Difference between revisions of "Os11g0490600"
(→Expression) |
(→Mutation) |
||
| Line 11: | Line 11: | ||
== Mutation== | == Mutation== | ||
| − | Mutations were also identified in the la1-Shiokari allele with a prostrate phenotype similar to la1-ZF802. | + | Mutations were also identified in the la1-Shiokari allele with a prostrate phenotype similar to la1-ZF802.[[File:Zhangshuling1.jpg]] |
| − | Sequence analysis revealed a single base substitute (TGG→TGA) at the second exon of LOC_Os03g03150 in tad1, which produces a premature stop codon. LOC_Os03g03150 is the TAD1 gene and the premature mutation is responsible for the phenotypes of the tad1 mutant plant. | + | Sequence analysis revealed a single base substitute (TGG→TGA) at the second exon of LOC_Os03g03150 in tad1, which produces a premature stop codon. LOC_Os03g03150 is the TAD1 gene and the premature mutation is responsible for the phenotypes of the tad1 mutant plant.[[File:Zhangshuling2.jpg]] |
| − | TAD1 is involved in regulating the exit of mitosis and it is indeed a functional cell-cycle switch protein homolo¬gous to Cdh1. | + | TAD1 is involved in regulating the exit of mitosis and it is indeed a functional cell-cycle switch protein homolo¬gous to Cdh1.[[File:Zhangshuling3.jpg]] |
| − | Mutation in LA1 results in a significant increase in PAT and thus impairs the IAA differential distribution in la1-ZF802, which ultimately leads to a reduced gravitropic | + | Mutation in LA1 results in a significant increase in PAT and thus impairs the IAA differential distribution in la1-ZF802, which ultimately leads to a reduced gravitropic response in the mutant shoots.[[File:Zhangshuling4.jpg]] |
| − | response in the mutant shoots. | ||
== Expression== | == Expression== | ||
Revision as of 03:08, 9 June 2014
Please input one-sentence summary here.
Contents
Annotated Information
Function
LAZY1 (LA1) gene regulates shoot gravitropism by which the rice tiller angle is controlled. LA1, a novel grass-specific gene, is temporally and spatially expressed, and plays a negative role in polar auxin transport (PAT). Loss-of-function of LA1 enhances PAT greatly and thus alters the endogenous IAA distribution in shoots, leading to thereduced gravitropism, and therefore the tiller-spreading phenotype of rice plants. LA1 is an essential regulator of tiller angle of rice, opening a promising way for breeders to develop elite rice cultivars and other cereal crops with optimal plant architecture. LA1ΔN100 is the truncated LA1 with a deletion of amino-acid residues1-100 that contain a predicted transmembrane domain. LA1ΔNLS refers to the LA1 truncated from amino-acid residues 286 to 312, a segment containing a putative nuclear localization signal (NLS) domain. The 996-1571 bp region of the LA1 gene was subcloned into the T-easy vector and used as templates to generate sense and antisense RNA probes. Rice LA1gene is responsible for the tiller-spreading phenotype of mutant plants.
Mutation
Mutations were also identified in the la1-Shiokari allele with a prostrate phenotype similar to la1-ZF802. Sequence analysis revealed a single base substitute (TGG→TGA) at the second exon of LOC_Os03g03150 in tad1, which produces a premature stop codon. LOC_Os03g03150 is the TAD1 gene and the premature mutation is responsible for the phenotypes of the tad1 mutant plant.
Sequence analysis revealed a single base substitute (TGG→TGA) at the second exon of LOC_Os03g03150 in tad1, which produces a premature stop codon. LOC_Os03g03150 is the TAD1 gene and the premature mutation is responsible for the phenotypes of the tad1 mutant plant.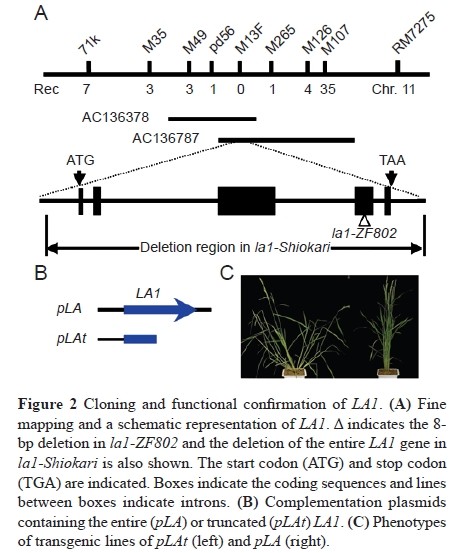 TAD1 is involved in regulating the exit of mitosis and it is indeed a functional cell-cycle switch protein homolo¬gous to Cdh1.
TAD1 is involved in regulating the exit of mitosis and it is indeed a functional cell-cycle switch protein homolo¬gous to Cdh1.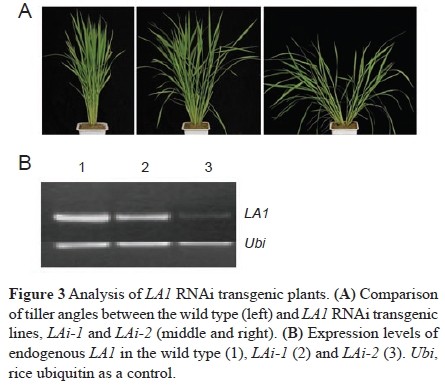 Mutation in LA1 results in a significant increase in PAT and thus impairs the IAA differential distribution in la1-ZF802, which ultimately leads to a reduced gravitropic response in the mutant shoots.
Mutation in LA1 results in a significant increase in PAT and thus impairs the IAA differential distribution in la1-ZF802, which ultimately leads to a reduced gravitropic response in the mutant shoots.
Expression
LA1 is a finely regulated temporally and spatially expressed gene, and that the region of its specific expression may play an important role in controlling the rice tiller angle.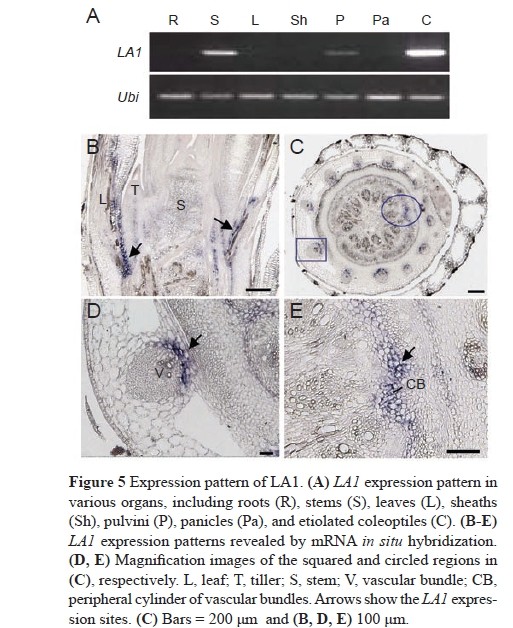


Evolution
LA1 may represent a new type of regulating proteins that shuttle between the plasma membrane and the nucleus. Further investigations of LA1 functions will allow for a better understanding of the mechanism underlying the monocotyledonous shoot gravitropism.
Labs working on this gene
1、State Key Laboratory of Plant Genomics and National Center for Plant Gene Research, Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, Beijing 100101, China; 2、State Key Laboratory of Rice Biology, China National Rice Research Institute, Chinese Academy of Agricultural Sciences, Hangzhou 310006, China
References
1. Morita MT, Tasaka M. Gravity sensing and signaling. Curr Opin Plant Biol 2004; 7:712-718. 2. Perbal G, Driss-Ecole D. Mechanotransduction in gravisensing cells. Trends Plant Sci 2003; 8:498-504. 3.Fukaki H, Wysocka-Diller J, Kato T, Fujisawa H, Benfey PN Tasaka M. Genetic evidence that the endodermis is essential for shoot gravitropism in Arabidopsis thaliana. Plant J 1998; 14:425-430. 4. Tsugeki R, Olson ML, Fedoroff NV. Transposon tagging and the study of root development in Arabidopsis. Gravit Space Biol Bull 1998; 11:79-87. 5. Moore I. Gravitropism: lateral thinking in auxin transport. CurrBiol 2002; 12:452-454. 6. Kim SK, Chang SC, Lee EJ, et al. Involvement of brassinosteroids in the gravitropic response of primary root of maize. Plant Physiol 2000; 123:997-1004. 7. Gutjahr C, Riemann M, Muller A, Duchting P, Weiler EW, Nick P. Cholodny-Went revisited: a role for jasmonate in gravitropism of rice coleoptiles. Planta 2005; 222:575-585. 8. Aloni R, Langhans M, Aloni E, Ullrich CI. Role of cytokinin in the regulation of root gravitropism. Planta 2004; 220:177-182.
Supplementary information
It is linked to the online version of the paper on the Cell Research website.
Structured Information
| Gene Name |
Os11g0490600 |
|---|---|
| Description |
Conserved hypothetical protein |
| Version |
NM_001074458.1 GI:115485564 GeneID:4350543 |
| Length |
5524 bp |
| Definition |
Oryza sativa Japonica Group Os11g0490600, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 11:19145382..19150905 |
| Sequence Coding Region |
19145410..19145415,19145782..19145851,19147818..19148686,19150064..19150349,19150566..19150585 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008404:19145382..19150905 source=RiceChromosome11 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008404:19145382..19150905 source=RiceChromosome11 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>atgaagctcttaggttggatgcatcgaaagctacggagtaataatgacgtgttcaaagagttcaacaccggaggaggtggggcctgcaactgcatcaccgggcttgcctcgcctgaccacgacaacgactacttctccggcgacgacgccgcccacgcctcgccgccggtcaccgccggcgacctcttcaccttcggcggcagcggccttctcaccatcggcacgctaggcatcgccgccgtcgccattcccagcggcggcgacgacgacgactacgacatcgacttcgaggtggacgccaccagcgacgacgacggcggcttcaccgtcgaggacgacgacgccgacgtcggcggcgccgtcacgcccaccttcaccttccccgcggcgacggcggcggaggcggtcgtcgccaccgtggagaaggcagtggccgcggtggaggcgatcgcggagaaggacgacgacaccaccacggaggacgacctgatggtggtgagcgccgagctggagaaggtgctcggcggcgtcgacgtggcgtcggcgcgggtgagcttcgccatgggcggtggcgtcgactgcccgctccagggcttcctgttcggctccccggtgagcgacgtcgagtcgcgcccggagtacctgcaggcgccgcgggactcgtccggctcctgcggcggcggcgggcggcgcacctcgctcggcgagctgttcatgcgcacccgcttcgccgacgagaaggtggcgctcgtcgccgtcgccgagggcgaggacggcgtcgccggcgacgacggcgctgctgctgccggcgtcggcggagacagagcggggaaaggcggcggctacaagacgatgaagaagaggaaggtgaaggacgagaaaggcggcggcggcgccgccggcggtggaatgccggcgacggtgacgaagagcaagtttcagaagatccttcaaatcttccacaggaaagtctaccccgagaacacactcctcacaaggaatctgaccaagaagagccgcaaccgcggcgccaccgataatggcggtggcgccgtggccaccggagaccccgacgggcctctggcctcgccggtgctccggtgccggaaggaccatcccatgaggggcttcggctgctgcaccaatggcgccttcggtgcatcgtcaccgggaggcaacgccgagatgaacggcaacaagagcggccactggatcaagactgatgccgactacttggtgctggaattataa</cdnaseq> |
| Protein Sequence |
<aaseq>MKLLGWMHRKLRSNNDVFKEFNTGGGGACNCITGLASPDHDNDY FSGDDAAHASPPVTAGDLFTFGGSGLLTIGTLGIAAVAIPSGGDDDDYDIDFEVDATS DDDGGFTVEDDDADVGGAVTPTFTFPAATAAEAVVATVEKAVAAVEAIAEKDDDTTTE DDLMVVSAELEKVLGGVDVASARVSFAMGGGVDCPLQGFLFGSPVSDVESRPEYLQAP RDSSGSCGGGGRRTSLGELFMRTRFADEKVALVAVAEGEDGVAGDDGAAAAGVGGDRA GKGGGYKTMKKRKVKDEKGGGGAAGGGMPATVTKSKFQKILQIFHRKVYPENTLLTRN LTKKSRNRGATDNGGGAVATGDPDGPLASPVLRCRKDHPMRGFGCCTNGAFGASSPGG NAEMNGNKSGHWIKTDADYLVLEL</aaseq> |
| Gene Sequence |
<dnaseqindica>29..34#401..470#2437..3305#4683..4968#5185..5204#atcttagcacgctaaaccggcttccaagatgaaggttagtgtttcatctattgaacatatccatgcttattagtttcgtaatgtagatactaagtttatgtatatgtagttctcttttggaacttgagttgttaatttctttttgtaaatattttttgtttggtttggttacgttgctggcctagctgatcatgctttctgggttgaataaatgtagctacaactattaatcctttattttgtttttccatcattgccgttgtcatcatctttcattgtcatcatcatcttctccttttgacgatcttcatcagatcttgatcaaaagtaatagaccatgcattttcatgattgatgaagttcataaaaatttctgatcctcatatgtgtatatgatcagctcttaggttggatgcatcgaaagctacggagtaataatgacgtgttcaaagagttcaacaccggaggaggtatgcacgcacaacttaattaatctattttttgtcctttttttcttatgtttaattgatgaagtataattgttttgaaaaatacagtagctcgtaacacaaacgcactcacctctagtactctctccagtgtttttatatattaaactaacgtcatataaaaaattgaagttagtatatatgtataatagttcatcggtacatgcagaaaaaaaaaagagtttgtctctggttttagatctgtaagcatgcccacaataatgagaaaaccactgcaaaattgttggccggccggtcccacatgtcattgaccagattagccgtctgaagtgctgaactgtatggaacactagcaatcatcaatgggagttttgcactcttgcgcttggtcacattggagatggcacatctatctataatatcacacatgccacttttcttccagcctcaatatctatagttccagcactagctagctagctagaccaactattgcatagcaagaaccctatattatcattttgcagataacattttcttttccatgtagtaacattgccatggaaagatcacgagggcatgtctatcttttgtgaaactctctctctctccttttgaaggcttcctttgagagttttctccttgttggactgttgctcttaggctcagcatgttaatatcagagttttcgcccgtttttcataaatcaattgtgccgttgtaagatttttgcgtccagttttccttataaactggacctctctgcctgtttcctcgagaaaaattgtccaagagcattctcgacgattcaacaactagaattcaacaaataaagatttttatatatggtgcatatggtacggtgagatattgtatttttagtgtgagataccaaaagagctacaacaaattaatacgataaaataagggaaatttcatggactaactagttagaaacatattttgagtaaatgaaaattttattttattaccttggtttacaaaagacttataataagtgctgtcaaacatctaaaatgttaaattcttaatagacgaaaagtattagaatttaaaacgtgaaaattatatactttttgatgaaaaatagtaggaatgtgaaaattatattagtagactttttttgatgaaaactctttcatatagttatatgttaattttttataactatatagtttgagaaagtaatcatataacttttgcattaaaaaatgtgtcaatgtccaaaacattatcttaattaacccggtccttctcctcccaaaggaacacataaaaaaggacaaaaaccaatacatcttgaaaaacttacatagaaaaagcttaatcatattttatatcaccctgatcccgttgcaacgcacatgaatgtaactatttctttttaaaatggtgttgtacttcataaatttgaaacaaaaaaaatacatataatgtaacccttttaatttctgaaacacatcaaaacttttctttctcaaaaaaaagaaaagaaaacacatgaaagcctgcataattttgcatgcaaatgatcgtactccttcctttttaaggttacaagactttcttacattacccaaatttatatagataataaatctagacacaaatatatgtgatttattaatatgtatatgaatgtgaacaatgccaaaaagtcttataatataaaatggagaaagtatttaatactccctcagtttctaaatatttgacaccattgattttttaaacatgtttaatcattcgtcttattcaaaaattttaagtaattattaattattttcctatcatttgattcattgttaaatatacttttatgtatacatataattttacgtatttcacaaaagtttttgataagacggacggtcaaacatgtgctaaaaagtcaactgtgtcagatatttagaaacggagggagtaattcactgtgtgactgcaggtggggcctgcaactgcatcaccgggcttgcctcgcctgaccacgacaacgactacttctccggcgacgacgccgcccacgcctcgccgccggtcaccgccggcgacctcttcaccttcggcggcagcggccttctcaccatcggcacgctaggcatcgccgccgtcgccattcccagcggcggcgacgacgacgactacgacatcgacttcgaggtggacgccaccagcgacgacgacggcggcttcaccgtcgaggacgacgacgccgacgtcggcggcgccgtcacgcccaccttcaccttccccgcggcgacggcggcggaggcggtcgtcgccaccgtggagaaggcagtggccgcggtggaggcgatcgcggagaaggacgacgacaccaccacggaggacgacctgatggtggtgagcgccgagctggagaaggtgctcggcggcgtcgacgtggcgtcggcgcgggtgagcttcgccatgggcggtggcgtcgactgcccgctccagggcttcctgttcggctccccggtgagcgacgtcgagtcgcgcccggagtacctgcaggcgccgcgggactcgtccggctcctgcggcggcggcgggcggcgcacctcgctcggcgagctgttcatgcgcacccgcttcgccgacgagaaggtggcgctcgtcgccgtcgccgagggcgaggacggcgtcgccggcgacgacggcgctgctgctgccggcgtcggcggagacagagcggggaaaggcggcggctacaagacgatgaagaagaggaaggtgaaggacgagaaaggcggcggcggcgccgccggcggtggaatgccggcgacggtgacgaagagcaagtttcagaaggtaacttttttttttgttgttgctagcattttaatttgctttgaagaaaactataaaatatctgcaattttggctgactatttcatgaggtttccttggatatgatcatttgattaattagttcgttcgattgctatagttttagttcatttttctttaattaatttgtaataaaactcaagagttattgtgagttttttttttattatttttgcttcagtacgtgcattttctactagtgaaacttcaatattaaccaggccagtcgacatttcttaattgtgatttgatttaagggtcaagattagaactatagattgaatatgcaaagtttcagctcggacgacatttgaacatttgactaattatatatttccgagttcaattgtttgccggtaaaattctgaacttgcagttggaacattcaagattcaactctgattcacatgatctcgaactcaatatctccaactttataaaaagacaaaaagattactgctcaacggtgagtttaatgaaggaatgctcatcacagcaaatccataattctatataactctctgtaatatgtaatggtaagaaacagatattcatgattaaatcatttttccccttcagaaagaaatactgatttttcgtggtacatccaaaattactccaaagtttacactgaatttcacgctgttcttggtgtttaagattaattttaggtagatattttctagtcccgcaacaaagctgacaatgaaaaccccaaaactaatcctaaatgaaattctaaaattaagaaaaacagctttagattataaattttaatagtaagtccaaggaaatagccagcaatgttaataagattgctgataagaatatataacagtttggagccaatccatttcacatgtgtatcctgctctgggacataagtagacacaaatcccgagttttgactatcatctcataatcggaagatctttcactgtgtcgtcagattcgagttaagtttctgtagaccaacttggtgtgtcaatggtgttgttaggagtggggatactgggggcagtgagttgacatgcgatttgtttgcaatctcctgtctaatgctatctttatggatcatttcatttgtcatttttcgtttgccccataaacaacaagggattcttgatttgatatattatacatccttctaggtgctctcttgcatttcttttccttggagagatgctccaaatatccctcaaaagaaattctagattccttttgctactacttggtacttacatttaattttgttacaacgaatgaaagaaactgagtttcagaaagtgatcaaaagttgacaatgttctgaaatgaggaaacatctgctcattgcagatccttcaaatcttccacaggaaagtctaccccgagaacacactcctcacaaggaatctgaccaagaagagccgcaaccgcggcgccaccgataatggcggtggcgccgtggccaccggagaccccgacgggcctctggcctcgccggtgctccggtgccggaaggaccatcccatgaggggcttcggctgctgcaccaatggcgccttcggtgcatcgtcaccgggaggcaacgccgagatgaacggcaacaagagcggccactggatcaagactgatgccgactgtgagtagcactgcacaccttggtgtctccatccatcttcaatggatctatctttgcaatcatgcattcagtcttgacagctacatctttccatactatttttggggaatcttccaagagctatccatcactagtttctgagttactgtgtgctgatgcacacactgcaaacattgtgctttttcactgaaaccctgtctctctgatctgatgcagacttggtgctggaattataatggggaaaagaagaggagctatttgtttcatcaagaattaaagcttgaatagtgagcatctcatatatatgaatatgtgcttgctgttacaatataatctctatctgtttggtatagaagggcttgaatgtgctgtatatgtcactatatattgggggggagaggattactatgagatcaactcataagtgtgtggtgtttatatacattgctctgtgaatgtatgacagagatcagaacttaatgtattgtatgctttcttgatgtgattggtgtttatgagcctagctgatggcaatattgtgtgtgggtccacccct</dnaseqindica> |
| External Link(s) |