Difference between revisions of "Os03g0597200"
(→Expression) |
(→Labs working on this gene) |
||
| Line 34: | Line 34: | ||
==Labs working on this gene== | ==Labs working on this gene== | ||
| − | + | *National Key Facility for Crop Gene Resources and Genetic Improvement, Institute of Crop Science, Chinese Academy of Agricultural Sciences, Beijing 100081, People’s Republic of China (N.S., F.-Q.W., G.-L.F., Y.L., X.-L.C.,X.Z., X.-P.G., Z.-J.C., C.-L.L., H.W., J.-M.W.); | |
| − | + | *National Key Laboratory for Crop Genetics and Germplasm Enhancement, Nanjing Agricultural University, Nanjing 210095, People’s Republic of China (M.-L.H., L.J., J.-M.W.); | |
| + | *Institute of Industrial Crops, Jiangsu Academy of Agricultural Sciences, Nanjing 210014, People’s Republic of China (M.-L.H., C.-K.Q.); | ||
| + | *State Key Laboratory of Rice Biology, International Atomic Energy Agency Collaborating Center, Institute of Nuclear Agricultural Sciences, Zhejiang University, Hangzhou 310029,People’s Republic of China (D.-X.W., X.-L.S.) | ||
==References== | ==References== | ||
Revision as of 13:27, 31 May 2014
Please input one-sentence summary here.
Contents
Annotated Information
The predicted ORF ofYSAencodes a 742-amino acid polypeptide with a calculated molecular mass of 82.5kD. Database searches using Pfam revealed that YSAencodes a member of the P subfamily of PPR-containing proteins. The gene product contains a tandem repeat of 15 PPR motifs with varying degrees of conservation. The second to 15th repeats are contiguous, and the sequence is not well conserved (Fig. 4B). The PPR motif is a degenerate 35-amino acid repeat often arranged in tandem arrays of two to 27 repeats per polypeptide. Thus YSA is a new member of the superfamily of PPR proteins.
Function
YSA is required for chloroplast development in early seedling leaves, and disruption of its function causes a seedling stage-specific albino phenotype, but the plant recovers and develops normal green leaves from the four-leaf stage onward.The ysa mutant develops albino leaves before the three-leaf stage, but the mutant gradually turns green and recovers to normal green at the six-leaf stage.(Su N, Hu ML, Wu DX, et al.2012) Functional studies of PPR proteins in higher plants remain very sparse. Accumulating data point to an involvement in posttranscriptional processes in organelles. Additional evidence for a role of PPR proteins in regulating organelle gene expression has also come from positional cloning of several cytoplasmic male sterility (CMS) restorer genes from petunia (Petunia hybrida; Bentolila et al., 2002) and radish (Raphanus sativus; Brown et al., 2003; Desloire et al., 2003; Koizuka et al., 2003). Genetic and biochemical data, and structural modeling of PPR tracts based on established tetratricopeptide repeat proteins together suggest that PPR proteins typically bind directly to specific organellar RNA sequences through a surface created by the stacked helical repeating units. However, still very little is known about the functions, substrates, and regulatory mechanisms for the vast majority of PPR proteins.The ysa mutant develops albino leaves before the three-leaf stage, but the mutant gradually turns green and recovers to normal green at the six-leaf stage. Further investigation showed that the change in leaf color in ysa mutant is associated with changes in chlorophyll content and chloroplast development. The albino phenotype of the young ysaseedlings is caused by a reduction in total chlorophyll content, rather than reduction of a particular pigment.
Expression
Tissue localization Molecular analysis of an F2 population from the cross Taiziyuzhu3ysaplaced the YSAlocus between the markers RM411 and RM8208 on chromosome 3 . The YSAlocus was further narrowed down to a 45-kb region, which includes 10 putative open reading frames(ORFs; Fig. 3, B–D). We sequenced all ORFs and found a 5-bp deletion in Os03g40020, causing a premature stop codon. that the 5-bp deletion in Os03g40020 is responsible for the albino phenotype of ysa mutant. light plays a major role in regulatingYSAexpression.YSA is highly expressed in young leaves and stems, but not in the roots (Fig. 1, A–D). Quantitative real-time reverse transcription (RT)-PCR analysis revealed that the expression of YSA peaked in the fourth leaf (Fig. 2E). Thus, the expression pattern of YSA is consistent with the seedling-stage-specific albino phenotype of ysa mutant and further supports the notion that YSA plays an important role in chloroplast development in the first few leaves of rice seedlings, but plays more minor roles in later stages.
Figure 1. Phenotypic analysis of the ysa mutant plants. A to C, Phenotypes of Pei'ai64S (left) and ysa mutant (right) seedlings at 1 (A), 2 (B), and 3 (C) weeks after sowing. D, The pigment contents in leaves of 1-week-old ysa mutants are much lower than that in Pei'ai64S. E, The pigment contents in leaves of 6-week-old ysa mutants are similar to that of Pei'ai64S plants. Chla, Chlorophyll a; Chlb, chlorophyll b; Chl, total chlorophyll; Car, carotenoid. Bars represent sds of three measurements. Student’s t test was performed on the raw data; asterisk indicates statistical significance at P < 0.01.
Subcellular Localization of YSA Protein
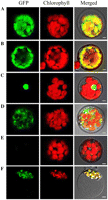
The YSA protein is predicted to localize to chloroplasts by ChloroP (Emanuelsson et al., 1999) and TargetP (Emanuelsson et al., 2000). To investigate the actual cellular localization of YSA, we constructed the green fluorescent signals of YSA-GFP fusion proteins colocalized with the autofluorescent signals of chlorophylls in the chloroplasts, consistent with the results obtained for GFP fused to the transit peptide of the small subunit of Arabidopsis ribulose bisphosphate carboxylase (Fig. 2, A and B). When GFP fused to the nuclear localization signal of the fibrillarin protein, GFP signals located specifically in the nucleus of Arabidopsis protoplasts (Fig. 2C). In addition, the protoplasts transformed with the empty GFP vector without a specific targeting sequence had green fluorescent signals in both the cytoplasm and the nucleus (Fig.2, D and E). To further confirm the subcellular localization of YSA protein, we transformed the plasmid containing the YSA-GFP fusion constructs into rice protoplasts. Confocal microscopy observations revealed that GFP-YSA was exclusively detected in the chloroplasts (Fig. 2F). These findings suggest that YSA protein is localized to the chloroplast.
Figure 2. Subcellular localization of YSA protein. Fluorescence signals were visualized using confocal laser-scanning microscopy. Green fluorescence shows GFP, red fluorescence indicates chloroplast autofluorescence, and yellow fluorescence indicates images with the two types of fluorescence merged. A, GFP signals of the YSA-GFP fusion protein. B, GFP signals from the transit peptide of ribulose bisphosphate carboxylase small subunit (control). C, GFP signals from the nuclear localization signal of fibrillarin (control). D, Empty GFP vector without a specific targeting sequence. E, Untransformed chloroplasts. F,Subcellular localization of YSA protein in rice protoplasts. Bars = 5mm. A to E are Arabidopsis protoplasts and F is rice protoplasts.
Evolution
Homology with Arabidopsis Similar to At3g53700: MEE40 (maternal effect embryo arrest 40) (HF=7e-1) You can also add sub-section(s) at will.
Labs working on this gene
- National Key Facility for Crop Gene Resources and Genetic Improvement, Institute of Crop Science, Chinese Academy of Agricultural Sciences, Beijing 100081, People’s Republic of China (N.S., F.-Q.W., G.-L.F., Y.L., X.-L.C.,X.Z., X.-P.G., Z.-J.C., C.-L.L., H.W., J.-M.W.);
- National Key Laboratory for Crop Genetics and Germplasm Enhancement, Nanjing Agricultural University, Nanjing 210095, People’s Republic of China (M.-L.H., L.J., J.-M.W.);
- Institute of Industrial Crops, Jiangsu Academy of Agricultural Sciences, Nanjing 210014, People’s Republic of China (M.-L.H., C.-K.Q.);
- State Key Laboratory of Rice Biology, International Atomic Energy Agency Collaborating Center, Institute of Nuclear Agricultural Sciences, Zhejiang University, Hangzhou 310029,People’s Republic of China (D.-X.W., X.-L.S.)
References
[1] Bentolila S, Alfonso AA, Hanson MR (2002) A pentatricopeptide repeat-containing gene restores fertility to cytoplasmic male-sterile plants. Proc Natl Acad Sci USA 99: 10887–10892
[2] Brown GG, Formanová N, Jin H, Wargachuk R, Dendy C, Patil P, Laforest M, Zhang J, Cheung WY, Landry BS (2003) The radish Rfo restorer gene of Ogura cytoplasmic male sterility encodes a protein with multiple pentatricopeptide repeats. Plant J 35: 262–272
[3] Desloire S, Gherbi H, Laloui W, Marhadour S, Clouet V, Cattolico L, Falentin C, Giancola S, Renard M, Budar F, et al. (2003) Identification of the fertility restoration locus, Rfo, in radish, as a member of the pentatricopeptide-repeat protein family. EMBO Rep 4: 588–594
[4] Koizuka N, Imai R, Fujimoto H, Hayakawa T, Kimura Y, Kohno-Murase J, Sakai T, Kawasaki S, Imamura J (2003) Genetic characterization of a pentatricopeptide repeat protein gene, orf687, that restores fertility in the cytoplasmic male-sterile Kosena radish. Plant J 34: 407–415
[5] Emanuelsson O, Nielsen H, von Heijne G (1999) ChloroP, a neural network-based method for predicting chloroplast transit peptides and their cleavage sites. Protein Sci 8: 978–984
[6] Emanuelsson O, Nielsen H, Brunak S, von Heijne G (2000) Predicting subcellular localization of proteins based on their N-terminal amino acid sequence. J Mol Biol 300: 1005–1016
[7] Su N, Hu ML, Wu DX, et al.(2012) Disruption of a rice pentatricopeptide repeat protein causes a seedling-specific albino phenotype and its utilization to enhance seed purity in hybrid rice production[J].Plant Physiology, 159(1):227-238.
Structured Information
| Gene Name |
Os03g0597200 |
|---|---|
| Description |
Protein prenyltransferase domain containing protein |
| Version |
NM_001057140.1 GI:115454008 GeneID:4333379 |
| Length |
5627 bp |
| Definition |
Oryza sativa Japonica Group Os03g0597200, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 3:22993430..22999056 |
| Sequence Coding Region |
22996530..22998758 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008396:22993430..22999056 source=RiceChromosome03 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008396:22993430..22999056 source=RiceChromosome03 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>atgccccgcgtttgcgccgcccctcgggcgccgccgccgccgtgcccgtgccatgtcggagtagggccgcttcggccgaggtggcgcgcctcccggcacggccctctccgggcggctggccaggagcagctcctcaccgccctgcgcgagcagccggaccccgacgcggcgctccggatgctcaacgcggcgctggcgcgggacgacttcgcgcccggccccgaggtctacgaggagatcatccgcaagctcggcgcggtcggggccctcgacctcatgaaggtgctcgtcgcggagatgcggcgggaggggcaccaggtgaaattgggcgtagtccactccttcttggacagctacgaggggcagcagctgttcgacgatgccgtcgacctgattctgaatcaactccaaccattgtttggcattcaggcagacaccgtggtgtacaatcaccttctcaatgttcttgtggaggggagcaaaatgaaactccttgaatcagtgtactcggagatgggtgctaggggaatcaagcctgatgttgtcacattcaacacactgatgaaggcgttgtgccgagcacatcaggtcaggactgcagttctcatgctcgaggaaatgtctagcagaggcgtggcgcctgacgagacgacgtttaccaccctgatgcaaggatttgtcgaggaggggagcatcgaggctgcactgagggtcaaagccaggatgttggaaatggggtgctcggcgacgaaggtgacggttaatgttttgattaatgggtactgcaagctagggagggtggaggatgctcttgggtatatacagcaggagattgccgatgggtttgagcctgaccagatcacatataacacttttgttaatggtctctgccaaaatgatcatgtcggccatgccctcaaagtcatggatgtgatggttcaggagggccatgatcctgatgttttcacctacaatatcgttgtgaattgcctttgtaaaaatggacagcttgaagaggcaaaaggaattctgaatcagatggtggatcggggttgtttgcctgacattaccacattcaacactctcattgctgccttatgcacggggaatcgacttgaggaagccttggaccttgcacgtcaggttacagtgaagggagtctctccagatgtttatactttcaatattctgattaacgcgctctgcaaagtaggcgatcctcatcttgcacttcgattgtttgaagagatgaagaacagtggatgcaccccggatgaagtaacatacaatactttgattgacaatctttgctcacttgggaagcttggtaaagcattggatttgttaaaagatatggagtccactggttgtcctcgaagtacaattacatataacactataattgacgggttatgcaagaaaatgagaatagaagaagcagaagaagtttttgatcaaatggatctgcaaggcatttcgaggaatgcaatcacattcaatactctcatcgatggtttgtgcaaggacaaaaagattgatgatgcttttgagcttattaatcaaatgataagtgaagggttgcaacctaacaatatcacttacaattctattctaactcattattgcaagcaaggtgacataaaaaaggctgcggatattttagaaactatgactgcaaatggatttgaagtggatgttgttacgtacggtactctgattaacggtctatgcaaggctggtaggacacaggttgctttgaaggtactcagaggtatgcggataaaagggatgaggcctactccaaaagcttacaatcctgtgctccagtctctcttcagacggaataatatcagagatgccctgagtcttttcagggagatggcagaggttggtgagcctcctgatgctttgacatataagattgtttttcgtgggctctgtcgtggtggagggcctattaaagaagcttttgatttcatgttggagatggttgataaggggttcataccagagttctcatccttccgtatgctagctgaaggtctattaaacctgggtatggatgattacttcattagagccattgaaataatcatggaaaaggtcgacctcagagagtctgatgtttctgcaataaggggatatctcaagatccgcaaattttatgatgcattagcaacctttggccgtttcctggagatcaacaaccctcaatggagttaccgatga</cdnaseq> |
| Protein Sequence |
<aaseq>MPRVCAAPRAPPPPCPCHVGVGPLRPRWRASRHGPLRAAGQEQL LTALREQPDPDAALRMLNAALARDDFAPGPEVYEEIIRKLGAVGALDLMKVLVAEMRR EGHQVKLGVVHSFLDSYEGQQLFDDAVDLILNQLQPLFGIQADTVVYNHLLNVLVEGS KMKLLESVYSEMGARGIKPDVVTFNTLMKALCRAHQVRTAVLMLEEMSSRGVAPDETT FTTLMQGFVEEGSIEAALRVKARMLEMGCSATKVTVNVLINGYCKLGRVEDALGYIQQ EIADGFEPDQITYNTFVNGLCQNDHVGHALKVMDVMVQEGHDPDVFTYNIVVNCLCKN GQLEEAKGILNQMVDRGCLPDITTFNTLIAALCTGNRLEEALDLARQVTVKGVSPDVY TFNILINALCKVGDPHLALRLFEEMKNSGCTPDEVTYNTLIDNLCSLGKLGKALDLLK DMESTGCPRSTITYNTIIDGLCKKMRIEEAEEVFDQMDLQGISRNAITFNTLIDGLCK DKKIDDAFELINQMISEGLQPNNITYNSILTHYCKQGDIKKAADILETMTANGFEVDV VTYGTLINGLCKAGRTQVALKVLRGMRIKGMRPTPKAYNPVLQSLFRRNNIRDALSLF REMAEVGEPPDALTYKIVFRGLCRGGGPIKEAFDFMLEMVDKGFIPEFSSFRMLAEGL LNLGMDDYFIRAIEIIMEKVDLRESDVSAIRGYLKIRKFYDALATFGRFLEINNPQWS YR</aaseq> |
| Gene Sequence |
<dnaseqindica>299..2527#ctcctgttcccctctgcctgccttcacggagaacacgccgccgcacgcccgcaaagttgtcgctccgccgccgggtcctgcggccacttcctccctctccctgtgcatgcgctctcttccccacctgtactttactttagctgctcctctgcccagttgcccacgacctgacgacccggacatggcgcaggctgaggcggggacgacgacatcgccggcgggttgacgcagaaaggagcgaccacccgagggctccgctggattttcaggtagctgagctgagctgaactgaaccccaatgccccgcgtttgcgccgcccctcgggcgccgccgccgccgtgcccgtgccatgtcggagtagggccgcttcggccgaggtggcgcgcctcccggcacggccctctccgggcggctggccaggagcagctcctcaccgccctgcgcgagcagccggaccccgacgcggcgctccggatgctcaacgcggcgctggcgcgggacgacttcgcgcccggccccgaggtctacgaggagatcatccgcaagctcggcgcggtcggggccctcgacctcatgaaggtgctcgtcgcggagatgcggcgggaggggcaccaggtgaaattgggcgtagtccactccttcttggacagctacgaggggcagcagctgttcgacgatgccgtcgacctgattctgaatcaactccaaccattgtttggcattcaggcagacaccgtggtgtacaatcaccttctcaatgttcttgtggaggggagcaaaatgaaactccttgaatcagtgtactcggagatgggtgctaggggaatcaagcctgatgttgtcacattcaacacactgatgaaggcgttgtgccgagcacatcaggtcaggactgcagttctcatgctcgaggaaatgtctagcagaggcgtggcgcctgacgagacgacgtttaccaccctgatgcaaggatttgtcgaggaggggagcatcgaggctgcactgagggtcaaagccaggatgttggaaatggggtgctcggcgacgaaggtgacggttaatgttttgattaatgggtactgcaagctagggagggtggaggatgctcttgggtatatacagcaggagattgccgatgggtttgagcctgaccagatcacatataacacttttgttaatggtctctgccaaaatgatcatgtcggccatgccctcaaagtcatggatgtgatggttcaggagggccatgatcctgatgttttcacctacaatatcgttgtgaattgcctttgtaaaaatggacagcttgaagaggcaaaaggaattctgaatcagatggtggatcggggttgtttgcctgacattaccacattcaacactctcattgctgccttatgcacggggaatcgacttgaggaagccttggaccttgcacgtcaggttacagtgaagggagtctctccagatgtttatactttcaatattctgattaacgcgctctgcaaagtaggcgatcctcatcttgcacttcgattgtttgaagagatgaagaacagtggatgcaccccggatgaagtaacatacaatactttgattgacaatctttgctcacttgggaagcttggtaaagcattggatttgttaaaagatatggagtccactggttgtcctcgaagtacaattacatataacactataattgacgggttatgcaagaaaatgagaatagaagaagcagaagaagtttttgatcaaatggatctgcaaggcatttcgaggaatgcaatcacattcaatactctcatcgatggtttgtgcaaggacaaaaagattgatgatgcttttgagcttattaatcaaatgataagtgaagggttgcaacctaacaatatcacttacaattctattctaactcattattgcaagcaaggtgacataaaaaaggctgcggatattttagaaactatgactgcaaatggatttgaagtggatgttgttacgtacggtactctgattaacggtctatgcaaggctggtaggacacaggttgctttgaaggtactcagaggtatgcggataaaagggatgaggcctactccaaaagcttacaatcctgtgctccagtctctcttcagacggaataatatcagagatgccctgagtcttttcagggagatggcagaggttggtgagcctcctgatgctttgacatataagattgtttttcgtgggctctgtcgtggtggagggcctattaaagaagcttttgatttcatgttggagatggttgataaggggttcataccagagttctcatccttccgtatgctagctgaaggtctattaaacctgggtatggatgattacttcattagagccattgaaataatcatggaaaaggtcgacctcagagagtctgatgtttctgcaataaggggatatctcaagatccgcaaattttatgatgcattagcaacctttggccgtttcctggagatcaacaaccctcaatggagttaccgatgaagcagaatacataactgggacaaattacttgaatagtattagggaaatctcaaagagtggatggaatttttgctggttgcttaggggaatgaaagtcttaaattgattataataggtgattgtgttcattcctcggtagggatgaagtcagagcatgaagaagctcatcttggtgcagaaacttagcttattggaacagaatctaggtgctaggtgctgtctgctgccagattagtacctagcgccttaacgaacagtggctgcagatcccatccgcttattttgatcaaatctgattcattttctattccctaataaaagcctgattcatcttcattgcatatggtcgaacctaaggctatcatgtacagttagatcccaacccttcgttctatgagatgttgtcccatagaagaaatatcttctgagtatatcctagtactctaaaatgttcactaatatgttcaacttaattagttttgtaacctcctaaaatagttctattagttttgtaaccttctaaaatatgaattagttttgatctggctgatcttcctttggttaggtactacaaattcttaattcagacacatatttggttttctgaaaatttcatctatatttggtcaggctggcatttgaagttcttattttagtcatatactttagttttttacaattttcatttgttaaagatgataatttatttgttagcacagagcatgtttagaaatctgaaatattaaaacatgcatgttctcatgaaaataaatgttagttttgtttaaattccaatccacatattttttaatcaatgtcagaaattaccatgcttcacttattgacctagtatatgtatagtatttgatggatcatgttgatttggatttgctctactaacttgttcctattccaacaaagatttatatgcatcttgtgttctaaaaatgctacatgtgtcaagttgaaggaaaattctagctatgtggtgtcctaattttggtagatggtacctagtaatagataacaattctgttttatagtgatgagtaaatttgactaaatcgagtctagaaagtgatataattttctggagaagtctttcttggtgatttgggaaagggccattacctatactgatatgaaatctggaactagaaagatctgaacatcaatgttctaaagttttttgtctgaatttcttgtgcagaatatgaaggaaggtggatctggaataggtatttacatgtcctgttcagattcctgcaatctgataaactactgcaatccaataaactcttgatctattgttttctggatttttttttgggtgtagagtattaggaaggtatttgctttgttacaggggttggttggatgttcagaaaaccaaaatctgaatacactaacacattagccagtgttttattttagtttatgttattctgaccacaacaccagcagcttcaattggtagtagaacacactctgatgcaattggtaggtgtacaacagataatttgccgtgaatgctacttaattcaaccattttttttccatacagtgctatacaatggcacacaaacatccttcagttggactcaaatttgtttcagctttgctattttacagtcacctgcagttttttctaatccatcagtcgattttgttggcagtagattctgcttcaagatctgtgcacgcagtggaggcacatttggttacaggctgcttcaccagacccgttaacccatgggccatggatgctcccctagtagcatatgttgttttttccaccaactgcctcttgtaaatcaagatgctcccctctccacatttgctgcagacttcctccccgtggatctgagacgaactccggcgaccggcggcgtctacgagccagcgttcaccaatttctcaggtatatgagtcgtcatctatgtgcatctgctactttttttttttggctgattggacgggttgagaaggaggtgtttggggggaagagagcagaataatcccattgcgaaacaagcgaggaaggcttgtcgatggtggcggcggtggccttgattatcgaagatgcatgcgtgagggaaactgtgtgccagggaggcaagcttggtggcacacatcgagtggtatatcttcaggttactgctagatcgaagatcctttggtgcttgatgtgtgctgttggtttggtagttttggtggtactgtggcaagtgaagacacgaaagaggagctagtagaggcttagagtctggagtcgtaatcgatgtgcaagagcatcaatgccatcagttcctctgggcagcacggctctgcacttttcatctgagaaacatatatattttgttgattctgattttgacgaattatccaaacatgcagtttttttgggttccgcgtgcaaagacgattttgatctcctggcagatttaagggtggtgattttgtttttgttttgcatgtagatgatgcattttgtaaatcattttaggttgccccttttttttttgtttttgcgagttcttactgaatttggataagttctggttgattggatcgacattctgtacatttgcaaggggattctatgggtttttgcttattgctaatcaaattattttgtgagttttggtttattggatggtgagaaagcaagattacatggattttgcttcattgctgattttgtcgaaatttatcaggcatgcggtttttggtttcttgaatacgaactacacggacagtactcctatattcttggtgattttgaagtgtatttgttattttggggaggtggaatggttaatattttgttcaaggggacacttctacagatgtatacattacttttgtcattttcagcataacaaagagatttggtagattcagaattcagatggcaactacacgtcgaatgatgtatgcgaagacatggggaattctgatgctgattccactgaagaaagctaaacataattttaatttatttaccaactcatttttctgtcaacacatttgctagatagttgctacccgataaacgacatcttgaaatctgagttatacttcagtctattttttttcc</dnaseqindica> |
| External Link(s) |