054 family protein |
| Version |
NM_001069854.2 GI:297609560 GeneID:4347213 |
| Length |
2215 bp |
| Definition |
Oryza sativa Japonica Group Os09g0449000, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 9:17439752..17441966 |
| Sequence Coding Region |
17439752..17440061,17440136..17440442,17440544..17441966 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008402:17439752..17441966 source=RiceChromosome09 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008402:17439752..17441966 source=RiceChromosome09 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>atggcgcctaagatggtgatcagcctggggagctcgcggcggcggaagcgcggcgagatgctgttccggttcgaggccttctgccagcccggctacccggcgaacttcgccggcgccggcggcttcagggacaacgtgaggacgctgctcggcttcgcgcacctggaggccggcgtccacggcgagaccaagtgctggtcgttccagctcgagctgcaccgccacccccccaccgtcgtgaggctcttcgtcgtcgaggaggaggtcgccgcctcgccgcaccgccagtgccacctctgccgccatattgggtgggggaggcatctgatatgcagcaagaggtatcacttcttgctgccgaggagggaatcggcggcggaagccgacggcctgtgcttcgcgatcaaccacggcggcggcggtggcgcggagaaagcgtcgtcgaaagggacgacgacgacggcctccagcagaggccacctgctacacggcgtcgtgcacctcaacggctacggccacctcgtcgccctccacggcctcgagggcggctccgacttcgtctccggccaccagatcatggacctctgggaccgcatttgctcagccttgcacgtaaggacggtgagcctggtggacacggcgaggaagggccacatggagctgaggctgctgcacggcgtcgcgtacggcgagacgtggttcgggcggtgggggtacaggtacggccggccgagctacggcgtcgcgctgccgtcgtaccggcagtcgctgcacgtgctcggctccatgccgctctgcgtgctggtgccgcacctgtcgtgcttcagccaggagctccccatggtggtcaccaagtaccaggccatcagcggccacaagctgctcagcctcggcgacctcctccgcttcatgctcgagctgcgcgcccgcctgccggccacctccgtcacggccatggactaccggggcatcatgtcggaggcctcgtgccggtggtcggcgaagcgcgtcgacatggcggcgcgcgccgtcgtggacgcgctccgccgcgccgagccggcggcgcggtgggtcacgcggcaggaggtgcgcgacgcggcgcgcgcctacatcggcgacacgggcctcctcgacttcgtgctcaagtccctcggcaaccacatcgtcggcaactacgtcgtgcgccgcaccatgaacccggtgaccaaggtgctcgagtactgcctcgaggacgtctccagcgtgctcccggcggtcgccgccggcggcggcgtgccggcgcagggcaagatgagggtgaggttccagctcacgcgtgcgcagctcatgagggacctggtgcacctgtaccggcacgtgctcaaggagcccagccaggcgctcaccggcggcgcgttcggcgcgatcccggtggcggtgcggatggtcctggacatcaagcacttcgtcaaagattaccacgaaggacaagccgcggcgagcagcaatggcggtggcggattcgggcatccccacatcaacctgtgctgcacgctgctcgtgagcaacgggagcccggagctagctccaccgtacgagacggtgaccctgccggcgcacgcgacggtgggcgagctgaagtgggaggcgcagagggtgttcagcgagatgtacctcggcctgaggagcttcgcggcggactccgtcgtcggggtcggcgccgaccaggagggcctcccggtgctcgggctggtcgacgtcggaagcgccgtcgtggtgcaagggagcgtgggcgagcagataaacggggaggaccacgagaggaaggaggaggcggcggcggcggccgtgtgcgaggggagcggcggcggcgagcgcgtcgtggactgcgcgtgcggcgcggtggacgacgacggcgagcgcatggcgtgctgcgacatctgcgaggcgtggcagcacacgcggtgcgccgggatcgcggacaccgaggacgcgccgcacgtcttcctctgcagccggtgcgacaacgacgtcgtgtcgttcccgtccttcaactgttag</cdnaseq> |
| Protein Sequence |
<aaseq>MAPKMVISLGSSRRRKRGEMLFRFEAFCQPGYPANFAGAGGFRD NVRTLLGFAHLEAGVHGETKCWSFQLELHRHPPTVVRLFVVEEEVAASPHRQCHLCRH IGWGRHLICSKRYHFLLPRRESAAEADGLCFAINHGGGGGAEKASSKGTTTTASSRGH LLHGVVHLNGYGHLVALHGLEGGSDFVSGHQIMDLWDRICSALHVRTVSLVDTARKGH MELRLLHGVAYGETWFGRWGYRYGRPSYGVALPSYRQSLHVLGSMPLCVLVPHLSCFS QELPMVVTKYQAISGHKLLSLGDLLRFMLELRARLPATSVTAMDYRGIMSEASCRWSA KRVDMAARAVVDALRRAEPAARWVTRQEVRDAARAYIGDTGLLDFVLKSLGNHIVGNY VVRRTMNPVTKVLEYCLEDVSSVLPAVAAGGGVPAQGKMRVRFQLTRAQLMRDLVHLY RHVLKEPSQALTGGAFGAIPVAVRMVLDIKHFVKDYHEGQAAASSNGGGGFGHPHINL CCTLLVSNGSPELAPPYETVTLPAHATVGELKWEAQRVFSEMYLGLRSFAADSVVGVG ADQEGLPVLGLVDVGSAVVVQGSVGEQINGEDHERKEEAAAAAVCEGSGGGERVVDCA CGAVDDDGERMACCDICEAWQHTRCAGIADTEDAPHVFLCSRCDNDVVSFPSFNC</aaseq> |
| Gene Sequence |
<dnaseqindica>1..310#385..691#793..2215#atggcgcctaagatggtgatcagcctggggagctcgcggcggcggaagcgcggcgagatgctgttccggttcgaggccttctgccagcccggctacccggcgaacttcgccggcgccggcggcttcagggacaacgtgaggacgctgctcggcttcgcgcacctggaggccggcgtccacggcgagaccaagtgctggtcgttccagctcgagctgcaccgccacccccccaccgtcgtgaggctcttcgtcgtcgaggaggaggtcgccgcctcgccgcaccgccagtgccacctctgccgccatattggtccgtcgaacaaactacaattaatcaatcaacctttacataggattgatccgatcgatgccatggtgttgtagggtgggggaggcatctgatatgcagcaagaggtatcacttcttgctgccgaggagggaatcggcggcggaagccgacggcctgtgcttcgcgatcaaccacggcggcggcggtggcgcggagaaagcgtcgtcgaaagggacgacgacgacggcctccagcagaggccacctgctacacggcgtcgtgcacctcaacggctacggccacctcgtcgccctccacggcctcgagggcggctccgacttcgtctccggccaccagatcatggacctctgggaccgcatttgctcagccttgcacgtaaggtagtagtagtatacatgtgcgtgtgcatgcatgcaagcaatgcaacgatgtcgggctgcgtgtgagaacatttgcttgggcatggtgtggtgtatgcaaggacggtgagcctggtggacacggcgaggaagggccacatggagctgaggctgctgcacggcgtcgcgtacggcgagacgtggttcgggcggtgggggtacaggtacggccggccgagctacggcgtcgcgctgccgtcgtaccggcagtcgctgcacgtgctcggctccatgccgctctgcgtgctggtgccgcacctgtcgtgcttcagccaggagctccccatggtggtcaccaagtaccaggccatcagcggccacaagctgctcagcctcggcgacctcctccgcttcatgctcgagctgcgcgcccgcctgccggccacctccgtcacggccatggactaccggggcatcatgtcggaggcctcgtgccggtggtcggcgaagcgcgtcgacatggcggcgcgcgccgtcgtggacgcgctccgccgcgccgagccggcggcgcggtgggtcacgcggcaggaggtgcgcgacgcggcgcgcgcctacatcggcgacacgggcctcctcgacttcgtgctcaagtccctcggcaaccacatcgtcggcaactacgtcgtgcgccgcaccatgaacccggtgaccaaggtgctcgagtactgcctcgaggacgtctccagcgtgctcccggcggtcgccgccggcggcggcgtgccggcgcagggcaagatgagggtgaggttccagctcacgcgtgcgcagctcatgagggacctggtgcacctgtaccggcacgtgctcaaggagcccagccaggcgctcaccggcggcgcgttcggcgcgatcccggtggcggtgcggatggtcctggacatcaagcacttcgtcaaagattaccacgaaggacaagccgcggcgagcagcaatggcggtggcggattcgggcatccccacatcaacctgtgctgcacgctgctcgtgagcaacgggagcccggagctagctccaccgtacgagacggtgaccctgccggcgcacgcgacggtgggcgagctgaagtgggaggcgcagagggtgttcagcgagatgtacctcggcctgaggagcttcgcggcggactccgtcgtcggggtcggcgccgaccaggagggcctcccggtgctcgggctggtcgacgtcggaagcgccgtcgtggtgcaagggagcgtgggcgagcagataaacggggaggaccacgagaggaaggaggaggcggcggcggcggccgtgtgcgaggggagcggcggcggcgagcgcgtcgtggactgcgcgtgcggcgcggtggacgacgacggcgagcgcatggcgtgctgcgacatctgcgaggcgtggcagcacacgcggtgcgccgggatcgcggacaccgaggacgcgccgcacgtcttcctctgcagccggtgcgacaacgacgtcgtgtcgttcccgtccttcaactgttag</dnaseqindica> |
| External Link(s) |