Difference between revisions of "Os03g0597200"
Yilutongxing (talk | contribs) (→Labs working on this gene) |
Yilutongxing (talk | contribs) (→Annotated Information) |
||
| Line 3: | Line 3: | ||
==Annotated Information== | ==Annotated Information== | ||
===Function=== | ===Function=== | ||
| − | YSA is required for chloroplast development in early seedling leaves, and disruption of its function causes a seedling stage-specific albino phenotype, but the plant recovers and develops normal green leaves from the four-leaf stage onward.The ysa mutant develops albino leaves before the three-leaf stage, but the mutant gradually turns green and recovers to normal green at the six-leaf stage. | + | YSA is required for chloroplast development in early seedling leaves, and disruption of its function causes a seedling stage-specific albino phenotype, but the plant recovers and develops normal green leaves from the four-leaf stage onward.The ysa mutant develops albino leaves before the three-leaf stage, but the mutant gradually turns green and recovers to normal green at the six-leaf stage.(Su N, Hu ML, Wu DX, et al.2012) |
Functional studies of PPR proteins in higher plants remain very sparse. Accumulating data point to an involvement in posttranscriptional processes in organelles. Additional evidence for a role of PPR proteins in regulating organelle gene expression has also come from positional cloning of several cytoplasmic male sterility (CMS) restorer genes from petunia (Petunia hybrida; Bentolila et al., 2002) and radish (Raphanus sativus; Brown et al., 2003; Desloire et al., 2003; Koizuka et al., 2003). Genetic and biochemical data, and structural modeling of PPR tracts based on established tetratricopeptide repeat proteins together suggest that PPR proteins typically bind directly to specific organellar RNA sequences through a surface created by the stacked helical repeating units. However, still very little is known about the functions, substrates, and regulatory mechanisms for the vast majority of PPR proteins.The ysa mutant develops albino leaves before the three-leaf stage, but the mutant gradually turns green and recovers to normal green at the six-leaf stage. Further investigation showed that the change in leaf color in ysa mutant is associated with changes in chlorophyll content and chloroplast development. | Functional studies of PPR proteins in higher plants remain very sparse. Accumulating data point to an involvement in posttranscriptional processes in organelles. Additional evidence for a role of PPR proteins in regulating organelle gene expression has also come from positional cloning of several cytoplasmic male sterility (CMS) restorer genes from petunia (Petunia hybrida; Bentolila et al., 2002) and radish (Raphanus sativus; Brown et al., 2003; Desloire et al., 2003; Koizuka et al., 2003). Genetic and biochemical data, and structural modeling of PPR tracts based on established tetratricopeptide repeat proteins together suggest that PPR proteins typically bind directly to specific organellar RNA sequences through a surface created by the stacked helical repeating units. However, still very little is known about the functions, substrates, and regulatory mechanisms for the vast majority of PPR proteins.The ysa mutant develops albino leaves before the three-leaf stage, but the mutant gradually turns green and recovers to normal green at the six-leaf stage. Further investigation showed that the change in leaf color in ysa mutant is associated with changes in chlorophyll content and chloroplast development. | ||
| Line 10: | Line 10: | ||
Tissue localization | Tissue localization | ||
| − | YSA is highly expressed in young leaves and stems, but not in the roots (Fig. 1, A–D). Quantitative real-time reverse transcription (RT)-PCR analysis revealed that the expression of YSA peaked in the fourth leaf (Fig. | + | YSA is highly expressed in young leaves and stems, but not in the roots (Fig. 1, A–D). Quantitative real-time reverse transcription (RT)-PCR analysis revealed that the expression of YSA peaked in the fourth leaf (Fig. 2E). Thus, the expression pattern of YSA is consistent with the seedling-stage-specific albino phenotype of ysa mutant and further supports the notion that YSA plays an important role in chloroplast development in the first few leaves of rice seedlings, but plays more minor roles in later stages. |
[[File:picture1.jpg]] | [[File:picture1.jpg]] | ||
Revision as of 08:11, 22 May 2014
Please input one-sentence summary here.
Contents
Annotated Information
Function
YSA is required for chloroplast development in early seedling leaves, and disruption of its function causes a seedling stage-specific albino phenotype, but the plant recovers and develops normal green leaves from the four-leaf stage onward.The ysa mutant develops albino leaves before the three-leaf stage, but the mutant gradually turns green and recovers to normal green at the six-leaf stage.(Su N, Hu ML, Wu DX, et al.2012) Functional studies of PPR proteins in higher plants remain very sparse. Accumulating data point to an involvement in posttranscriptional processes in organelles. Additional evidence for a role of PPR proteins in regulating organelle gene expression has also come from positional cloning of several cytoplasmic male sterility (CMS) restorer genes from petunia (Petunia hybrida; Bentolila et al., 2002) and radish (Raphanus sativus; Brown et al., 2003; Desloire et al., 2003; Koizuka et al., 2003). Genetic and biochemical data, and structural modeling of PPR tracts based on established tetratricopeptide repeat proteins together suggest that PPR proteins typically bind directly to specific organellar RNA sequences through a surface created by the stacked helical repeating units. However, still very little is known about the functions, substrates, and regulatory mechanisms for the vast majority of PPR proteins.The ysa mutant develops albino leaves before the three-leaf stage, but the mutant gradually turns green and recovers to normal green at the six-leaf stage. Further investigation showed that the change in leaf color in ysa mutant is associated with changes in chlorophyll content and chloroplast development.
Expression
Tissue localization YSA is highly expressed in young leaves and stems, but not in the roots (Fig. 1, A–D). Quantitative real-time reverse transcription (RT)-PCR analysis revealed that the expression of YSA peaked in the fourth leaf (Fig. 2E). Thus, the expression pattern of YSA is consistent with the seedling-stage-specific albino phenotype of ysa mutant and further supports the notion that YSA plays an important role in chloroplast development in the first few leaves of rice seedlings, but plays more minor roles in later stages.
Figure 1. Phenotypic analysis of the ysa mutant plants. A to C, Phenotypes of Pei'ai64S (left) and ysa mutant (right) seedlings at 1 (A), 2 (B), and 3 (C) weeks after sowing. D, The pigment contents in leaves of 1-week-old ysa mutants are much lower than that in Pei'ai64S. E, The pigment contents in leaves of 6-week-old ysa mutants are similar to that of Pei'ai64S plants. Chla, Chlorophyll a; Chlb, chlorophyll b; Chl, total chlorophyll; Car, carotenoid. Bars represent sds of three measurements. Student’s t test was performed on the raw data; asterisk indicates statistical significance at P < 0.01.
Subcellular Localization of YSA Protein
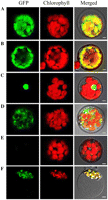
The YSA protein is predicted to localize to chloroplasts by ChloroP (Emanuelsson et al., 1999) and TargetP (Emanuelsson et al., 2000). To investigate the actual cellular localization of YSA, we constructed the green fluorescent signals of YSA-GFP fusion proteins colocalized with the autofluorescent signals of chlorophylls in the chloroplasts, consistent with the results obtained for GFP fused to the transit peptide of the small subunit of Arabidopsis ribulose bisphosphate carboxylase (Fig. 2, A and B). When GFP fused to the nuclear localization signal of the fibrillarin protein, GFP signals located specifically in the nucleus of Arabidopsis protoplasts (Fig. 2C). In addition, the protoplasts transformed with the empty GFP vector without a specific targeting sequence had green fluorescent signals in both the cytoplasm and the nucleus (Fig.2, D and E). To further confirm the subcellular localization of YSA protein, we transformed the plasmid containing the YSA-GFP fusion constructs into rice protoplasts. Confocal microscopy observations revealed that GFP-YSA was exclusively detected in the chloroplasts (Fig. 2F). These findings suggest that YSA protein is localized to the chloroplast.
Evolution
pentatricopeptide (PPR) repeat-containing protein; similar to pentatricopeptide (PPR) repeat-containing protein [Arabidopsis thaliana] (TAIR:AT5G65820.1); similar to OJ991113_30.18 [Oryza sativa (japonica cultivar-group)] (GB:CAE02059.2); similar to Os08g0525500 [Oryza sativa (japonica cultivar-group)] (GB:NP_001062291.1); contains InterPro domain Pentatricopeptide repeat; (InterPro:IPR002885); contains InterPro domain Protein prenyltransferase; (InterPro:IPR008940); contains InterPro domain Tetratricopeptide-like helical; (InterPro:IPR011990)
Homology with Arabidopsis Similar to At3g53700: MEE40 (maternal effect embryo arrest 40) (HF=7e-1) You can also add sub-section(s) at will.
Labs working on this gene
Please input related labs here. Institute of Crop Science, National Key Facility for Crop Gene Resources and Genetic Improvement, Chinese Academy of Agricultural Sciences, Beijing 10081, China
References
Please input cited references here.
Structured Information
| Gene Name |
Os03g0597200 |
|---|---|
| Description |
Protein prenyltransferase domain containing protein |
| Version |
NM_001057140.1 GI:115454008 GeneID:4333379 |
| Length |
5627 bp |
| Definition |
Oryza sativa Japonica Group Os03g0597200, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 3:22993430..22999056 |
| Sequence Coding Region |
22996530..22998758 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008396:22993430..22999056 source=RiceChromosome03 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008396:22993430..22999056 source=RiceChromosome03 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>atgccccgcgtttgcgccgcccctcgggcgccgccgccgccgtgcccgtgccatgtcggagtagggccgcttcggccgaggtggcgcgcctcccggcacggccctctccgggcggctggccaggagcagctcctcaccgccctgcgcgagcagccggaccccgacgcggcgctccggatgctcaacgcggcgctggcgcgggacgacttcgcgcccggccccgaggtctacgaggagatcatccgcaagctcggcgcggtcggggccctcgacctcatgaaggtgctcgtcgcggagatgcggcgggaggggcaccaggtgaaattgggcgtagtccactccttcttggacagctacgaggggcagcagctgttcgacgatgccgtcgacctgattctgaatcaactccaaccattgtttggcattcaggcagacaccgtggtgtacaatcaccttctcaatgttcttgtggaggggagcaaaatgaaactccttgaatcagtgtactcggagatgggtgctaggggaatcaagcctgatgttgtcacattcaacacactgatgaaggcgttgtgccgagcacatcaggtcaggactgcagttctcatgctcgaggaaatgtctagcagaggcgtggcgcctgacgagacgacgtttaccaccctgatgcaaggatttgtcgaggaggggagcatcgaggctgcactgagggtcaaagccaggatgttggaaatggggtgctcggcgacgaaggtgacggttaatgttttgattaatgggtactgcaagctagggagggtggaggatgctcttgggtatatacagcaggagattgccgatgggtttgagcctgaccagatcacatataacacttttgttaatggtctctgccaaaatgatcatgtcggccatgccctcaaagtcatggatgtgatggttcaggagggccatgatcctgatgttttcacctacaatatcgttgtgaattgcctttgtaaaaatggacagcttgaagaggcaaaaggaattctgaatcagatggtggatcggggttgtttgcctgacattaccacattcaacactctcattgctgccttatgcacggggaatcgacttgaggaagccttggaccttgcacgtcaggttacagtgaagggagtctctccagatgtttatactttcaatattctgattaacgcgctctgcaaagtaggcgatcctcatcttgcacttcgattgtttgaagagatgaagaacagtggatgcaccccggatgaagtaacatacaatactttgattgacaatctttgctcacttgggaagcttggtaaagcattggatttgttaaaagatatggagtccactggttgtcctcgaagtacaattacatataacactataattgacgggttatgcaagaaaatgagaatagaagaagcagaagaagtttttgatcaaatggatctgcaaggcatttcgaggaatgcaatcacattcaatactctcatcgatggtttgtgcaaggacaaaaagattgatgatgcttttgagcttattaatcaaatgataagtgaagggttgcaacctaacaatatcacttacaattctattctaactcattattgcaagcaaggtgacataaaaaaggctgcggatattttagaaactatgactgcaaatggatttgaagtggatgttgttacgtacggtactctgattaacggtctatgcaaggctggtaggacacaggttgctttgaaggtactcagaggtatgcggataaaagggatgaggcctactccaaaagcttacaatcctgtgctccagtctctcttcagacggaataatatcagagatgccctgagtcttttcagggagatggcagaggttggtgagcctcctgatgctttgacatataagattgtttttcgtgggctctgtcgtggtggagggcctattaaagaagcttttgatttcatgttggagatggttgataaggggttcataccagagttctcatccttccgtatgctagctgaaggtctattaaacctgggtatggatgattacttcattagagccattgaaataatcatggaaaaggtcgacctcagagagtctgatgtttctgcaataaggggatatctcaagatccgcaaattttatgatgcattagcaacctttggccgtttcctggagatcaacaaccctcaatggagttaccgatga</cdnaseq> |
| Protein Sequence |
<aaseq>MPRVCAAPRAPPPPCPCHVGVGPLRPRWRASRHGPLRAAGQEQL LTALREQPDPDAALRMLNAALARDDFAPGPEVYEEIIRKLGAVGALDLMKVLVAEMRR EGHQVKLGVVHSFLDSYEGQQLFDDAVDLILNQLQPLFGIQADTVVYNHLLNVLVEGS KMKLLESVYSEMGARGIKPDVVTFNTLMKALCRAHQVRTAVLMLEEMSSRGVAPDETT FTTLMQGFVEEGSIEAALRVKARMLEMGCSATKVTVNVLINGYCKLGRVEDALGYIQQ EIADGFEPDQITYNTFVNGLCQNDHVGHALKVMDVMVQEGHDPDVFTYNIVVNCLCKN GQLEEAKGILNQMVDRGCLPDITTFNTLIAALCTGNRLEEALDLARQVTVKGVSPDVY TFNILINALCKVGDPHLALRLFEEMKNSGCTPDEVTYNTLIDNLCSLGKLGKALDLLK DMESTGCPRSTITYNTIIDGLCKKMRIEEAEEVFDQMDLQGISRNAITFNTLIDGLCK DKKIDDAFELINQMISEGLQPNNITYNSILTHYCKQGDIKKAADILETMTANGFEVDV VTYGTLINGLCKAGRTQVALKVLRGMRIKGMRPTPKAYNPVLQSLFRRNNIRDALSLF REMAEVGEPPDALTYKIVFRGLCRGGGPIKEAFDFMLEMVDKGFIPEFSSFRMLAEGL LNLGMDDYFIRAIEIIMEKVDLRESDVSAIRGYLKIRKFYDALATFGRFLEINNPQWS YR</aaseq> |
| Gene Sequence |
<dnaseqindica>299..2527#ctcctgttcccctctgcctgccttcacggagaacacgccgccgcacgcccgcaaagttgtcgctccgccgccgggtcctgcggccacttcctccctctccctgtgcatgcgctctcttccccacctgtactttactttagctgctcctctgcccagttgcccacgacctgacgacccggacatggcgcaggctgaggcggggacgacgacatcgccggcgggttgacgcagaaaggagcgaccacccgagggctccgctggattttcaggtagctgagctgagctgaactgaaccccaatgccccgcgtttgcgccgcccctcgggcgccgccgccgccgtgcccgtgccatgtcggagtagggccgcttcggccgaggtggcgcgcctcccggcacggccctctccgggcggctggccaggagcagctcctcaccgccctgcgcgagcagccggaccccgacgcggcgctccggatgctcaacgcggcgctggcgcgggacgacttcgcgcccggccccgaggtctacgaggagatcatccgcaagctcggcgcggtcggggccctcgacctcatgaaggtgctcgtcgcggagatgcggcgggaggggcaccaggtgaaattgggcgtagtccactccttcttggacagctacgaggggcagcagctgttcgacgatgccgtcgacctgattctgaatcaactccaaccattgtttggcattcaggcagacaccgtggtgtacaatcaccttctcaatgttcttgtggaggggagcaaaatgaaactccttgaatcagtgtactcggagatgggtgctaggggaatcaagcctgatgttgtcacattcaacacactgatgaaggcgttgtgccgagcacatcaggtcaggactgcagttctcatgctcgaggaaatgtctagcagaggcgtggcgcctgacgagacgacgtttaccaccctgatgcaaggatttgtcgaggaggggagcatcgaggctgcactgagggtcaaagccaggatgttggaaatggggtgctcggcgacgaaggtgacggttaatgttttgattaatgggtactgcaagctagggagggtggaggatgctcttgggtatatacagcaggagattgccgatgggtttgagcctgaccagatcacatataacacttttgttaatggtctctgccaaaatgatcatgtcggccatgccctcaaagtcatggatgtgatggttcaggagggccatgatcctgatgttttcacctacaatatcgttgtgaattgcctttgtaaaaatggacagcttgaagaggcaaaaggaattctgaatcagatggtggatcggggttgtttgcctgacattaccacattcaacactctcattgctgccttatgcacggggaatcgacttgaggaagccttggaccttgcacgtcaggttacagtgaagggagtctctccagatgtttatactttcaatattctgattaacgcgctctgcaaagtaggcgatcctcatcttgcacttcgattgtttgaagagatgaagaacagtggatgcaccccggatgaagtaacatacaatactttgattgacaatctttgctcacttgggaagcttggtaaagcattggatttgttaaaagatatggagtccactggttgtcctcgaagtacaattacatataacactataattgacgggttatgcaagaaaatgagaatagaagaagcagaagaagtttttgatcaaatggatctgcaaggcatttcgaggaatgcaatcacattcaatactctcatcgatggtttgtgcaaggacaaaaagattgatgatgcttttgagcttattaatcaaatgataagtgaagggttgcaacctaacaatatcacttacaattctattctaactcattattgcaagcaaggtgacataaaaaaggctgcggatattttagaaactatgactgcaaatggatttgaagtggatgttgttacgtacggtactctgattaacggtctatgcaaggctggtaggacacaggttgctttgaaggtactcagaggtatgcggataaaagggatgaggcctactccaaaagcttacaatcctgtgctccagtctctcttcagacggaataatatcagagatgccctgagtcttttcagggagatggcagaggttggtgagcctcctgatgctttgacatataagattgtttttcgtgggctctgtcgtggtggagggcctattaaagaagcttttgatttcatgttggagatggttgataaggggttcataccagagttctcatccttccgtatgctagctgaaggtctattaaacctgggtatggatgattacttcattagagccattgaaataatcatggaaaaggtcgacctcagagagtctgatgtttctgcaataaggggatatctcaagatccgcaaattttatgatgcattagcaacctttggccgtttcctggagatcaacaaccctcaatggagttaccgatgaagcagaatacataactgggacaaattacttgaatagtattagggaaatctcaaagagtggatggaatttttgctggttgcttaggggaatgaaagtcttaaattgattataataggtgattgtgttcattcctcggtagggatgaagtcagagcatgaagaagctcatcttggtgcagaaacttagcttattggaacagaatctaggtgctaggtgctgtctgctgccagattagtacctagcgccttaacgaacagtggctgcagatcccatccgcttattttgatcaaatctgattcattttctattccctaataaaagcctgattcatcttcattgcatatggtcgaacctaaggctatcatgtacagttagatcccaacccttcgttctatgagatgttgtcccatagaagaaatatcttctgagtatatcctagtactctaaaatgttcactaatatgttcaacttaattagttttgtaacctcctaaaatagttctattagttttgtaaccttctaaaatatgaattagttttgatctggctgatcttcctttggttaggtactacaaattcttaattcagacacatatttggttttctgaaaatttcatctatatttggtcaggctggcatttgaagttcttattttagtcatatactttagttttttacaattttcatttgttaaagatgataatttatttgttagcacagagcatgtttagaaatctgaaatattaaaacatgcatgttctcatgaaaataaatgttagttttgtttaaattccaatccacatattttttaatcaatgtcagaaattaccatgcttcacttattgacctagtatatgtatagtatttgatggatcatgttgatttggatttgctctactaacttgttcctattccaacaaagatttatatgcatcttgtgttctaaaaatgctacatgtgtcaagttgaaggaaaattctagctatgtggtgtcctaattttggtagatggtacctagtaatagataacaattctgttttatagtgatgagtaaatttgactaaatcgagtctagaaagtgatataattttctggagaagtctttcttggtgatttgggaaagggccattacctatactgatatgaaatctggaactagaaagatctgaacatcaatgttctaaagttttttgtctgaatttcttgtgcagaatatgaaggaaggtggatctggaataggtatttacatgtcctgttcagattcctgcaatctgataaactactgcaatccaataaactcttgatctattgttttctggatttttttttgggtgtagagtattaggaaggtatttgctttgttacaggggttggttggatgttcagaaaaccaaaatctgaatacactaacacattagccagtgttttattttagtttatgttattctgaccacaacaccagcagcttcaattggtagtagaacacactctgatgcaattggtaggtgtacaacagataatttgccgtgaatgctacttaattcaaccattttttttccatacagtgctatacaatggcacacaaacatccttcagttggactcaaatttgtttcagctttgctattttacagtcacctgcagttttttctaatccatcagtcgattttgttggcagtagattctgcttcaagatctgtgcacgcagtggaggcacatttggttacaggctgcttcaccagacccgttaacccatgggccatggatgctcccctagtagcatatgttgttttttccaccaactgcctcttgtaaatcaagatgctcccctctccacatttgctgcagacttcctccccgtggatctgagacgaactccggcgaccggcggcgtctacgagccagcgttcaccaatttctcaggtatatgagtcgtcatctatgtgcatctgctactttttttttttggctgattggacgggttgagaaggaggtgtttggggggaagagagcagaataatcccattgcgaaacaagcgaggaaggcttgtcgatggtggcggcggtggccttgattatcgaagatgcatgcgtgagggaaactgtgtgccagggaggcaagcttggtggcacacatcgagtggtatatcttcaggttactgctagatcgaagatcctttggtgcttgatgtgtgctgttggtttggtagttttggtggtactgtggcaagtgaagacacgaaagaggagctagtagaggcttagagtctggagtcgtaatcgatgtgcaagagcatcaatgccatcagttcctctgggcagcacggctctgcacttttcatctgagaaacatatatattttgttgattctgattttgacgaattatccaaacatgcagtttttttgggttccgcgtgcaaagacgattttgatctcctggcagatttaagggtggtgattttgtttttgttttgcatgtagatgatgcattttgtaaatcattttaggttgccccttttttttttgtttttgcgagttcttactgaatttggataagttctggttgattggatcgacattctgtacatttgcaaggggattctatgggtttttgcttattgctaatcaaattattttgtgagttttggtttattggatggtgagaaagcaagattacatggattttgcttcattgctgattttgtcgaaatttatcaggcatgcggtttttggtttcttgaatacgaactacacggacagtactcctatattcttggtgattttgaagtgtatttgttattttggggaggtggaatggttaatattttgttcaaggggacacttctacagatgtatacattacttttgtcattttcagcataacaaagagatttggtagattcagaattcagatggcaactacacgtcgaatgatgtatgcgaagacatggggaattctgatgctgattccactgaagaaagctaaacataattttaatttatttaccaactcatttttctgtcaacacatttgctagatagttgctacccgataaacgacatcttgaaatctgagttatacttcagtctattttttttcc</dnaseqindica> |
| External Link(s) |