Difference between revisions of "Os05g0497200"
Jin Xiaoyang (talk | contribs) |
Jin Xiaoyang (talk | contribs) (→References) |
||
| Line 37: | Line 37: | ||
==References== | ==References== | ||
| − | + | <ref name="ref1">Ren D, Li Y, Zhao F, et al. MULTI-FLORET SPIKELET1, which encodes an AP2/ERF protein, determines spikelet meristem fate and sterile lemma identity in rice[J]. Plant physiology, 2013, 162(2): 872-884.</ref> | |
| − | + | <ref name="ref2">Sharoni A M, Nuruzzaman M, Satoh K, et al. Gene structures, classification and expression models of the AP2/EREBP transcription factor family in rice[J]. Plant and cell physiology, 2011, 52(2): 344-360.</ref> | |
| − | + | <ref name="ref3">Lee D Y, Lee J, Moon S, et al. The rice heterochronic gene SUPERNUMERARY BRACT regulates the transition from spikelet meristem to floral meristem[J]. The Plant Journal, 2007, 49(1): 64-78.</ref> | |
| − | + | <ref name="ref4">Lee D Y, An G. Two AP2 family genes, supernumerary bract (SNB) and Osindeterminate spikelet 1 (OsIDS1), synergistically control inflorescence architecture and floral meristem establishment in rice[J]. The Plant Journal, 2012, 69(3): 445-461.</ref> | |
| − | + | <ref name="ref5">Yoshida A, Suzaki T, Tanaka W, Hirano HY (2009) The homeotic gene long sterile lemma (G1) specifies sterile lemma identity in the rice spikelet. Proc Natl Acad Sci USA 106: 20103–20108 6.Hong L, Qian Q, Zhu K, Tang D, Huang Z, Gao L, Li M, Gu M, Cheng Z (2010) ELE restrains empty glumes from developing into lemmas. J Genet Genomics 37: 101–115</ref> | |
| − | + | <references /> | |
| − | |||
| − | |||
| − | |||
==Structured Information== | ==Structured Information== | ||
Revision as of 13:30, 31 May 2014
Please input one-sentence summary here.
Contents
Annotated Information
MULTI-FLORET SPIKELET1, which encodes an AP2/ERF protein, determines spikelet meristem fate and sterile lemma identity in rice.
Function
MFS1 encodes an ERF domain protein. MFS1 regulates palea development, specifies sterile lemma identity,and affects spikelet meristem determinacy.
A rice (Oryza sativa) spikelet mutant, multi-floret spikelet1 (mfs1) shows pleiotropic defects in spikelet development.Mfs1 has delayed transformation of spikelet meristems to floral meristems, which resulted in an extra hull-like organ and an elongated rachilla.(Figure.1) In addition, the sterile lemma was homeotically converted to the rudimentary glume and the body of the palea was degenerated in mfs1. These results suggest that the MULTI-FLORET SPIKELET1 (MFS1) gene plays an important role in the regulation of spikelet meristem determinacy and floral organ identity. MFS1 also positively regulates the expression of LONG STERILE LEMMA and the INDETERMINATE SPIKELET1 (IDS1)-like genes SUPERNUMERARY BRACT and OsIDS1.MFS1 belongs to an unknown function clade in the APETALA2/ethylene-responsive factor (AP2/ERF) family. The MFS1-green fluorescent protein fusion protein is localized in the nucleus.MFS1 messenger RNA is expressed in various tissues, especially in the spikelet and floral meristems.

Expression
The MFS1-green fluorescent protein fusion protein is localized in the nucleus. MFS1 messenger RNA is expressed in various tissues,especially in the spikelet and floral meristems. Quantitative reverse transcription-PCR (qPCR) analysis showed that MFS1 was universally expressed in various tissues, including roots, stems, leaves, and panicles, with higher levels in young panicles (2 cm or less) than in the other tissues examined (Fig. 2A).Furthermore, the MFS1 expression pattern was investigated by in situ hybridization. First, MFS1 was highly expressed in the meristems of branches and spikelets (Fig. 2, B–D). Next, strong signals were observed at the sites of initiation of the sterile lemma primordium (Fig. 2E). When the lemma and palea primordia formed, abundant MFS1 transcripts were detected in the lemma, palea, and floral meristem (Fig.2, D, F, and G). Subsequently, the expression of MFS1 was primarily restricted to the lemma, palea, lodicule, and stamen (Fig. 2, G and H). After the formation of pistil, MFS1 signals disappeared from the lemma and palea but were retained in the lodicule, stamen, and pistil (Fig. 2, I and J).

Evolution
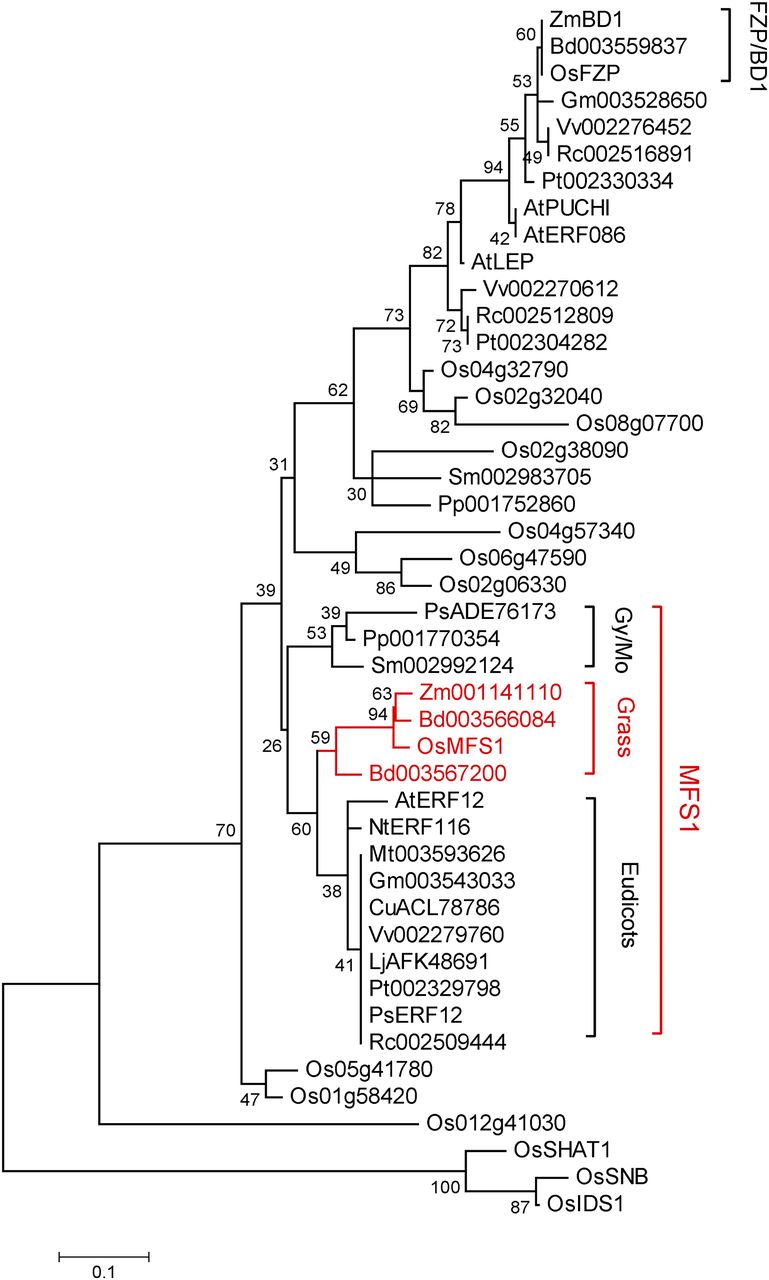
MFS1 Encodes an ERF Domain Protein The AP2/ERF gene family is plant specific and includes four subfamilies: AP2, RAV, DREB, and ERF. Phylogenetic analysis showed that MFS1 and its orthologs from moss, gymnosperms, dicots, and grasses constitute an MFS1-like clade, whereas the well-known ERF domain proteins FZP and BD1 and their orthologs constitute another clade in the ERF subfamily (Fig. 3). These results suggested that MFS1-like and FZP/BD1-like genes diverged before the emergence of gymnosperms and that the MFS1-like genes differ from the well-known AP2/ERF genes. In addition, phylogenetic analysis also showed that the other known AP2 domain genes (SNB, OsIDS1, and SHAT1) have a distant evolutionary relationship with the MFS1-like and FZP/BD1-like genes.
Sequence analysis showed that all MFS1-like proteins contain a highly conserved ERF domain, located close to their N terminus. Meanwhile, a conserved C-terminal domain was identified in MFS1-like proteins from grasses and dicots, which share the DLNEPP185-190 motif. A unique site (V37) and a motif (SPWH132-135) were also identified in MFS1-like proteins from grasses .In addition, the MFS1 gene shared low sequence similarity with the known AP2/ERF genes outside the AP2/ERF domain.
MFS1 Affects the Expression of Genes Related to Spikelet Development
Given that the mfs1-1 mutant exhibited spikelet defects,the expression levels of the IDS1-like genes SNB and OsIDS1, which are closely associated with the transition and determinacy of spikelet meristem in rice were examined. SNB transcripts accumulated primarily in young panicles less than 2 cm long, and their levels were lower in the mfs1-1 mutant than in the wild type (Fig. 4A). Then, levels of SNB transcripts were dramatically decreased in panicles longer than 2 cm, and no difference in the levels of SNB expression was found between wild-type and mfs1-1 panicles with a length 2 to 5 cm (Fig. 4A). OsIDS1 transcripts were first detected in young panicles less than 0.5 cm, and they were more abundant in wild-type panicles between 0.5 and 5 cm in length (Fig. 4A). Compared with that in the wild type, OsIDS1 expression showed no obvious change in panicles with a length less than 0.5 cm, whereas it dramatically decreased in mfs1-1 panicles 0.5 to 5 cm long (Fig. 4A). These results imply that MFS1 positively regulated the expression of the IDS1-like genes SNB and OsIDS1.(Figure 4.)
Using qPCR to examine the expression of the G1 gene, which has been shown to be involved in the specification of sterile lemma identity. In the wild type, a high level of G1 expression was detected in panicles shorter than 2 cm, but the mRNA levels were significantly reduced in those that were 2 to 5 cm (Fig. 4A). In the mfs1-1 mutant, G1 showed lower expression levels in young panicles shorter than 5 cm (Fig. 4A). In situ hybridization indicated that in the wild type, the G1 signals were strongly detected in sterile lemmas during the stage of sterile lemma primordia differentiation and formation, and subsequently, they decreased markedly when sterile lemmas started to elongate (Fig. 4, B–F). G1 expression was faint in the sterile lemma primordia of the mfs1-1 spikelet during the stages analyzed (Fig. 4, G–K), consistent with the results of qPCR analysis. These findings suggest that MFS1 positively regulates G1 expression.

Labs working on this gene
Rice Research Institute (D.R., Yu.L., F.Z., X.S., J.S., N.W., S.G., Yi.L., C.Z., Z.Y., G.H.), Chongqing Key Laboratory of Application and Safety Control of Genetically Modified Crops (D.R., Yu.L., F.Z., X.S., J.S., N.W., S.G., C.Z., Z.Y., G.H.), and Engineering Research Center of South Upland Agriculture, Ministry of Education (Yi.L., G.H.), Southwest University, Chongqing 400715, China; and Engineering Research Center of South Upland Agriculture, Ministry of Education, Chongqing 400715, China (Yu.L., F.Z., X.S., N.W., S.G., Yi.L., C.Z., Z.Y., G.H.)
References
- ↑ Ren D, Li Y, Zhao F, et al. MULTI-FLORET SPIKELET1, which encodes an AP2/ERF protein, determines spikelet meristem fate and sterile lemma identity in rice[J]. Plant physiology, 2013, 162(2): 872-884.
- ↑ Sharoni A M, Nuruzzaman M, Satoh K, et al. Gene structures, classification and expression models of the AP2/EREBP transcription factor family in rice[J]. Plant and cell physiology, 2011, 52(2): 344-360.
- ↑ Lee D Y, Lee J, Moon S, et al. The rice heterochronic gene SUPERNUMERARY BRACT regulates the transition from spikelet meristem to floral meristem[J]. The Plant Journal, 2007, 49(1): 64-78.
- ↑ Lee D Y, An G. Two AP2 family genes, supernumerary bract (SNB) and Osindeterminate spikelet 1 (OsIDS1), synergistically control inflorescence architecture and floral meristem establishment in rice[J]. The Plant Journal, 2012, 69(3): 445-461.
- ↑ Yoshida A, Suzaki T, Tanaka W, Hirano HY (2009) The homeotic gene long sterile lemma (G1) specifies sterile lemma identity in the rice spikelet. Proc Natl Acad Sci USA 106: 20103–20108 6.Hong L, Qian Q, Zhu K, Tang D, Huang Z, Gao L, Li M, Gu M, Cheng Z (2010) ELE restrains empty glumes from developing into lemmas. J Genet Genomics 37: 101–115
Structured Information
| Gene Name |
Os05g0497200 |
|---|---|
| Description |
Similar to Ethylene-responsive transcription factor 11 (Ethylene-responsive element binding factor 11) (EREBP-11) (AtERF11) |
| Version |
NM_001062476.2 GI:297604695 GeneID:4339208 |
| Length |
915 bp |
| Definition |
Oryza sativa Japonica Group Os05g0497200, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome |
{{{Chromosome}}} |
| Location |
Chromosome 5:24475113..24476027 |
| Sequence Coding Region |
24475273..24475842 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008398:24475113..24476027 source=RiceChromosome05 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008398:24475113..24476027 source=RiceChromosome05 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>atggagctggacatgggagcgggaggaggcggtggagtagtgggaggtgggcgagcggaggcgcactaccgcggggtgaggaagcggccgtggggccggtacgcggcggagatccgggacccgtggaagaagacgcgggtgtggctcggcacctacgacacgcccgtcgaggccgcgctcgcctacgaccgcgccgccgtcgcgctccgcggcgtcaaggcgcggaccaacttcggcagcggcagcagcggtggtggtggcgtcggcgggcacggccatggccacagccacgcccagctgccgcagcttcaccaccgcatgcacccgccgcggccgccgcagggccctggtcacttcggcgggctcgacatcagccacccttcgccgtggcactatgtctacttcccggcgagggtgcaggcgatggcgccggcggcggctggccatgtcgcggcgcacgtcgccgcgtcgctgccgtcgacgacgctggagctccggacggggccgagcgccggcgagctcccgttcgacctcaacgagccgccgccggcgctgctgttcggctcgtga</cdnaseq> |
| Protein Sequence |
<aaseq>MELDMGAGGGGGVVGGGRAEAHYRGVRKRPWGRYAAEIRDPWKK TRVWLGTYDTPVEAALAYDRAAVALRGVKARTNFGSGSSGGGGVGGHGHGHSHAQLPQ LHHRMHPPRPPQGPGHFGGLDISHPSPWHYVYFPARVQAMAPAAAGHVAAHVAASLPS TTLELRTGPSAGELPFDLNEPPPALLFGS</aaseq> |
| Gene Sequence |
<dnaseqindica>186..755#ggcgcgcgcaaacgcaaagccgacgcgagagcgagcggccaccgcgcgctagtagtagtagcgagccactatagatttggctcccgtataaagggtcgctgagacttgcttgtcgctggcagtggcagctgccgatccgtccgtcaaggcacagcgcaagcacaggctctcgctcgtggaggcgaatggagctggacatgggagcgggaggaggcggtggagtagtgggaggtgggcgagcggaggcgcactaccgcggggtgaggaagcggccgtggggccggtacgcggcggagatccgggacccgtggaagaagacgcgggtgtggctcggcacctacgacacgcccgtcgaggccgcgctcgcctacgaccgcgccgccgtcgcgctccgcggcgtcaaggcgcggaccaacttcggcagcggcagcagcggtggtggtggcgtcggcgggcacggccatggccacagccacgcccagctgccgcagcttcaccaccgcatgcacccgccgcggccgccgcagggccctggtcacttcggcgggctcgacatcagccacccttcgccgtggcactatgtctacttcccggcgagggtgcaggcgatggcgccggcggcggctggccatgtcgcggcgcacgtcgccgcgtcgctgccgtcgacgacgctggagctccggacggggccgagcgccggcgagctcccgttcgacctcaacgagccgccgccggcgctgctgttcggctcgtgatctcgacacgtacgtgagtagagcatcactgggtagtactaggaccaaggatcaagcttagagatcactagtagataggggtaaccgattaatgtgggagtggagagcactggtccggctgtgtacatacgtactactatttaaggaaagtttagcgttc</dnaseqindica> |
| External Link(s) |