Difference between revisions of "Os01g0253300"
Ibpchenwenbo (talk | contribs) m (→Protein structure) |
Ibpchenwenbo (talk | contribs) m (→References) |
||
| Line 57: | Line 57: | ||
<ref name="ref4"> | <ref name="ref4"> | ||
Shoji K, Iwasaki T, Matsuki R, Miyao M, Yamamoto N. Cloning of a cDNA encoding an importin-alpha and down-regulation of the gene by light in rice leaves. Gene. 1998 Jun 8;212(2):279-86.</ref> | Shoji K, Iwasaki T, Matsuki R, Miyao M, Yamamoto N. Cloning of a cDNA encoding an importin-alpha and down-regulation of the gene by light in rice leaves. Gene. 1998 Jun 8;212(2):279-86.</ref> | ||
| − | + | <ref name="ref5"> | |
| + | Chang CW1, Couñago RL, Williams SJ, Bodén M, Kobe B.Crystal structure of rice importin-α and structural basis of its interaction with plant-specific nuclear localization signals.Plant Cell. 2012 Dec;24(12):5074-88. doi: 10.1105/tpc.112.104422. Epub 2012 Dec 18.</ref> | ||
<ref name="ref6"> | <ref name="ref6"> | ||
Iwasaki T, Matsuki R, Shoji K, Sanmiya K, Miyao M, Yamamoto N. A novel importin alpha from rice, a component involved in the process of nuclear protein transport. FEBS Lett. 1998 May 29;428(3):259-62.</ref> | Iwasaki T, Matsuki R, Shoji K, Sanmiya K, Miyao M, Yamamoto N. A novel importin alpha from rice, a component involved in the process of nuclear protein transport. FEBS Lett. 1998 May 29;428(3):259-62.</ref> | ||
Revision as of 14:18, 6 June 2014
Please input one-sentence summary here.
Contents
Annotated Information
Function

Os01g0253300 encodes importin subunit alpha-1a,a member of importin family, which functions a essential role in nuclear protein import by specifically and directly binding to substrates containing either a simple or bipartite NLS motif[2].In vivo assay suggests that importin alpha-1a can replace vertebrate importin a in the mediation of nuclear import of NLS substrates, implying that rice importin a1 functions as a NLS receptor in the process of nuclear import of proteins(Fig-1)[1],which indicates the importin family has a conserved transportation function both in plants and mammals.The importin alpha-1a also has a function in virus infection by interact with mungbean yellow mosaic virus capsid protein[3].
Expression

Transcript analyses of rice importin alpha-1a by Chang-Jie Jiang et al shows importin alpha-1a is highly expressed in callus, followed by root and etiolated leaf and lowly expressed in green leaf(Fig-2)[2].The expression level of importin alpha-1a is greatly increased after darkt treatment in green leaf[Fig-3][4] .However it has a constitute expression pattern in non-photosynthetic tissues[Fig-4][3].Together,it indicates a complex regulation of the importin alpha-1a transcription.
Fig-4.Importin-atranscript levels in different rice tissues.(A) RNA was prepared from various tissues, roots (Root), mature leaf blades (M-Leaf), flag leaf blades (F-Leaf), stems (Stem) and suspension-cultured cells (Callus), and the transcript levels for importin-α in their tissues were examined by RNA blot analysis. Mature leaves, flag leaves and stems of field-grown plants were collected at noon on a sunny day in July. (B) The effect of illumination on the transcript levels for importin-α was examined in roots and suspension-cultured cells. Roots of water-cultured seedlings and suspension-cultured cells were exposed to white light (white fluorescent lamps; 90 μmol m−2 s−1) for 24 h (L), kept in complete darkness for 24 h (D) or kept in darkness for 24 h and then re-illuminated for 3 h (+3hL).From ref [3]
Protein structure
1.The structure of rice importin alpha-1a comprises 10 ARM repeats (green, cartoon representation) and two NLS-like sequences from the N-terminal IBB domain(shown in yellow stick )[FIG-5][5].

Fig-5:crystal structure of importin alpha-1a.From ref[5]
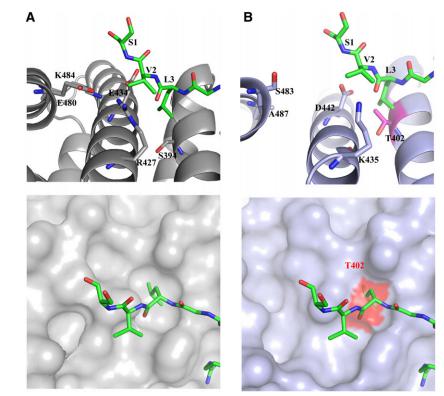
2.The crystal structures of their complexes with rice importin alpha-1a show that they bind to the minor NLS binding site[Fig6A]. By contrast, the crystal structures of their complexes with mouse (Mus musculus) importin alpha-1ashow preferential binding to the major NLS binding site[Fig6B]. The results reveal the molecular basis of a number of features of the classical nuclear transport pathway specific to plants[5].
Fig-6:Differential Binding of Plant-Specific NLSs to rImpa1a and mImpa.[5]
Evolution
The phylogenetic tree of importin alpha shows that the rice importin alpha-1a protein is the most distant member from various organisms[Fig-7][6].we could find another EST from Arabidopsis (accession number F15465), which has about 60% identity with #61L at the levelof both nucleotide and amino acid sequences. It might be interpreted that importin alpha-1a homologous to rice #61L(Refere to importin alpha-1a) is also present in Arabidopsis, and that the #61L protein and the homologue are functionally differentiated from the known importin alpha-1a.
Fig-7:(Phylogenetic tree of importinK. The tree was constructed by the UPGMA method using the GENETYX-MAC 7.3 software(Software Development Co., Tokyo) with default parameters. The accession number of the putative open reading frame T10M13.16 which is predicted ) From[6]
Labs working on this gene
- National Institute of Agrobiological Resources, Tsukuba, Ibaraki 305-8602, Japan.
- Friedrich Miescher Institute, Maulbeerstrasse 66, 4058 Basel, Switzerland.
- Central Research Institute of Electric Power Industry, Chiba, Japan.
- School of Chemistry and Molecular Biosciences and Institute for Molecular Bioscience, University of Queensland, Brisbane Qld 4072, Australia.
- School of Chemistry and Molecular Biosciences, Institute for Molecular Bioscience, and Australian Infectious Diseases Research Centre, University of Queensland, Brisbane, Qld 4072, Australia.
- The State Key Laboratory of Protein and Plant Gene Research, College of Life Sciences, Peking University, Beijing, China.
- Department of Biochemistry and Graduate Institute of Life Sciences, National Defense Medical Center, Taipei, Taiwan 114, Republic of China.
References
- ↑ 1.0 1.1 Jiang CJ, Imamoto N, Matsuki R, Yoneda Y, Yamamoto N. Functional characterization of a plant importin alpha homologue. J Biol Chem. 1998 Sep 11.
- ↑ 2.0 2.1 Jiang CJ, Shoji K, Matsuki R, Baba A, Inagaki N, Ban H, Iwasaki T, Imamoto N, Yoneda Y, Deng XW, Yamamoto N. Molecular cloning of a novel importin alpha homologue from rice, by which constitutive photomorphogenic 1 (COP1) nuclear localization signal (NLS)-protein is preferentially nuclear imported. J Biol Chem. 2001 Mar 23;276(12):9322-9. Epub 2000 Dec 20.
- ↑ 3.0 3.1 3.2 Guerra-Peraza O, Kirk D, Seltzer V, Veluthambi K, Schmit AC, Hohn T, Herzog E. Coat proteins of Rice tungro bacilliform virus and Mungbean yellow mosaic virus contain multiple nuclear-localization signals and interact with importin alpha. J Gen Virol. 2005 Jun;86(Pt 6):1815-26.
- ↑ 4.0 4.1 Shoji K, Iwasaki T, Matsuki R, Miyao M, Yamamoto N. Cloning of a cDNA encoding an importin-alpha and down-regulation of the gene by light in rice leaves. Gene. 1998 Jun 8;212(2):279-86.
- ↑ 5.0 5.1 5.2 5.3 Chang CW1, Couñago RL, Williams SJ, Bodén M, Kobe B.Crystal structure of rice importin-α and structural basis of its interaction with plant-specific nuclear localization signals.Plant Cell. 2012 Dec;24(12):5074-88. doi: 10.1105/tpc.112.104422. Epub 2012 Dec 18.
- ↑ 6.0 6.1 Iwasaki T, Matsuki R, Shoji K, Sanmiya K, Miyao M, Yamamoto N. A novel importin alpha from rice, a component involved in the process of nuclear protein transport. FEBS Lett. 1998 May 29;428(3):259-62.
Structured Information
| Gene Name |
Os01g0253300 |
|---|---|
| Description |
Importin alpha-1a subunit |
| Version |
NM_001049146.1 GI:115435705 GeneID:4327117 |
| Length |
5116 bp |
| Definition |
Oryza sativa Japonica Group Os01g0253300, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 1:8381095..8386210 |
| Sequence Coding Region |
8381261..8381470,8382333..8382429,8382526..8382617,8382709..8382840,8382955..8383140 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008394:8381095..8386210 source=RiceChromosome01 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008394:8381095..8386210 source=RiceChromosome01 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>atgtcgctgcgcccgagcgagcgggtggaggtgcggcggaaccgctacaaggtggcggtggacgccgaggaggggaggcggcggcgcgaggacaacatggtggagatccgcaagagccgccgcgaggagagcctcctcaagaagcgccgcgaggggctccaggcccaggcccccgtccccgcctccgccgcaaccggcgtcgataagaagctcgaaagccttcctgctatgattggtggagtttattcggacgataacaaccttcagcttgaggccacaacacagttccgcaagttgctttctatagagaggagccctccaattgaagaggttattcaatcaggcgttgttcctagattcgtgcagtttcttaccagagaggatttcccccagctacagttcgaagctgcgtgggcacttacaaacattgcatctggcacttcggagaatacaaaggttgtcattgatcatggggctgtaccaatatttgtgaagcttcttggttcttctagtgatgatgttcgtgagcaggctgtatgggcattgggcaatgttgctggtgattcccctaagtgccgtgaccttgttcttgccaatggtgcattgctgcctctgctagcacagttgaatgagcacactaaactctctatgctaaggaatgcaacttggactttatcgaacttctgcagaggaaaaccacagccatcatttgaacagactaggcctgctcttccagcactggcacggcttattcactccaatgatgaggaagttctcactgatgcgtgctgggctctttcatatttatctgatggcactaatgacaagatccaagccgtgattgaagctggtgtttgcccccgtcttgtggagcttctcctccatccatcaccttcagtgcttatacccgcactacgaactgttggaaatattgtaactggagatgacgcgcaaactcagtgcatcattgatcatcaagccctcccttgcctcctaagtctgttgacacaaaatctcaagaaaagcatcaagaaagaggcttgctggactatttcaaatattactgctggtaacaaagatcagatacaggctgtcataaatgctggcattattggtcctctggtgaatctgctccaaacggcagagttcgatatcaagaaggaagctgcatgggccatctcaaatgctacttcaggcggtagccatgatcaaatcaagtatctggtgagcgagggctgcatcaagccattgtgtgatcttcttatttgcccagatataagaatcgttacagtttgtttggagggtctcgagaacattcttaaagtaggagaaaccgacaagaccttggctgcaggtgatgtgaatgtcttttcccagatgattgatgaagcggaaggtttggaaaagattgagaacttgcagagccatgataacaatgagatctatgaaaaagctgtgaagattcttgaggcctactggatggatgaggaggatgatacaatgggagctaccacggtagctgctccccaaggtgctacttttgactttggccaaggcggtggtgctgctcaattcaaataa</cdnaseq> |
| Protein Sequence |
<aaseq>MSLRPSERVEVRRNRYKVAVDAEEGRRRREDNMVEIRKSRREES LLKKRREGLQAQAPVPASAATGVDKKLESLPAMIGGVYSDDNNLQLEATTQFRKLLSI ERSPPIEEVIQSGVVPRFVQFLTREDFPQLQFEAAWALTNIASGTSENTKVVIDHGAV PIFVKLLGSSSDDVREQAVWALGNVAGDSPKCRDLVLANGALLPLLAQLNEHTKLSML RNATWTLSNFCRGKPQPSFEQTRPALPALARLIHSNDEEVLTDACWALSYLSDGTNDK IQAVIEAGVCPRLVELLLHPSPSVLIPALRTVGNIVTGDDAQTQCIIDHQALPCLLSL LTQNLKKSIKKEACWTISNITAGNKDQIQAVINAGIIGPLVNLLQTAEFDIKKEAAWA ISNATSGGSHDQIKYLVSEGCIKPLCDLLICPDIRIVTVCLEGLENILKVGETDKTLA AGDVNVFSQMIDEAEGLEKIENLQSHDNNEIYEKAVKILEAYWMDEEDDTMGATTVAA PQGATFDFGQGGGAAQFK</aaseq> |
| Gene Sequence |
<dnaseqindica>167..376#1239..1335#1432..1523#1615..1746#1861..2046#2130..2293#2400..2478#3370..3498#3988..4115#4234..4597#gtgtgagctttaactccccccctttccccgcggccactagggtttctcctcctctttcctctcctcctccgccccgacgcctcgctagggtttttgcctctccctccgccgccgccgctgccgcgaggagaggagggggggagagcacccagccggcgagccagccatgtcgctgcgcccgagcgagcgggtggaggtgcggcggaaccgctacaaggtggcggtggacgccgaggaggggaggcggcggcgcgaggacaacatggtggagatccgcaagagccgccgcgaggagagcctcctcaagaagcgccgcgaggggctccaggcccaggcccccgtccccgcctccgccgcaaccggcgtcgataagaaggtgagtagtatactcctcgattcccccccgctcccctcccctcccctcccctgcctccgatgacgctatcccatcgtgtcctggatcccatattttaggtaggtgttggattggattgcactttttgagagggtttttgattcgtctgcggcatatttgtttaagtttgttattatacccgcatgtctactagcgtgcgggtgagggtaggttcctgctagatgtaggcaacacttagcaccacgtttcgatgcctcgattagtcacttgatttctgattctccttctgttgacaatattacaagtactgttagttggcacggttgcgaactcctagcgtttgaaatgtttccaggtacatgcctgctagctccagtgatctagttttagttttaggacatgtagattgtggtcttaggatcaaatcttccacggattcactgcggataattagttaggtgaccaggaactgtaataatgattgatcttcctgataatttttagtgtacatcttgctgttgtttcatcttatgttttgatagaagcatacaaatgagctgatgccatattatatctttatagctattttttgtgctatcttgaaattgagggtccagtgcaactttccttatgactgaaccttgtctttagtacatggaagcacataaatcatatagtagtgtttctggtcatgtttctagagacgtcctacattgataacgcatacatttatgttactgttaaactacctctggtttctgttctttgcaccacccctaaaagagaaggaaactgcttttaaaccctttttggggtttaatacgggaagttgagtatttgatatctctgttgaatgttgtagctcgaaagccttcctgctatgattggtggagtttattcggacgataacaaccttcagcttgaggccacaacacagttccgcaagttgctttctataggtacaaacctcttaaatgttttgggctgttaccttgtgaaaataatagttgaacaaaagttaattactaattacacccattacaatttttaattagagaggagccctccaattgaagaggttattcaatcaggcgttgttcctagattcgtgcagtttcttaccagagaggatttcccccagctacaggtgtgtgccaacacaatattattttcattatctggtaggttcaaataaaaatggatttattactgtggtgcttttgaattaaattttgtagttcgaagctgcgtgggcacttacaaacattgcatctggcacttcggagaatacaaaggttgtcattgatcatggggctgtaccaatatttgtgaagcttcttggttcttctagtgatgatgttcgtgagcaggtatccgatcatctcctgacttgtctttgtgctttctgtacttccttgatattattttgattgagtgcattgttggagttattaatgtttttacctcttgaaaaaaaattgaaggctgtatgggcattgggcaatgttgctggtgattcccctaagtgccgtgaccttgttcttgccaatggtgcattgctgcctctgctagcacagttgaatgagcacactaaactctctatgctaaggaatgcaacttggactttatcgaacttctgcagaggaaaaccacagccatcatttgaacaggtttggacaaagtttctttgtactctactgtagttagtagggaaagacagttccttaacatgatgtttattgggtgtcaatagactaggcctgctcttccagcactggcacggcttattcactccaatgatgaggaagttctcactgatgcgtgctgggctctttcatatttatctgatggcactaatgacaagatccaagccgtgattgaagctggtgtttgcccccgtcttgtggagcttctcctgtaagtgacaacattgcattacaattttctgccattttcctttagattcttatttgctactctcattataccctttttaatattttatactgtttacaacatgaagccatccatcaccttcagtgcttatacccgcactacgaactgttggaaatattgtaactggagatgacgcgcaaactcaggtattgtgttgattttttgcatctctatatctgacagattttgtatcaattgatatcaactactaagactcatgaatcttgagcttaatatttagaggttgtgttcatcacagttgatattaaattttaggcttgtggaaaatgtttaatatcatagttgatattaaatctagaggttgcatctaatgtggaaaatgttttgtaaaaactattgaaaaataaatgcacatatttgtcatatgtagatgcacatgtttggcaatttgacctactctttacgcccatcaattaaagatttgtctagtttgtgtcacttaaatacttgagatttgtcctctcacagtatcacttagatctatactctatactactttaaaaggtagtagtggtggtggcagtgaatctgccagcactacctctatcactaatgtatgggccctccatgttgggtgaaacccacatgctcaataaatttaaacgtcctccaaaagataatgtccatattgtttggtaagatctacagggaaggaatccataaaggggttgaaaagagccagtctacatattcttcaaatgatgagcatgcagtgtgcacagcatgatttttttttcattgcaacgcataggcaatttgctagtttgcctaaaaattcattgcctgctttgcagcaagtcatagctgctgtacttgtttcagccaaacttggcttttcttattgtattcttattattgtttgagttttaggagagctgagaatttttctgctatgctaaagctggtgttcatataatatgaaatatctactgtagatcacagcatgaaagttaattgcttttttgacatccatattgccatctcatcatctccccccaccccctttttttgtacagtgcatcattgatcatcaagccctcccttgcctcctaagtctgttgacacaaaatctcaagaaaagcatcaagaaagaggcttgctggactatttcaaatattactgctggtaacaaagatcagatacaggtatgtgtacagacttaaaaaatactgacatgctcatcattctactaagtcatgaaggatttgtgtagtgcatacttagtagttttaattgtgttccctctccaaaccataaacaccacacaatgttcatattccatgcaagctttcaggtcaacttttcctttcatgatgattacacttgttagcagactcatccttgttgtacaaagactgagaacatggtagtaacttaaagaaacccttatgtatgatttgctcgcaaagtatcaatggcagcctaatcactagtccaaatgttgctgctgggacctggggtcctagaaggggaggggatgggagcatatggatgattgattcatttacacggttgggaacttgtgcacttagctgcaatctttcgtccttaacctataatagcctatagctatagatcttgttgattgtgcatgttctgaaatgttcaacaatttgctgcattgactaccataggctgtcataaatgctggcattattggtcctctggtgaatctgctccaaacggcagagttcgatatcaagaaggaagctgcatgggccatctcaaatgctacttcaggcggtagccatgatcaaatcaagtatgtttactcaccttgcaccagttctattcagctttatgaaatgcaaatataaccgctttcatagtttcatgttaagcttcatttggatttgatattgtatggtttgttaaaataggtatctggtgagcgagggctgcatcaagccattgtgtgatcttcttatttgcccagatataagaatcgttacagtttgtttggagggtctcgagaacattcttaaagtaggagaaaccgacaagaccttggctgcaggtgatgtgaatgtcttttcccagatgattgatgaagcggaaggtttggaaaagattgagaacttgcagagccatgataacaatgagatctatgaaaaagctgtgaagattcttgaggcctactggatggatgaggaggatgatacaatgggagctaccacggtagctgctccccaaggtgctacttttgactttggccaaggcggtggtgctgctcaattcaaataaatacaggtactctcaatcctcaacatggtttgtgtttattgtgggttgtttccactcttaccataaatcctttctgttctggtatcattcacttgaggcaacaagtaatgatttcaaatttcctcgacagatttgaccagaggtggtaaattgtggccactgtgcggggtactgaattgttctggtggttcaataagctaaaggtttcaagggggccagtgttacattatttccctactggtcacggatatcatatcactacgtctgtaacagggattaaagctttgaagtctgcgtttttcatcaggctactactatgtagtgtttgcatcttttagtatggtgtgcaacattttgctacttgtatagcctgagtcgaagcttctgcaaagccgtcacaacacattcctgaacccgtctggttttgagtttcgtcaggagtgtgctgagttgcagtttgtccggacatgtaaaccattatgccttttaaattatatattgctagggtttgttcggttc</dnaseqindica> |
| External Link(s) |