Difference between revisions of "Os02g0220400"
Lymzzu2009 (talk | contribs) |
Lymzzu2009 (talk | contribs) |
||
| Line 1: | Line 1: | ||
| − | The CYTOKININ-RESPONSIVE GATA TRANSCRIPTION FACTOR1/GNC-like (''CGA1/GNL'') transcription factor was originally identified in Arabidopsis (At4g26150) due to rapidly increased expression following cytokinin application and similarity to a paralogous gene produced through genome duplication<ref name=" | + | The CYTOKININ-RESPONSIVE GATA TRANSCRIPTION FACTOR1/GNC-like (''CGA1/GNL'') transcription factor was originally identified in Arabidopsis (At4g26150) due to rapidly increased expression following cytokinin application and similarity to a paralogous gene produced through genome duplication<ref name="p1" /><ref name="p2" /><ref name="p3" /><ref name="p4" />. |
==Annotated Information== | ==Annotated Information== | ||
| Line 5: | Line 5: | ||
===Function=== | ===Function=== | ||
| − | Transgenic Arabidopsis plants with altered expression of ''CGA1'' have been shown to exhibit differences in germination, chlorophyll content, chloroplast number, leaf size, flowering time, and senescence<ref name=" | + | Transgenic Arabidopsis plants with altered expression of ''CGA1'' have been shown to exhibit differences in germination, chlorophyll content, chloroplast number, leaf size, flowering time, and senescence<ref name="p4" /><ref name="p5" /><ref name="p6" />. This includes recent reports showing that ectopic overexpression promotes chloroplast biogenesis in cells where they are not typically found. These data indicate that GNC and CGA1 function as key transcriptional regulators of chloroplast biogenesis in Arabidopsis. |
| − | A new research demonstrates that the conserved GATA transcription factor Cga1 (Os02g12790) regulates chloroplast development and plant architecture in rice (Oryza sativa)<ref name=" | + | A new research demonstrates that the conserved GATA transcription factor Cga1 (Os02g12790) regulates chloroplast development and plant architecture in rice (Oryza sativa)<ref name="p7" />. |
===Mutations=== | ===Mutations=== | ||
| Line 53: | Line 53: | ||
<references> | <references> | ||
| − | <ref name=" | + | <ref name="p1"> Reyes JC, Muro-Pastor MI, Florencio FJ (2004) The GATA family of transcription factors in Arabidopsis and rice. Plant Physiol 134: 1718–1732.</ref> |
| − | <ref name=" | + | <ref name="p2"> Kiba T, Naitou T, Koizumi N, Yamashino T, Sakakibara H, Mizuno T (2005) Combinatorial microarray analysis revealing Arabidopsis genes implicated in cytokinin responses through the His→Asp phosphorelay circuitry. Plant Cell Physiol 46: 339–355. </ref> |
| − | <ref name=" | + | <ref name="p3"> Naito T, Kiba T, Koizumi N, Yamashino T, Mizuno T (2007) Characterization of a unique GATA family gene that responds to both light and cytokinin in |
Arabidopsis thaliana. Biosci Biotechnol Biochem 71: 1557–1560. </ref> | Arabidopsis thaliana. Biosci Biotechnol Biochem 71: 1557–1560. </ref> | ||
| − | <ref name=" | + | <ref name="p4"> Mara CD, Irish VF (2008) Two GATA transcription factors are downstream effectors of floral homeotic gene action in Arabidopsis. Plant Physiol 147: 707–718. </ref> |
| − | <ref name=" | + | <ref name="p5"> Richter R, Behringer C, Müller IK, Schwechheimer C (2010) The GATAtype transcription factors GNC and GNL/CGA1 repress gibberellin signaling downstream from DELLA proteins and PHYTOCHROMEINTERACTING FACTORS. Genes Dev 24: 2093–2104. </ref> |
| − | <ref name=" | + | <ref name="p6"> Hudson D, Guevara D, Yaish MW, Hannam C, Long N, Clarke JD, Bi YM, Rothstein SJ (2011) GNC and CGA1 modulate chlorophyll biosynthesis and glutamate synthase (GLU1/Fd-GOGAT) expression in Arabidopsis. PLoS ONE 6: e26765. </ref> |
| − | <ref name=" | + | <ref name="p7"> Hudson, D., Guevara, D. R., Hand, A. J., Xu, Z., Hao, L., Chen, X., Zhu, T., Bi, Y. M., and Rothstein, S. J. (2013). Rice cytokinin GATA transcription Factor1 regulates chloroplast development and plant architecture. Plant physiology 162, 132-144. </ref> |
</references> | </references> | ||
Revision as of 16:41, 13 May 2014
The CYTOKININ-RESPONSIVE GATA TRANSCRIPTION FACTOR1/GNC-like (CGA1/GNL) transcription factor was originally identified in Arabidopsis (At4g26150) due to rapidly increased expression following cytokinin application and similarity to a paralogous gene produced through genome duplication[1][2][3][4].
Contents
Annotated Information
Function
Transgenic Arabidopsis plants with altered expression of CGA1 have been shown to exhibit differences in germination, chlorophyll content, chloroplast number, leaf size, flowering time, and senescence[4][5][6]. This includes recent reports showing that ectopic overexpression promotes chloroplast biogenesis in cells where they are not typically found. These data indicate that GNC and CGA1 function as key transcriptional regulators of chloroplast biogenesis in Arabidopsis.
A new research demonstrates that the conserved GATA transcription factor Cga1 (Os02g12790) regulates chloroplast development and plant architecture in rice (Oryza sativa)[7].
Mutations
Transgenic rice with altered expression of Cga1 exhibits differences in chlorophyll, chloroplast number, and starch content, which has also been reported in Arabidopsis. However, we also observed a dosage-dependent influence on phenotype, with strong overexpression causing a semidwarf phenotype, similar to the GA mutant Green Revolution varieties (Figure1).
Figure 1. Transgenic modification to Cga1 expression alters chlorophyll content and plant architectureCite error: Invalid <ref> tag;
name cannot be a simple integer. Use a descriptive title.
Novel evidence was presented that altering Cga1 expression in rice significantly influences tillering, biomass, and yield (Figure1, 2).
Figure 2. Cga1 expression influences starch content and grain productionCite error: Invalid <ref> tag;
name cannot be a simple integer. Use a descriptive title.
Changes are demonstrated in the expression of important nucleus-encoded, chloroplast-localized genes involved in chlorophyll binding, photosynthesis, and amino acid and starch biosynthesis in the Cga1 transgenics.
Altering expression of the rice homolog to the FILAMENTOUS TEMPERATURE SENSITIVE-Z (FtsZ) gene involved in chloroplast division provides a potential mechanism for controlling chloroplast number.
Growing the transgenic lines under different N conditions indicates that Cga1 is able to maintain chloroplast development under reduced N conditions, leading to an increased harvest index despite reduced plant size.
Expression
Rice tissues and patterns of expression for Cga1 were analyzed and established in wild-type Kaybonnet rice. Cga1 exhibited the strongest expression in green leaf tissue, with little and no expression in roots and floral organs, respectively (Figure3A).
The expression of Cga1 following a number of treatments was also analyzed. Differences in Cga1 expression throughout the course of the day was also observed. (Figure3B). Light was found to significantly increase Cga1 expression, whereas periods of darkness reduced expression (Figure3, B and C). Cga1 was highly upregulated (approximately 5-fold) by the synthetic cytokinin benzyladenine (BA; figure3C). Nitrate (NO3 -) also significantly increased Cga1 expression, although to a lesser extent than BA (Figure3C).
Figure 3. Expression of rice Cga1(Os02g12790)Cite error: Invalid <ref> tag;
name cannot be a simple integer. Use a descriptive title.
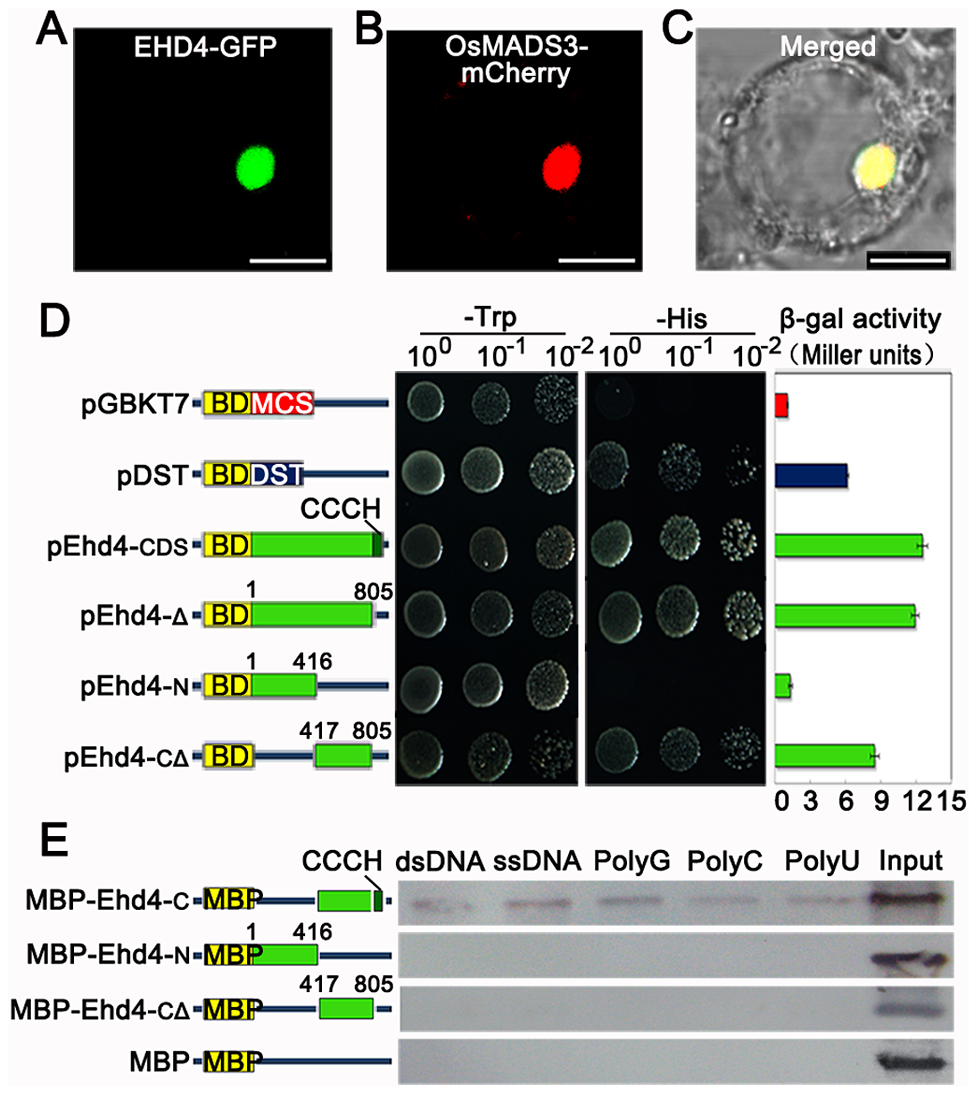
Subcellular Localization
Nucleus
Labs working on this gene
Department of Molecular and Cellular Biology, University of Guelph, Guelph, Ontario, Canada
Syngenta Biotechnology, Inc., Research Triangle Park,North Carolina
References
- ↑ Reyes JC, Muro-Pastor MI, Florencio FJ (2004) The GATA family of transcription factors in Arabidopsis and rice. Plant Physiol 134: 1718–1732.
- ↑ Kiba T, Naitou T, Koizumi N, Yamashino T, Sakakibara H, Mizuno T (2005) Combinatorial microarray analysis revealing Arabidopsis genes implicated in cytokinin responses through the His→Asp phosphorelay circuitry. Plant Cell Physiol 46: 339–355.
- ↑ Naito T, Kiba T, Koizumi N, Yamashino T, Mizuno T (2007) Characterization of a unique GATA family gene that responds to both light and cytokinin in Arabidopsis thaliana. Biosci Biotechnol Biochem 71: 1557–1560.
- ↑ 4.0 4.1 Mara CD, Irish VF (2008) Two GATA transcription factors are downstream effectors of floral homeotic gene action in Arabidopsis. Plant Physiol 147: 707–718.
- ↑ Richter R, Behringer C, Müller IK, Schwechheimer C (2010) The GATAtype transcription factors GNC and GNL/CGA1 repress gibberellin signaling downstream from DELLA proteins and PHYTOCHROMEINTERACTING FACTORS. Genes Dev 24: 2093–2104.
- ↑ Hudson D, Guevara D, Yaish MW, Hannam C, Long N, Clarke JD, Bi YM, Rothstein SJ (2011) GNC and CGA1 modulate chlorophyll biosynthesis and glutamate synthase (GLU1/Fd-GOGAT) expression in Arabidopsis. PLoS ONE 6: e26765.
- ↑ Hudson, D., Guevara, D. R., Hand, A. J., Xu, Z., Hao, L., Chen, X., Zhu, T., Bi, Y. M., and Rothstein, S. J. (2013). Rice cytokinin GATA transcription Factor1 regulates chloroplast development and plant architecture. Plant physiology 162, 132-144.
Structured Information
| Gene Name |
Os02g0220400 |
|---|---|
| Description |
Similar to GATA transcription factor 16 |
| Version |
NM_001052851.1 GI:115445072 GeneID:4328751 |
| Length |
1681 bp |
| Definition |
Oryza sativa Japonica Group Os02g0220400, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 2:6734753..6736433 |
| Sequence Coding Region |
6734994..6735464,6735659..6736039,6736187..6736396 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008395:6734753..6736433 source=RiceChromosome02 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008395:6734753..6736433 source=RiceChromosome02 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>atgtctactatctacatgagccagctacctgctactctccctctaatggagggggatcaggatcaggggctctacccagccttccatagagcaaaggaccctcctatcttgttccctttcatgatcgacagcgccgtcgagcaccaagggcaaatctatggagatcagggcttgaggaggcagcaggttttgggtgaatccaatcaacagttcaatgatcacatgatgatgggcggatcagatgtcttcctcacaccgtctccgttccgaccaaccatccaaagcatcggcagcgacatgatccagcgatcatcttatgatccatacgatatcgagagtaacaacaagcagcatgccaatggatcaaccagcaagtggatgtcgacgccgccaatgaagatgaggatcataaggaagggggcggcaaccgatcctgagggcggggcggtgagaaagccaaggagaagagcacaagcgcaccaggatgagagccagcaacaactgcagcaagctttgggtgtcgttagagtgtgctcggactgcaacaccaccaagacccccttgtggagaagtggtccttgtggccccaagtccctttgcaacgcgtgtggcatcaggcaaaggaaggcgcggcgggcgatggccgctgctgccaacggcggagcggcggtggcgccggcaaagagcgtggccgcggcgccggtgaacaataagccggcggcgaagaaggagaagagggcggcggacgtcgaccggtcgctgccgttcaagaaacggtgcaagatggtcgatcacgttgctgctgccgtcgctgccaccaagcccacggctgctggagaagtagtggccgccgctccgaaggaccaagatcacgtcatcgtcgtcggtggcgagaacgccgccgccacctccatgccggcacagaacccgatatccaaggcggcggcgaccgccgctgccgccgccgcctctccggcgttcttccacggcctccctcgcgacgagatcaccgacgccgccatgctgctcatgaccctatcctgtggcctcgtccacagctag</cdnaseq> |
| Protein Sequence |
<aaseq>MSTIYMSQLPATLPLMEGDQDQGLYPAFHRAKDPPILFPFMIDS AVEHQGQIYGDQGLRRQQVLGESNQQFNDHMMMGGSDVFLTPSPFRPTIQSIGSDMIQ RSSYDPYDIESNNKQHANGSTSKWMSTPPMKMRIIRKGAATDPEGGAVRKPRRRAQAH QDESQQQLQQALGVVRVCSDCNTTKTPLWRSGPCGPKSLCNACGIRQRKARRAMAAAA NGGAAVAPAKSVAAAPVNNKPAAKKEKRAADVDRSLPFKKRCKMVDHVAAAVAATKPT AAGEVVAAAPKDQDHVIVVGGENAAATSMPAQNPISKAAATAAAAAASPAFFHGLPRD EITDAAMLLMTLSCGLVHS</aaseq> |
| Gene Sequence |
<dnaseqindica>970..1440#395..775#38..247#cttctctcccatctctttcctcctcctcctctctgatatgtctactatctacatgagccagctacctgctactctccctctaatggagggggatcaggatcaggggctctacccagccttccatagagcaaaggaccctcctatcttgttccctttcatgatcgacagcgccgtcgagcaccaagggcaaatctatggagatcagggcttgaggaggcagcaggttttgggtgaatccaatcaacaggtgaggggatcgatcacatacatgtgtagaacaagctagctacttatacacatgtagatcagtgcttctttcttagtatatatgctgcttgccctaataattttagcttatactcctcttctttttctcttttttttgctcttgcagttcaatgatcacatgatgatgggcggatcagatgtcttcctcacaccgtctccgttccgaccaaccatccaaagcatcggcagcgacatgatccagcgatcatcttatgatccatacgatatcgagagtaacaacaagcagcatgccaatggatcaaccagcaagtggatgtcgacgccgccaatgaagatgaggatcataaggaagggggcggcaaccgatcctgagggcggggcggtgagaaagccaaggagaagagcacaagcgcaccaggatgagagccagcaacaactgcagcaagctttgggtgtcgttagagtgtgctcggactgcaacaccaccaagacccccttgtggagaagtggtccttgtggccccaaggtgagatttacttgctcttacacccctattaacaactgcccaaactcatgtgttgatctgtctgtctgggtgctatatgctaccttactatgtgcattttctctttgttttgttacaccagatcatcatgcatatgttaaagattgttgatttccttggtttaaattgtgtgttgtgctatgcatatggtgcagtccctttgcaacgcgtgtggcatcaggcaaaggaaggcgcggcgggcgatggccgctgctgccaacggcggagcggcggtggcgccggcaaagagcgtggccgcggcgccggtgaacaataagccggcggcgaagaaggagaagagggcggcggacgtcgaccggtcgctgccgttcaagaaacggtgcaagatggtcgatcacgttgctgctgccgtcgctgccaccaagcccacggctgctggagaagtagtggccgccgctccgaaggaccaagatcacgtcatcgtcgtcggtggcgagaacgccgccgccacctccatgccggcacagaacccgatatccaaggcggcggcgaccgccgctgccgccgccgcctctccggcgttcttccacggcctccctcgcgacgagatcaccgacgccgccatgctgctcatgaccctatcctgtggcctcgtccacagctagctagctagctgatcaaaactagctagctactagtaccgttaatttgatgagggcaacaaccagagtactatgtaccactactagcaatattttgtgtgtgccttgtgatcttttgttgttttgtgttgttgaggagatcactagatcaggatgaaggagagatagtgatcacatgtctaaggacgaaataaacgagaacaaactcgctagctagctactagccgggatcaggattatattt</dnaseqindica> |
| External Link(s) |