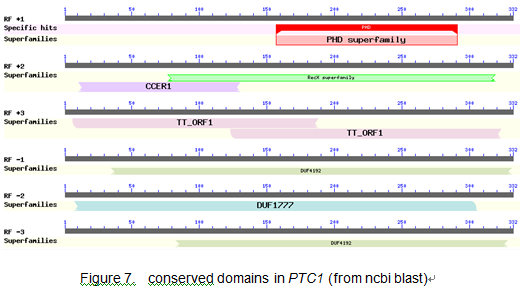
Difference between revisions of "Os09g0449000"
(→Mutation) |
(→Expression) |
||
| Line 41: | Line 41: | ||
Please input expression information | Please input expression information | ||
| − | According to the RiceGE (Gene Expression Atlas Data Sources), PTC1 expression is highest in the 10- to 15-cm inflorescence (stages 7–10), gradually decreasing in the 15- to 20-cm inflorescence (stages 10–12). Reverse transcription (RT)-PCR and quantitative RT-PCR (RT-qPCR) results indicated that PTC1was preferentially expressed in the anther from stage 8 to stage 9 (Fig. | + | According to the RiceGE (Gene Expression Atlas Data Sources), PTC1 expression is highest in the 10- to 15-cm inflorescence (stages 7–10), gradually decreasing in the 15- to 20-cm inflorescence (stages 10–12). Reverse transcription (RT)-PCR and quantitative RT-PCR (RT-qPCR) results indicated that PTC1was preferentially expressed in the anther from stage 8 to stage 9 (Fig.6, A and B). Strikingly, ptc1 displayed an increased level of the mutantPTC1transcript (Fig.6, A and B). This suggests either auto down-regulation of the PTC1 transcript or that the mutant transcript may be more stable and accumulates more than the wild type. In transgenic plants expressing the PTC1pro:GUS fusion (driven by the 3,043-bpPTC1upstream region), GUS staining was specifically detected in the anther from stages 8 to 9 and at a low level in stage 10 (Fig.6, C and D). No GUS expression was seen in the palea, lemma, pistil (Fig.6, C and D) or other organs. Maximal GUS staining, albeit at a very low level, was seen in the tapetal tissues, but GUS was also apparent within the microspores (Fig.6E), indicating that PTC1 plays a specific function in tapetal cells during anther development. |
| − | [[File: | + | [[File:yuan6.jpg]] |
===Microarray analysis=== | ===Microarray analysis=== | ||
Revision as of 14:03, 23 May 2014
Please input one-sentence summary here.
Contents
Annotated Information
Function
Please input function information here. PERSISTANT TAPETAL CELL1(PTC1), a key regulator in monocot rice (Oryza sativa), which controls programmed tapetal development and functional pollen formation. The evolutionary significance of PTC1was revealed by partial genetic complementation of the homologous mutation MALE STERILITY1(MS1) in the dicot Arabidopsis (Arabidopsis thaliana). PTC1 encodes a PHD-finger (for plant homeodomain) protein, which is expressed specifically in tapetal cells and microspores during anther development in stages 8 and 9, when the wild-type tapetal cells initiate a typical apoptosis-like cell death [1]. Even though ptc1 mutants show phenotypic similarity to ms1in a lack of tapetal DNA fragmentation, delayed tapetal degeneration, as well as abnormal pollen wall formation and aborted microspore development, the ptc1 mutant displays a previously unreported phenotype of uncontrolled tapetal proliferation and subsequent commencement of necrosis-like tapetal death. Moreover, the regulatory role of PTC1in anther development was revealed by comparison with MS1 and other rice anther developmental regulators. PTC1 may functions downstream of GAMYB[2], and in parallel with TDR in regulating programmed anther development and pollen formation[3].
Mutation
Please input mutation information ptc1 displayed complete male sterility and normal female development when tested by reciprocal cross analysis. All of the F1 progeny were fertile, with an approximate 3:1 ratio for phenotypic segregation in F2 plants. Compared with wild-type plants, ptc1 exhibited normal vegetative development (Fig.1,A). During the reproductive stage, ptc1 plants developed normal panicles and floral organs (Fig.1,B–D), but ptc1 anthers were smaller, white, and lacked viable pollen grains (Fig.1, D and E).
Histological analysis indicated that early events in anther and pollen development progressed normally until after meiosis and microspore release. The epidermis, endothecium, middle layer, tapetum, and pollen mother cells formed normally in ptc1 anthers at stage 6 before meiosis (Fig. 2, A and E). During meiosis (stage 8a), both the wild-type and ptc1 pollen mother cells became associated with the tapetal layer and the tapetum appeared vacuolated (Fig. 2, B and F); subsequently (during stage 8b), tetrads with four haploid microspores formed and the middle cell layer became very thin and degenerated in both wildtype and ptc1 cells (Fig. 2, C and G). From stage 9 (free microspore released from the tetrad) to early stage 10 (vacuolated microspore), the wild-type tapetum became condensed, less vacuolated, and stained strongly (Fig. 2, D and I). In contrast, ptc1 tapetal cells were more vacuolated and less stained (Fig. 2, H and M). From late stage 10 to stage 11 (mitosis I), the wild-type tapetum appeared to undergo degeneration, and the cytoplasmic constituents remained densely stained and clearly discernible (Fig.2, J and K); however, the ptc1 tapetum became abnormally enlarged, and cytosolic constituents seemed to be secreted into the anther locule (Fig. 2, N and O).Meanwhile, the wild-type microspores maintained a defined shape (Fig. 2, J and K), while the ptc1 microspores degenerated (Fig. 2, N–P).At stage 13, when the mature pollen grains were formed, the wild-type tapetal layer and middle layer were completely degraded, and round, densely stained pollen grains were observed (Fig. 2L). However, at this stage in the ptc1 anther, only cell debris of both tapetal cells and pollen grains remained (Fig. 2P).
Transmission electron microscopy was used to gain greater insight into the ptc1 tapetal and microspore abnormalities. From stage 9 onward, the wild-type tapetal cytoplasm became condensed, with swollen and less defined organelles, which showed a loss of membrane integrity and a diffuse cellular organization, indicative of apoptotic degeneration (Fig. 3, Aand B). Cytoplasmic condensation was reduced in the ptc1 tapetum, with membrane and organellar integrity maintained (Fig. 3C). The ptc1 tapetum had a high density of organelles, particularly mitochondria and endoplasmic reticulum, which appeared morphologically intact, giving the appearance of a highly active tissue without any signs of tapetal breakdown and apoptosis (Fig. 3, D, G, and H). Strikingly, extensive tapetal proliferation was also observed in ptc1, resulting in tapetal extrusions into the anther locule (Fig. 3,E–H). These extruded regions appeared to have a continuous outer membrane with the surrounding tapetal cells and contained an extremely high density of organelles, lipid bodies, and electron-dense materials (Fig. 3, G and H), implying that unregulated proliferation of the tapetum cellular contents was occurring.
Wild-type microspore developed a well-organized pollen wall, with extensive banding of electron-transparent material (Fig. 4, C and D), whereas ptc1 developed a thin and abnormal pollen exine, with fewer electron-transparent channels across the pollen wall (Fig. 4, G and H). The ptc1 mutant appeared to have a reduced bacular structure, with no spine structures as well as an abnormal nexine, which was uneven on the internal microspore surface (Fig. 4, G and H). This was supported by scanning electron microscopy (SEM), which indicated that the surface of wild-type pollen grains exhibited a regular pattern (Fig. 4, I and K) with spine distribution on the exine (Fig. 4, C and D), whereas ptc1 pollen showed a lack of the elabora
Tapetal DNA fragmentation was not detected at stage 7 in both the wild type and ptc1 (Fig.5, A and C). However, at early stage 9, when the microspore was released from the tetrad, DNA fragmentation signal could be seen in the wild type (Fig.5B).No TUNEL signal was detected in the ptc1 mutant (Fig. 5, C and D), implying that regulated DNA fragmentation does not occur in the ptc1 tapetal cells.
Expression
Please input expression information
According to the RiceGE (Gene Expression Atlas Data Sources), PTC1 expression is highest in the 10- to 15-cm inflorescence (stages 7–10), gradually decreasing in the 15- to 20-cm inflorescence (stages 10–12). Reverse transcription (RT)-PCR and quantitative RT-PCR (RT-qPCR) results indicated that PTC1was preferentially expressed in the anther from stage 8 to stage 9 (Fig.6, A and B). Strikingly, ptc1 displayed an increased level of the mutantPTC1transcript (Fig.6, A and B). This suggests either auto down-regulation of the PTC1 transcript or that the mutant transcript may be more stable and accumulates more than the wild type. In transgenic plants expressing the PTC1pro:GUS fusion (driven by the 3,043-bpPTC1upstream region), GUS staining was specifically detected in the anther from stages 8 to 9 and at a low level in stage 10 (Fig.6, C and D). No GUS expression was seen in the palea, lemma, pistil (Fig.6, C and D) or other organs. Maximal GUS staining, albeit at a very low level, was seen in the tapetal tissues, but GUS was also apparent within the microspores (Fig.6E), indicating that PTC1 plays a specific function in tapetal cells during anther development.
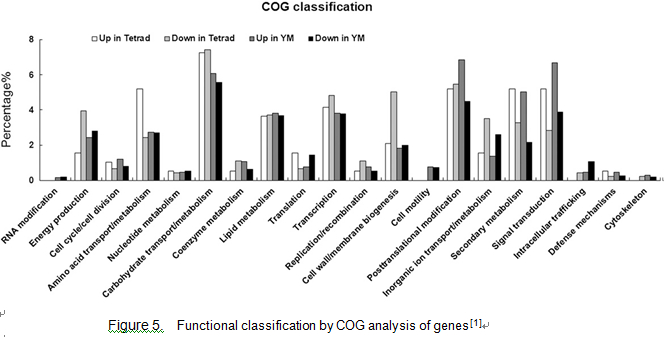
Microarray analysis
Please input Microarray analysis here. Microarray analysis revealed 2,943 genes with a 2-fold or greater expression change over the two developmental stages in ptc1 anthers compared with the wild type (570 down-regulated and 278 up-regulated at stage 8; 1,252 down-regulated and 843 up-regulated at stage 9. The 2,417 genes can be group into four COG (for Cluster of Orthologous Groups of proteins) categories: I, information storage and processing (144 genes); II, cellular processes and signaling (363 genes); III, metabolism (551 genes); and IV, poorly characterized genes (1,359 genes)
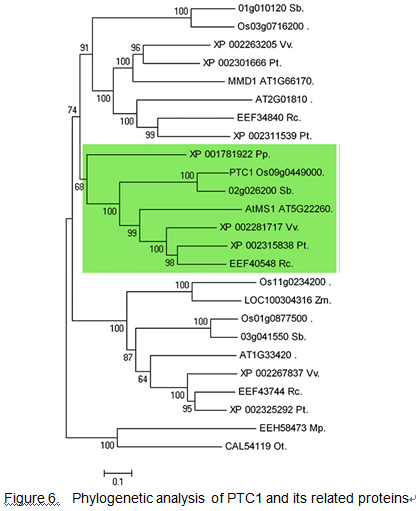
Evolution
Please input evolution information here.
You can also add sub-section(s) at will. To gain additional insights into the phylogenetic elationship between PTC1 and its close homologs, BLASTP searches were conducted with the full-length amino acid sequence of PTC1. An unrooted tree of PTC1 and its 25 homologs from moss to flowering plants was constructed. PTC1 was located in the same clade with Arabidopsis MS1.
Labs working on this gene
Please input related labs here.
School of Life Sciences and Biotechnology and State Key Laboratory of Hybrid Rice, Shanghai Jiao Tong University, Shanghai,China
Plant Sciences Division, School of Biosciences, University of Nottingham, Loughborough, Leicstershire, United Kingdom
References
Please input cited references here.
[1] Hui Li, Zheng Yuan, Gema Vizcay-Barrena. et al.PERSISTENT TAPETAL CELL1 Encodes a PHD-Finger Protein That Is Required for Tapetal Cell Death and Pollen Development in Rice. Plant Physiology 156: 615–630 (2011)
[2] Aya K, Ueguchi-Tanaka M, Kondo M . et al. Gibberellin modulates anther development in rice via the transcriptional regulation of GAMYB. Plant Cell 21: 1453–1472 (2009)
[3] Li N, Zhang DS, Liu HS. et al. The rice tapetum degeneration retardation gene is required for tapetum degradation and anther development. Plant Cell 18: 2999–3014 (2006)
Structured Information
| Gene Name |
Os09g0449000 |
|---|---|
| Description | |
| Version |
NM_001069854.2 GI:297609560 GeneID:4347213 |
| Length |
2215 bp |
| Definition |
Oryza sativa Japonica Group Os09g0449000, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 9:17439752..17441966 |
| Sequence Coding Region |
17439752..17440061,17440136..17440442,17440544..17441966 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008402:17439752..17441966 source=RiceChromosome09 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008402:17439752..17441966 source=RiceChromosome09 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>atggcgcctaagatggtgatcagcctggggagctcgcggcggcggaagcgcggcgagatgctgttccggttcgaggccttctgccagcccggctacccggcgaacttcgccggcgccggcggcttcagggacaacgtgaggacgctgctcggcttcgcgcacctggaggccggcgtccacggcgagaccaagtgctggtcgttccagctcgagctgcaccgccacccccccaccgtcgtgaggctcttcgtcgtcgaggaggaggtcgccgcctcgccgcaccgccagtgccacctctgccgccatattgggtgggggaggcatctgatatgcagcaagaggtatcacttcttgctgccgaggagggaatcggcggcggaagccgacggcctgtgcttcgcgatcaaccacggcggcggcggtggcgcggagaaagcgtcgtcgaaagggacgacgacgacggcctccagcagaggccacctgctacacggcgtcgtgcacctcaacggctacggccacctcgtcgccctccacggcctcgagggcggctccgacttcgtctccggccaccagatcatggacctctgggaccgcatttgctcagccttgcacgtaaggacggtgagcctggtggacacggcgaggaagggccacatggagctgaggctgctgcacggcgtcgcgtacggcgagacgtggttcgggcggtgggggtacaggtacggccggccgagctacggcgtcgcgctgccgtcgtaccggcagtcgctgcacgtgctcggctccatgccgctctgcgtgctggtgccgcacctgtcgtgcttcagccaggagctccccatggtggtcaccaagtaccaggccatcagcggccacaagctgctcagcctcggcgacctcctccgcttcatgctcgagctgcgcgcccgcctgccggccacctccgtcacggccatggactaccggggcatcatgtcggaggcctcgtgccggtggtcggcgaagcgcgtcgacatggcggcgcgcgccgtcgtggacgcgctccgccgcgccgagccggcggcgcggtgggtcacgcggcaggaggtgcgcgacgcggcgcgcgcctacatcggcgacacgggcctcctcgacttcgtgctcaagtccctcggcaaccacatcgtcggcaactacgtcgtgcgccgcaccatgaacccggtgaccaaggtgctcgagtactgcctcgaggacgtctccagcgtgctcccggcggtcgccgccggcggcggcgtgccggcgcagggcaagatgagggtgaggttccagctcacgcgtgcgcagctcatgagggacctggtgcacctgtaccggcacgtgctcaaggagcccagccaggcgctcaccggcggcgcgttcggcgcgatcccggtggcggtgcggatggtcctggacatcaagcacttcgtcaaagattaccacgaaggacaagccgcggcgagcagcaatggcggtggcggattcgggcatccccacatcaacctgtgctgcacgctgctcgtgagcaacgggagcccggagctagctccaccgtacgagacggtgaccctgccggcgcacgcgacggtgggcgagctgaagtgggaggcgcagagggtgttcagcgagatgtacctcggcctgaggagcttcgcggcggactccgtcgtcggggtcggcgccgaccaggagggcctcccggtgctcgggctggtcgacgtcggaagcgccgtcgtggtgcaagggagcgtgggcgagcagataaacggggaggaccacgagaggaaggaggaggcggcggcggcggccgtgtgcgaggggagcggcggcggcgagcgcgtcgtggactgcgcgtgcggcgcggtggacgacgacggcgagcgcatggcgtgctgcgacatctgcgaggcgtggcagcacacgcggtgcgccgggatcgcggacaccgaggacgcgccgcacgtcttcctctgcagccggtgcgacaacgacgtcgtgtcgttcccgtccttcaactgttag</cdnaseq> |
| Protein Sequence |
<aaseq>MAPKMVISLGSSRRRKRGEMLFRFEAFCQPGYPANFAGAGGFRD NVRTLLGFAHLEAGVHGETKCWSFQLELHRHPPTVVRLFVVEEEVAASPHRQCHLCRH IGWGRHLICSKRYHFLLPRRESAAEADGLCFAINHGGGGGAEKASSKGTTTTASSRGH LLHGVVHLNGYGHLVALHGLEGGSDFVSGHQIMDLWDRICSALHVRTVSLVDTARKGH MELRLLHGVAYGETWFGRWGYRYGRPSYGVALPSYRQSLHVLGSMPLCVLVPHLSCFS QELPMVVTKYQAISGHKLLSLGDLLRFMLELRARLPATSVTAMDYRGIMSEASCRWSA KRVDMAARAVVDALRRAEPAARWVTRQEVRDAARAYIGDTGLLDFVLKSLGNHIVGNY VVRRTMNPVTKVLEYCLEDVSSVLPAVAAGGGVPAQGKMRVRFQLTRAQLMRDLVHLY RHVLKEPSQALTGGAFGAIPVAVRMVLDIKHFVKDYHEGQAAASSNGGGGFGHPHINL CCTLLVSNGSPELAPPYETVTLPAHATVGELKWEAQRVFSEMYLGLRSFAADSVVGVG ADQEGLPVLGLVDVGSAVVVQGSVGEQINGEDHERKEEAAAAAVCEGSGGGERVVDCA CGAVDDDGERMACCDICEAWQHTRCAGIADTEDAPHVFLCSRCDNDVVSFPSFNC</aaseq> |
| Gene Sequence |
<dnaseqindica>1..310#385..691#793..2215#atggcgcctaagatggtgatcagcctggggagctcgcggcggcggaagcgcggcgagatgctgttccggttcgaggccttctgccagcccggctacccggcgaacttcgccggcgccggcggcttcagggacaacgtgaggacgctgctcggcttcgcgcacctggaggccggcgtccacggcgagaccaagtgctggtcgttccagctcgagctgcaccgccacccccccaccgtcgtgaggctcttcgtcgtcgaggaggaggtcgccgcctcgccgcaccgccagtgccacctctgccgccatattggtccgtcgaacaaactacaattaatcaatcaacctttacataggattgatccgatcgatgccatggtgttgtagggtgggggaggcatctgatatgcagcaagaggtatcacttcttgctgccgaggagggaatcggcggcggaagccgacggcctgtgcttcgcgatcaaccacggcggcggcggtggcgcggagaaagcgtcgtcgaaagggacgacgacgacggcctccagcagaggccacctgctacacggcgtcgtgcacctcaacggctacggccacctcgtcgccctccacggcctcgagggcggctccgacttcgtctccggccaccagatcatggacctctgggaccgcatttgctcagccttgcacgtaaggtagtagtagtatacatgtgcgtgtgcatgcatgcaagcaatgcaacgatgtcgggctgcgtgtgagaacatttgcttgggcatggtgtggtgtatgcaaggacggtgagcctggtggacacggcgaggaagggccacatggagctgaggctgctgcacggcgtcgcgtacggcgagacgtggttcgggcggtgggggtacaggtacggccggccgagctacggcgtcgcgctgccgtcgtaccggcagtcgctgcacgtgctcggctccatgccgctctgcgtgctggtgccgcacctgtcgtgcttcagccaggagctccccatggtggtcaccaagtaccaggccatcagcggccacaagctgctcagcctcggcgacctcctccgcttcatgctcgagctgcgcgcccgcctgccggccacctccgtcacggccatggactaccggggcatcatgtcggaggcctcgtgccggtggtcggcgaagcgcgtcgacatggcggcgcgcgccgtcgtggacgcgctccgccgcgccgagccggcggcgcggtgggtcacgcggcaggaggtgcgcgacgcggcgcgcgcctacatcggcgacacgggcctcctcgacttcgtgctcaagtccctcggcaaccacatcgtcggcaactacgtcgtgcgccgcaccatgaacccggtgaccaaggtgctcgagtactgcctcgaggacgtctccagcgtgctcccggcggtcgccgccggcggcggcgtgccggcgcagggcaagatgagggtgaggttccagctcacgcgtgcgcagctcatgagggacctggtgcacctgtaccggcacgtgctcaaggagcccagccaggcgctcaccggcggcgcgttcggcgcgatcccggtggcggtgcggatggtcctggacatcaagcacttcgtcaaagattaccacgaaggacaagccgcggcgagcagcaatggcggtggcggattcgggcatccccacatcaacctgtgctgcacgctgctcgtgagcaacgggagcccggagctagctccaccgtacgagacggtgaccctgccggcgcacgcgacggtgggcgagctgaagtgggaggcgcagagggtgttcagcgagatgtacctcggcctgaggagcttcgcggcggactccgtcgtcggggtcggcgccgaccaggagggcctcccggtgctcgggctggtcgacgtcggaagcgccgtcgtggtgcaagggagcgtgggcgagcagataaacggggaggaccacgagaggaaggaggaggcggcggcggcggccgtgtgcgaggggagcggcggcggcgagcgcgtcgtggactgcgcgtgcggcgcggtggacgacgacggcgagcgcatggcgtgctgcgacatctgcgaggcgtggcagcacacgcggtgcgccgggatcgcggacaccgaggacgcgccgcacgtcttcctctgcagccggtgcgacaacgacgtcgtgtcgttcccgtccttcaactgttag</dnaseqindica> |
| External Link(s) |