Difference between revisions of "Os10g0505700"
(→References) |
Yilutongxing (talk | contribs) (→Expression) |
||
| Line 7: | Line 7: | ||
===Expression=== | ===Expression=== | ||
| − | + | Wild-type rice nsLTP2 was cloned into the thioredoxin | |
| + | fusion vector (pET32a).This wild-type plasmid was | ||
| + | used as a template for constructing the mutant proteins, | ||
| + | L8A, I15A, F36A, F39A, Y45A, Y48A, and V49A. All | ||
| + | mutants were constructed with the QuickChange | ||
| + | PCR-mediated site-directed mutagenesis kit (Stratagene). | ||
| + | Primers for each mutant were purchased from MDBio | ||
| + | (Taiwan), and mutant construction was based on the | ||
| + | procedure reported previously. | ||
| + | |||
| + | Wild-type rice nsLTP2 and its mutants were expressed | ||
| + | in the pET32a vector transformed into E. coli BL21 | ||
| + | (DE3) (Novagene).E. coli cells were incubated in LB | ||
| + | medium containing ampicillin at 37 8 C. Isopropyl- b -D- | ||
| + | thiogalactopyranoside (1 mM) was used to induce pro- | ||
| + | tein expression. The expressed fusion protein was isolated | ||
| + | by a Co-NTA affinity column (BD Biosciences, San Jones, | ||
| + | USA) with the elution buffer (50 mM Tris pH 7.5, | ||
| + | 150 mM NaCl, and 50 mM imidazole). The purified fusion | ||
| + | protein was then cleaved with Factor Xa and purified by | ||
| + | reverse-phase high-performance liquid chromatogra- | ||
| + | phy.The final product was verified by analysis on a | ||
| + | Micro Mass Quattro Ultima mass spectrometer with elec- | ||
| + | tron spray ionization (Table I). All recombinant proteins | ||
| + | had high purity ( > 98%), and molecular weights of pro- | ||
| + | teins observed in mass spectra corresponded well with | ||
| + | the theoretical molecular masses deduced from the pro- | ||
| + | tein sequence. All mutants were expressed in the oxidized | ||
| + | form, in which the eight cysteine residues were linked in | ||
| + | four intermolecular disulfide bonds. | ||
===Evolution=== | ===Evolution=== | ||
Revision as of 03:54, 7 June 2014
Please input one-sentence summary here.
Contents
Annotated Information
Function
Lipid transfer proteins (LTPs) are typically present in the majority of prokaryotic and eukaryotic cells [1] such as bacteria, yeasts, plants and animals [2]. Plant nsLTPs are basic (with pI 8–10) disulfide-rich proteins divided into two subfamilies; nsLTP1 (~9 kDa) and nsLTP2 (~7 kDa) [3] and [4]. While both proteins have comparable lipid transfer activity, greater stability of nsLTP2 has been proved [5]. Due to their main biological activity, lipid transportation across bio-membranes in vitro, nsLTPs have received much attention from pharmaceutical viewpoints. Owing to the ability of LTPs in promoting the movement of lipids other than phospholipids, clarification of the underlying mechanisms and specificities would be essential to development of LTP-mediated transport systems and controlled release of low molecular weight drugs. A potential role of LTPs in designing efficient drug delivery systems has been suggested [6] and [7]. Their actual biological roles are diverse, including participating in developmental processes and pathogen resistance [8]. Of the two main isoforms, plant nsLTP1 has been studied more frequently and its structure, function and binding properties are well characterized [9] and [10]. Computational studies on nsLTP1 revealed that the insertion of various lipids into the cavity does not necessarily induce significant structural changes with an exception of a prostaglandin. Proteins from various species also varied in terms of molecular surfaces and electrostatic potentials as well as the ability to bind negatively charged lipids. The lack of specificity of ligand binding is suggested to result from nonspecific character of van der Waals interactions [10]. Rice nsLTP2, builds of 69 amino acids and has smaller size, higher structural stability, different disulfide bond pattern and less than 30% sequence similarity with nsLTP1 [11]. Samuel et. al. have performed molecular docking of stearic acid into both LTP1 and LTP2 cavity, demonstrating marked differences between the two isoforms in accommodation of the fatty acid chain in the protein active site. These authors have also reported that there is a tunnel-like hydrophobic cavity running through the whole molecule of rice LTP2 [11]. Despite established beneficial capacities of nsLTP2 to drug delivery [12] and [13] it has not been identified how the plasticity of the protein cavity may help in fitting of diverse molecular shapes and sizes.
Expression
Wild-type rice nsLTP2 was cloned into the thioredoxin fusion vector (pET32a).This wild-type plasmid was used as a template for constructing the mutant proteins, L8A, I15A, F36A, F39A, Y45A, Y48A, and V49A. All mutants were constructed with the QuickChange PCR-mediated site-directed mutagenesis kit (Stratagene). Primers for each mutant were purchased from MDBio (Taiwan), and mutant construction was based on the procedure reported previously.
Wild-type rice nsLTP2 and its mutants were expressed in the pET32a vector transformed into E. coli BL21 (DE3) (Novagene).E. coli cells were incubated in LB medium containing ampicillin at 37 8 C. Isopropyl- b -D- thiogalactopyranoside (1 mM) was used to induce pro- tein expression. The expressed fusion protein was isolated by a Co-NTA affinity column (BD Biosciences, San Jones, USA) with the elution buffer (50 mM Tris pH 7.5, 150 mM NaCl, and 50 mM imidazole). The purified fusion protein was then cleaved with Factor Xa and purified by reverse-phase high-performance liquid chromatogra- phy.The final product was verified by analysis on a Micro Mass Quattro Ultima mass spectrometer with elec- tron spray ionization (Table I). All recombinant proteins had high purity ( > 98%), and molecular weights of pro- teins observed in mass spectra corresponded well with the theoretical molecular masses deduced from the pro- tein sequence. All mutants were expressed in the oxidized form, in which the eight cysteine residues were linked in four intermolecular disulfide bonds.
Evolution
Structure of Rice nsLTP2
The rice nsLTP2 is a predominantly α-helical protein consisting of three prominent helices within the N-terminal 40 amino acids. The well conserved cysteine residues form four disulfide bonds to stabilize the three-dimensional fold of the protein. The C-terminal amino acid residues, Lys41–His69, constitute a less structured region of the molecule with a high density of positively charged residues. The r.m.s.d. values for the backbone and all heavy atoms were 1.09 ± 0.20 and 1.54 ± 0.25 Å, respectively. The first 40 amino acids (Ala1–Ala40), constituting the rigid portion of the molecule, have r.m.s.d. values of 0.65 ± 0.1 Å for the backbone and 0.95 ± 0.15 Å for all heavy atoms. Superposition of the 15 NMR structures are shown as a stereo representation in Fig. 3 A. Three helices of rice nsLTP2 positioned at Cys3–Ala16, Thr22–Ala31, and Gln33-Ala40 are colored green,red, and purple, respectively. Helices II and III are connected by a 90° turn to form a very rigid and unique structural motif. The curved helix I accommodates two disulfide bonds (Cys3–Cys35 and Cys11–Cys25). The flexible portion of the polypeptide contains two single-turn helices at positions Tyr45–Tyr48 and Ala54–Val58. A series of hydrophobic residues distributed throughout the nsLTP2 sequence combine to form a hydrophobic cavity. A continuous stretch of hydrophobic residues, Cys61–Ile65, near the C terminus forms a flexible cap over the hydrophobic cavity. The C-terminal region also contains two cysteines bridged to the rigid portion of the molecule (Cys26–Cys61 and Cys37–Cys68). These two disulfide bonds help to maintain the correct orientation of the hydrophobic cap. The final energy-minimized average structure of rice nsLTP2 is shown in Fig.3 B. A ProCheck analysis of the three-dimensional structure revealed that only Ser59 and Ser60 are in the disallowed region, corresponding to 3.6% of the residues in the protein (19). These residues constitute a portion in the flexible C terminus that makes a very sharp turn to cover the hydrophobic cavity.
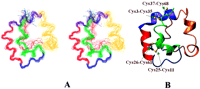
Figure 1, stereo representation of the 15 best superimposed NMR structures of rice nsLTP2 (only the backbone atoms are shown for clarity). B, solution structure of rice nsLTP2. Disulfide bonds involved in the three-dimensional structure are shown in ball-and-stick representation. Helix I (green) and helix II (red) are connected through a loop (Gly17–Pro21). Helix II and helix III (purple) are joined by a sharp 90° turn. The region containing two single turn helices and the C terminus are shown inorange and brown, respectively.
Labs working on this gene
Please input related labs here.
References
[1] R.C. Crain, D.B. Zilversmit, Two nonspecific phospholipid exchange proteins from beef liver. 1. Purification and characterization, Biochemistry 19 (1980) 1433–1439. [2] D.G. Rueckert, K. Schmidt, Lipid transfer proteins, Chem. Phys. Lipids 56 (1990) 1–20. [3] J.C. Kader, Lipid-transfer proteins: a puzzling family of plant proteins, Trends Plant Sci. 2 (1997) 66–70. [4] K.F. Lin, Y.N. Liu, S.T.D. Hsu, D. Samuel, C.S. Cheng, A.M.J.J. Bonvin, P.C. Lyu, Characterization and structural analyses of nonspecific lipid transfer protein 1 from mung bean, Biochemistry 44 (2005) 5703–5712. [5] J.P. Douliez, C. Pato, H. Rabesona, D. Mollé, D. Marion, Disulfide bond assign- ment, lipid transfer activity and secondary structure of a 7 kDa plant lipid transfer protein, LTP2, Eur. J. Biochem. 268 (2001) 1400–1403. [6] C. Pato, M.L. Borgne, G. Le Baut, P. Le Pape, D. Marion, J.P. Douliez, Potential application of plant lipid transfer proteins for drug delivery, Biochem. Pharmacol. 62 (2001) 555–560. [7] Q. Wang, Y. Du, L. Fan, Properties of chitosan/poly (vinyl alcohol) films for drug controlled release, J. Appl. Polym. Sci. 96 (2005) 808–813. [8] A. Carvalho de Oliveira, V. Moreira Gomes, Role of plant lipid transfer proteins in plant cell physiology—a concise review, Peptides 28 (2007) 1144–1153. [9] C.S. Cheng, D. Samuel, Y.J. Liu, J.C. Shyu, S.M. Lai, K.F. Lin, P.C. Lyu, Binding mechanism of nonspecific lipid transfer proteins and their role in plant defense, Biochemistry 43 (2004) 13628–13636. [10] L.F. Pacios, C. Gomez-Casado, L. Tordesillas, A. Palacin, R. Sanchez-Monge, A. Diaz-Perales, Computational study of ligand binding in Lipid Transfer Proteins: Structures, interfaces, and free energies of protein-lipid complexes, J. Comput. Chem. 33 (2012) 1831–1844. [11] D. Samuel, Y.J. Liu, C.S. Cheng, P.C. Lyu, Solution structure of plant nonspecific lipid transfer protein-2 from rice (Oryza sativa), J. Biol. Chem. 277 (2002) 35267–35273. [12] J.P. Douliez, T. Michon, K. Elmorjani, D. Marian, Structure, biological and technological functions of lipid transfer proteins and indolines, the major lipid binding proteins from cereal kernels, J. Cereal Sci. 32 (2000) 1–20. [13] C. Chenga, M. Chenb, Y. Liua, L. Huanga, K. Lina, P. Lyu, Evaluation of plant non- specific lipid-transfer proteins for potential application in drug delivery, Enzyme Microb. Technol. 35 (2004) 532–539.
Structured Information
| Gene Name |
Os10g0505700 |
|---|---|
| Description |
Similar to Nonspecific lipid-transfer protein 2 (nsLTP2) (7 kDa lipid transfer protein) |
| Version |
NM_001071543.1 GI:115482829 GeneID:4349061 |
| Length |
523 bp |
| Definition |
Oryza sativa Japonica Group Os10g0505700, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 10:19790378..19790900 |
| Sequence Coding Region |
19790543..19790818 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008403:19790378..19790900 source=RiceChromosome10 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008403:19790378..19790900 source=RiceChromosome10 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>atggcgaagtgggcggcgatcatggcggtgctgctgctggcggcggcgtgggcgccggcgccggcgacggcgcagtgcaacgccgggcagctggcgatctgcgcgggcgcgatcatcggcgggtcgacgccgtcggcgtcgtgctgctccaacctgcgcgcgcagagggggtgcttctgccagtacgcgcgcaacccggcgtacgcctcctacatcaacagcgccaacgcccgcaagaccctcacctcctgcggcatcgccatcccccgctgctag</cdnaseq> |
| Protein Sequence |
<aaseq>MAKWAAIMAVLLLAAAWAPAPATAQCNAGQLAICAGAIIGGSTP SASCCSNLRAQRGCFCQYARNPAYASYINSANARKTLTSCGIAIPRC</aaseq> |
| Gene Sequence |
<dnaseqindica>83..358#agcagccagcagccaactgcatcgatatcgatcgatcactcgaccgaccatctcacaagcaaagcaaaagctcgtggcaacaatggcgaagtgggcggcgatcatggcggtgctgctgctggcggcggcgtgggcgccggcgccggcgacggcgcagtgcaacgccgggcagctggcgatctgcgcgggcgcgatcatcggcgggtcgacgccgtcggcgtcgtgctgctccaacctgcgcgcgcagagggggtgcttctgccagtacgcgcgcaacccggcgtacgcctcctacatcaacagcgccaacgcccgcaagaccctcacctcctgcggcatcgccatcccccgctgctaggcacgctcgatctcccgccgccgcgcgccgccgctcgccggcgccggcgccggccatggtgcgtggcaaatatatatatatatatatatatatatatatatatatatatatatactgtgtgtacgtgcgcttgaataaaggacgtgagttaatttgatcggtgtc</dnaseqindica> |
| External Link(s) |