Difference between revisions of "Os07g0211500"
| Line 1: | Line 1: | ||
| − | Rc is a domestication-related gene required for red pericarp in rice. | + | Rc is a domestication-related gene required for red pericarp in rice (Oryza sativa) and it encodes a basic helix-loop-helix (bHLH) protein that was fine-mapped to an 18.5-kb region on rice chromosome 7 using a cross between Oryza rufipogon (red pericarp) and O. sativa cv Jefferson (white pericarp). |
===characteristic=== | ===characteristic=== | ||
| Line 18: | Line 18: | ||
===Expression=== | ===Expression=== | ||
| − | + | RT-PCR experiments confirmed that the Rc gene was expressed in both red- and white-grained rice but that a shortened transcript was present in white varieties. | |
===Nucleotide Polymorphisms=== | ===Nucleotide Polymorphisms=== | ||
| − | + | Phylogenetic analysis, supported by comparative mapping in rice and maize (Zea mays), showed that Rc, a positive regulator of proanthocyanidin, is orthologous with INTENSIFIER1, a negative regulator of anthocyanin production in maize, and is not in the same clade as rice bHLH anthocyanin regulators.[[File:mapping of Rc.jpg]] | |
===Allele Distribution=== | ===Allele Distribution=== | ||
| − | + | Sequencing of the alleles from both mapping parents as well as from two independent genetic stocks of Rc revealed that the dominant red allele differed from the recessive white allele by a 14-bp deletion within exon 6 that knocked out the bHLH domain of the protein. | |
===Haplotype Diversity=== | ===Haplotype Diversity=== | ||
Revision as of 08:18, 10 June 2014
Rc is a domestication-related gene required for red pericarp in rice (Oryza sativa) and it encodes a basic helix-loop-helix (bHLH) protein that was fine-mapped to an 18.5-kb region on rice chromosome 7 using a cross between Oryza rufipogon (red pericarp) and O. sativa cv Jefferson (white pericarp).
Contents
characteristic
Mutation

 Grain color has been the target of selection in domesticated Asian rice (as well as other domesticated grains), and the vast majority of O. sativa varieties have white (nonpigmented) grains, compared to the dark red grains that characterize the wild relative and other wild species. The genetic basis of white grains in O. sativa has been pinpointed to loss-of-function mutations in the Rc gene, a regulatory protein in the proanthocyanidin synthesis pathway; these loss-of-function mutations prevent the development of a pigmented pericarp layer. A survey of 337 white pericarp rice cultivars showed that the majority (>97%) have a 14-bp deletion in exon 7 of the gene, resulting in a premature stop codon and a non-functional allele referred to as rc.
Grain color has been the target of selection in domesticated Asian rice (as well as other domesticated grains), and the vast majority of O. sativa varieties have white (nonpigmented) grains, compared to the dark red grains that characterize the wild relative and other wild species. The genetic basis of white grains in O. sativa has been pinpointed to loss-of-function mutations in the Rc gene, a regulatory protein in the proanthocyanidin synthesis pathway; these loss-of-function mutations prevent the development of a pigmented pericarp layer. A survey of 337 white pericarp rice cultivars showed that the majority (>97%) have a 14-bp deletion in exon 7 of the gene, resulting in a premature stop codon and a non-functional allele referred to as rc.
Our findings for the pericarp color gene, Rc, indicate that remarkably similar genetic mechanisms can be at play. We find that two of the three white pericarp O. glaberrima varieties in our sample harbor a unique point mutation that is predicted to result in a premature stop codon in exon 7 of the Rc gene.
Instead, two samples contained a novel point mutation predicted to result in a premature stop-codon in exon 7, hereafter referred to as allele rc-g1 (Fig. 1). This mutation is 146 bp upstream of the Rc-s point mutation and 201 bp upstream of the initiation of the 14-bp rc deletion. The third white pericarp O. glaberrima contained no identifiable insertions, deletions, or premature stop-codons, nor does it differ from the red-pericarp O. glaberrima sequences for any predicted amino-acid changes. Comparison of this sample to the most closely related red-pericarp sample showed that they were identical for the sequenced region upstream of the start codon (>1.5 kb) and for the first intron; both of these regions are potentially important as cis-regulatory regions in plants.
This study also shows that white pericarps might be achieved by mutations in cis-regulatory regions of Rc or potentially mutations in other genes, a pattern that was not observed in an extensive survey of O. sativa cultivars.
Function
Expression
RT-PCR experiments confirmed that the Rc gene was expressed in both red- and white-grained rice but that a shortened transcript was present in white varieties.
Nucleotide Polymorphisms
Phylogenetic analysis, supported by comparative mapping in rice and maize (Zea mays), showed that Rc, a positive regulator of proanthocyanidin, is orthologous with INTENSIFIER1, a negative regulator of anthocyanin production in maize, and is not in the same clade as rice bHLH anthocyanin regulators.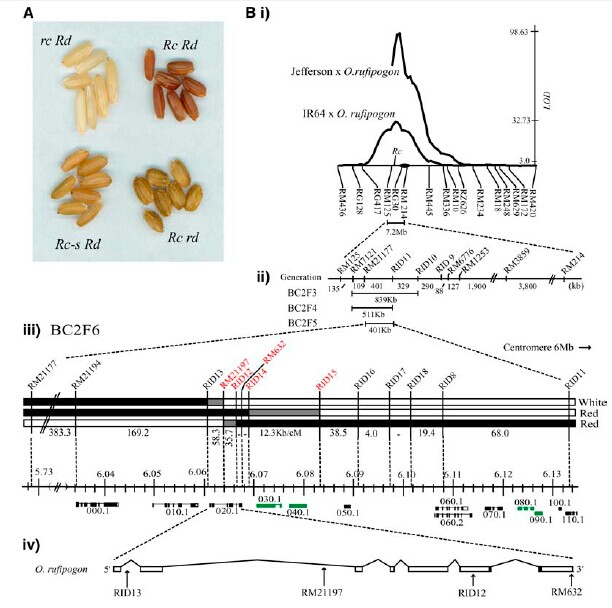
Allele Distribution
Sequencing of the alleles from both mapping parents as well as from two independent genetic stocks of Rc revealed that the dominant red allele differed from the recessive white allele by a 14-bp deletion within exon 6 that knocked out the bHLH domain of the protein.
Haplotype Diversity
Recombination
Evolution
Repeated phenotypic evolution can occur at the interspecific level, where it is manifest as the appearance of the same trait in multiple domesticated species, and at the intraspecific level, where the same trait arises multiple times within a single crop. In rice, the opportunity exists to examine the genetic basis of repeated trait evolution at both the interspecific and intraspecific level.
At the intraspecific level, repeated phenotypic evolution during domestication can be explored in two contexts; one is the repeated evolution of a trait in varieties resulting from a single domestication event, the other is repeated evolution of a trait in varieties that result from multiple, independent domestication events within the same species.
Labs working on this gene
- Department of Biology, Washington University in St. Louis, St. Louis, Missouri, USA
- Plant Science Department, South Dakota State University, Brookings, South Dakota 57007, †Biosciences Research Laboratory, U.S.
- Department of Agriculture–Agricultural Research Service, Fargo, North Dakota 58105, ‡Northern Crop Science Laboratory, U.S.
- Department of Agriculture–Agricultural Research Service, Fargo, North Dakota 58102, §National Institute of Agrobiologica
- Sciences, Tsukuba, Ibaraki 305-8602, Japan
- Agricultural College, Yangzhou University, Yangzhou 225008, China
- Department of Plant Breeding and Genetics, Cornell University, Ithaca, New York 14953-1901
- Department of Plant Biology, Cornell University, Ithaca, New York 14853
- Department of Plant Breeding and Genetics, Cornell University, Ithaca, New York, United States of America
- International Rice Research Institute, Los Ban˜ os, Philippines*
- Indonesian Center for Agricultural Biotechnology and Genetic Resources Research and Development, Bogor, Indonesia
- Department of Agronomy, Chungbuk National University, Chongju, Republic of Korea
- National Institute of Agricultural Biotechnology, Suwon, Republic of Korea
- Department of Biological Statistics and Computational Biology, Cornell University, Ithaca, New York, United States of America
- Plant Genome Research Unit, National Institute of Agrobiological Sciences, 2-1-2 Kannondai, Tsukuba, Ibaraki, 305-8602 Japan
- QTL Genomics Research Center, National Institute of Agrobiological Sciences, 2-1-2 Kannondai, Tsukuba, Ibaraki, 305-8602 Japan
- Genetic Diversity Department, National Institute of Agrobiological Sciences, Tsukuba, Ibaraki 305-8602, Japan
- Department of Biological Science and Technology, Tokyo University of Science, Noda, Chiba 278-8510, Japan
- Research Institute for Biresources, Okayama University, Kurashiki, Okayama 710-0046, Japan
- National Agricultural Research Center for Kyushu Okinawa Region, National Agriculture and Bio-oriented Research Organization, Nishigoshi, Kikuchi, Kumamoto 861-1192, Japan
- National Institute for Basic Biology, Okazaki, Aichi 444-8585, Japan
- Plant Breeding Laboratory, Graduate School of Agriculture, Hokkaido University, Sapporo, Hokkaido 060-8589, Japan
References
- ↑ Xing-You Gu; Michael E. Foley; David P. Horvath; James V. Anderson; Jiuhuan Feng; Lihua Zhang; Chase R. Mowry; Heng Ye; Jeffery C. Suttle; Koh-ichi Kadowaki; Zhongxiang Chen. Association Between Seed Dormancy and Pericarp Color Is Controlled by a Pleiotropic Gene That Regulates Abscisic Acid and Flavonoid Synthesis in Weedy Red Rice;Genetics, 2011, 189(4): 1515-1524.
Cite error: <ref> tag with name "ref2" defined in <references> is not used in prior text.
Cite error: <ref> tag with name "ref3" defined in <references> is not used in prior text.
Cite error: <ref> tag with name "ref4" defined in <references> is not used in prior text.
Cite error: <ref> tag with name "ref5" defined in <references> is not used in prior text.
Cite error: <ref> tag with name "ref6" defined in <references> is not used in prior text.
Structured Information
| Gene Name |
Os07g0211500 |
|---|---|
| Description |
Pericarp Color;red grain color gene |
| Version |
DQ204735.1 GI:78057266 |
| Length |
833bp |
| Definition |
Oryza sativa (japonica cultivar-group) cultivar H75 brown pericarp and seed coat (Rc) gene, complete cds. |
| Source |
Oryza sativa Japonica Group (Japanese rice) ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 7 |
| Sequence Coding Region |
{{{CDS}}} |
| Expression | |
| Genome Context |
{{{GCID}}} |
| Gene Structure |
<gbrowseImage2> name=NC_008397:19856181..19857859 source=RiceChromosome07 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq> 1 atggccggcg gcgaggcgca tgcggcgctg caggcggtgg cgcagagcct ccggtggacc 61 tacagcctcc tctggcagct ctgcccccac caagggagctcg
421 ctggtgtggg gggaggggca ctacaacggc gccgtcaaga cgcggaagtc gacggtgatg
481 cagccgccgc cggcggagga ggaggacgac gccgaccacg cggcgcgcca ccggagccgg
541 cagctgaggg agctctacga ctggctgcag caggccgggg agaactccag cggcggcgtg
601 cagacgtcgt cgacgacggc gagccggcgg ccgggggcgg ctctgtcgcc ggaggacctg
661 acggagacgg agtggttctt cctcatgtcg gcatcctact ccttccctcc cggcatcgggtta
3421 cctggaaggg catttgcaag gagaggccat gtatggctca ctggagcaaa tgaagttgac
3481 agcaaagtat tcctaagagc aattcttgcc aagacagttgtg tgcattcctg ttgtcgatgg cgtcctggaa attggaacta cggaaaaggtggaggaag atatgggcct gattcagtat gcaaggggca
4201 tcttcatgga tcaacatggc atccacatga agcctaccct ctcacagcac tcaacatcca
4261 acccagtcac ccactgtact catcagcatc caatccaggt tcagatgcaa ctaggtatca
4321 ccagccaaac aaagtttgat tattcagatg agctcaatgc agatgaggag aatgatgaca
4381 cagaagaaga gggcatgtca ggttcagaca ctaacaacac tgacactgaa aggaattcag
4441 gccagctgca acttcaaatg caagaccaac tgaacatggt gagcaatgac caccagacaa
4501 taccaaataa tgcagtttcc agtgagctaa tgcagtgtga gatgtcagaa gtggtaagag
4561 atggctgctc aaataatatt ttagaggatg aaatccaaat gctgatggat tgccaaaaca
4621 gtaattgtca gttaaatttg caagggccag atgagccttg tcactcttgg cattttctct
4681 gcgaggagtt acaaaatgat taccagccagctactgaaga tcaagtggca tcacctgaaa atacccatta
4921 cccaaaaaca ctcatgacaa tcctacatta caacacgctg cgacagcaag agatgaacat
4981 caagaactac ttgccagttt cagagaaatc atcattctcc agatggacta ctcctgaagg
5041 aagtgatgac aacaagacca tgatcagtcc aggcaccaca cagagaatgc tcaagagcat
5101 cctgatgatt gttcccagta gtcactgcag ttacagggga gcagaaacac ctgaatcaag
5161 gggcgggaaa ggcgcaagtg gaacgcgaaa agtcggtgcc atccaaggtg atttcagtgc
5221 caaccatgtg ctgaaagaga ggagaagaag agagaagctc aatgagaagt tcataattct
5281 gcgatctttg gtacctttca tgacaaagatggac aaggcgtcga tactaggcga
6001 cacgatcgag tacgtgaagc agctaaggaa ccgcatacaa gagctcgagt cgtcgtcgtc
6061 gtcgtcacga gcagccgccc gggcgccatc ggcggcggcc gccgggaggc ggaggaagag
6121 atccgccgcc gccgccactg ccacggcggc ggaagggatg agcagcagca atggccgcaa
6181 tggcggcgag gcggcggagg tggtgcaggt gtccatcatc gagagcgacg cgctgctgga
6241 gctccggtgc ggttgcggcg gcggcggcgg cggtgtggtg ctgctccggg tgatgcaggc
6301 gatgcaggag ctccagctgg aggtcaccgc cgtccaggcc tcgtgcgccg gtggcgagct
6361 gctcgccgag ctgcgcgcca aggtcgtcgt tatgatcctg atctgcatga aaatgcaaat
6421 gcaaatgcaa atgcagaatt aa</cdnaseq>
|
| Protein Sequence |
<aaseq>MAGGEAHAALQAVAQSLRWTYSLLWQLCPHQGSSLVWGEGHYNG AVKTRKSTVMQPPPAEEEDDADHAARHRSRQLRELYDWLQQAGENSSGGVQTSSTTAS
RRPGAALSPEDLTETEWFFLMSASYSFPPGIGLPGRAFARRGHVWLTGANEVDSKVFL
RAILAKTVVCIPVVDGVLEIGTTEKVEEDMGLIQYARGIFMDQHGIHMKPTLSQHSTS
NPVTHCTHQHPIQVQMQLGITSQTKFDYSDELNADEENDDTEEEGMSGSDTNNTDTER
NSGQLQLQMQDQLNMVSNDHQTIPNNAVSSELMQCEMSEVVRDGCSNNILEDEIQMLM
DCQNSNCQLNLQGPDEPCHSWHFLCEELQNDYQPATEDQVASPENTHYPKTLMTILHY
NTLRQQEMNIKNYLPVSEKSSFSRWTTPEGSDDNKTMISPGTTQRMLKSILMIVPSSH
CSYRGAETPESRGGKGASGTRKVGAIQGDFSANHVLKERRRREKLNEKFIILRSLVPF
MTKMDKASILGDTIEYVKQLRNRIQELESSSSSSRAAARAPSAAAAGRRRKRSAAAAT
ATAAEGMSSSNGRNGGEAAEVVQVSIIESDALLELRCGCGGGGGGVVLLRVMQAMQEL
QLEVTAVQASCAGGELLAELRAKVVVMILICMKMQMQMQMQN"</aaseq>
|
| Gene Sequence |
<dnaseqindica>1321..1386#668..1152#450..531# 1 atggccggcg gcgaggcgca tgcggcgctg caggcggtgg cgcagagcct ccggtggacc 61 tacagcctcc tctggcagct ctgcccccac caagggtacc taccctacct acctacgaca
121 cgatgcacag tgttcatcca tggccggcca tggcggatcg tcgtcgttgt cgatgatcat
181 cgaaggaagc tagaggatat ggctcaatac tttgataata tatatactga tctctccgta
241 caacaaaaat ataaaaattc tagctagtat cgaatgagac atatgctatg ctagtactac
301 gaatctaaaa agatgtacat attttgattc gtattattag gatatatcac gagtttttat
361 attttgagac ggatgtaata attctgaatt tagttgtgat ccatggcatg caggagctcg
421 ctggtgtggg gggaggggca ctacaacggc gccgtcaaga cgcggaagtc gacggtgatg
481 cagccgccgc cggcggagga ggaggacgac gccgaccacg cggcgcgcca ccggagccgg
541 cagctgaggg agctctacga ctggctgcag caggccgggg agaactccag cggcggcgtg
601 cagacgtcgt cgacgacggc gagccggcgg ccgggggcgg ctctgtcgcc ggaggacctg
661 acggagacgg agtggttctt cctcatgtcg gcatcctact ccttccctcc cggcatcggg
721 tatataataa aaaatataga tataaatatt taagcatgca tgcataaatt aaaccacact
781 tcttgttacg tgttcttggc aaaatgatga acaattacca ctaattaatt ggagccagaa
841 accctaaaga tttacccacc tggttaatta atcggtgtgt tgatccacgc atgcatgcat
901 gcagaaaatc aagatcagga tagctccttt tcttttgcag gttaattagc tagatcttca
961 cgtataatta gctagctaga ttttaaaata taatttattc aatttgattt atgattttta
1021 ttttttattt caaatagata caactgtata caaaatttta ttttggtaca tacctccgat
1081 ccaactacat cagaggtaaa aaaaaaatta aaccgttgga attgattaga acaagatcgt
1141 gcggtcaaat tatatcataa ctaacttttc tgattctcta aagcatagag atgtatatat
1201 acatcgtatt attaggctct atatttcctg attaacacta gatgcatata taattttgat
1261 agtcaaaata tacttttgat aggctctaaa gaaaaactta ataacatgta ctccctccat
1321 atacttttga tagtcatatt tcatcttgac acacagatca agtataagta attctactta
1381 tcatccattt aaacacgcta ctagttattc ctcataaaca agcgattcat taatatttac
1441 atttctcgat gcttgtgtag ccaatattgt gtggaagaat ggaatgtcat taagaggata
1501 ggttgttgga ttgaaatatg cctatcaaaa ataaattttt agatttgaaa atatgcctat
1561 caaaagtaga tggagggagt attaattaat gtgaatttcc aatcctactg ttgtgatatt
1621 aggctttgta ccttcttgtc caggaggtat atatatggct cttttaagga tgggagaaaa
1681 tatcatcttt aatacaacta tatatggctt ttgtttgata aatacaactt ttattttgta
1741 tgaatacaaa tatattgata aatatccacc attataatcc taacccatta ggatcatatg
1801 gtgtatattt ttttaactat ttgtttttta taaattaata ttaagagatc acaataaaaa
1861 tatagtatta tgaaagtact cttaacaaca tatccaatga taaaattatt attattacaa
1921 aatatagtgg tcaaattgta tagaattcaa tagcctgatt ttatgacgtc aagtaaatta
1981 aataaagaat gaaggtagtg ctagagtgat caaacaatat ctctcctaaa atatgtccta
2041 taagttttac tccataaatc caagggtcaa aagttgttgg gttatttttt tagataataa
2101 catactaccc cttttcaaaa tgtatgattc tattgacttt ttgcacaaca tttaaccatt
2161 tgtcatatta aaaattagta taaacatcta aaaatataag ttacaattat attttatttg
2221 atgataaaac aactcacaac aaaataaata atatttatat aatctttttg gaataaaacg
2281 aatgatcaaa cattattcaa aaagtcaatg gtatagtacg ttttgaaatt gatagactat
2341 gagagcaaaa ttttgagata acatggaaaa ttatcctctt agacattgca ctgtgtaata
2401 attaataata atgaatgaaa ggctaagact tttcttccac cttatataag tggttgaata
2461 tatagcaatc acatcattac atgattttgt aaccaaccgt ctctatagct ccgatacagt
2521 gctagtttca catcgtaata attaaagagt ataataataa atcgaggtgt acttctcatc
2581 gatgaagtga tgtgccgctt agctaaatta aactcgtatg cgaaaaatca gtatatgtcc
2641 ggttaatttc taagagagag attgagagag aataattgcg cccctccaaa tccccctctt
2701 ggacgttagg gagctatata gacggtattg ctaagtgcga tgtgtacata acgtacctgt
2761 cgtaggaaca tttctcatcc aaattaagta gtaatgcatg gcatgaaatc catttttgta
2821 ttttgcatgg caaagaatga caacaaggaa tacactagct agccctgccc tttttcaatt
2881 taatttaaca tcaaacttag ttattgtatt tcttttgtca gaatagcatg cattgcatac
2941 tctttaaaaa taattaatta gtgtatttta ctagtcttac aaaagtatca agagagacaa
3001 ctaattatag ttgggagaca ccaaacttgt ttttaataat gacaattaaa accctacctc
3061 tacatccaac atagacgtac atagtccgaa ggcgccaaat atttgtacat ttagctacca
3121 gatttcagta cgagttctca cattataatt ttgatttttt tatttttttt ataaacaatc
3181 tggtaccctt ttatgtctgg aaggaaaaaa aaaatctaaa ttgcaacatt ttagtcggtg
3241 agaatggtac tctgtcctag ctactttcta cacatgagag agagagagag agagagagag
3301 agagccttta attgcccttg cccatgcatc tttctttgca cacatgtatg cttttcacat
3361 tgtcatgagg agagaacttg ttaagttgca cacatgtgtg ctttgcatgt cttcaggtta
3421 cctggaaggg catttgcaag gagaggccat gtatggctca ctggagcaaa tgaagttgac
3481 agcaaagtat tcctaagagc aattcttgcc aaggttcagc catcaccttc tcttacctat
3541 ttttcactct gaatgccaac agtgctttgc acattgtagt ctgtttgcag actgcaaatg
3601 atgaccataa tcagatcaga aaataaaata atattatata ctttttgagc cagctagcaa
3661 gaatatgtaa caataattct cctttttttt tcttgttctt ttccctgatg tggtgcataa
3721 caaataacca aactgatgaa tggcagagtg ctggtatcca ggtatttgcc tctaaaagta
3781 gctacacgtt tactatgaaa ttttgtggct tttgttcatc tttggatgca gtggccatta
3841 tctaaaaact atgaatttcc agactgcagt ttttatctaa ttttgtgact ttgtacatca
3901 gacagttgtg tgcattcctg ttgtcgatgg cgtcctggaa attggaacta cggaaaaggt
3961 gatttcgtat attatcagct gacaatctaa ttatatgggc catataatta agtataaatc
4021 aaaatacctc ataatatatt ataaagtatc taatgtgatt atgtgaatat tggctatttc
4081 aatgtaattt gatatatgaa actgataatc ctctgaaact ccgtaaggat caaactaatc
4141 aaaatgtata tattttcaag gtggaggaag atatgggcct gattcagtat gcaaggggca
4201 tcttcatgga tcaacatggc atccacatga agcctaccct ctcacagcac tcaacatcca
4261 acccagtcac ccactgtact catcagcatc caatccaggt tcagatgcaa ctaggtatca
4321 ccagccaaac aaagtttgat tattcagatg agctcaatgc agatgaggag aatgatgaca
4381 cagaagaaga gggcatgtca ggttcagaca ctaacaacac tgacactgaa aggaattcag
4441 gccagctgca acttcaaatg caagaccaac tgaacatggt gagcaatgac caccagacaa
4501 taccaaataa tgcagtttcc agtgagctaa tgcagtgtga gatgtcagaa gtggtaagag
4561 atggctgctc aaataatatt ttagaggatg aaatccaaat gctgatggat tgccaaaaca
4621 gtaattgtca gttaaatttg caagggccag atgagccttg tcactcttgg cattttctct
4681 gcgaggagtt acaaaatgat taccagccag gtattacatt tgagaagata atccttcaaa
4741 agcacccttg ttccaaaaat atatatttgt actcttcaca caagcactgc catttttttt
4801 cttttttgca tacatcctca attcttgcat ttcttttcca tatatttgat acaactgtct
4861 ccatttccct tctgtcacag ctactgaaga tcaagtggca tcacctgaaa atacccatta
4921 cccaaaaaca ctcatgacaa tcctacatta caacacgctg cgacagcaag agatgaacat
4981 caagaactac ttgccagttt cagagaaatc atcattctcc agatggacta ctcctgaagg
5041 aagtgatgac aacaagacca tgatcagtcc aggcaccaca cagagaatgc tcaagagcat
5101 cctgatgatt gttcccagta gtcactgcag ttacagggga gcagaaacac ctgaatcaag
5161 gggcgggaaa ggcgcaagtg gaacgcgaaa agtcggtgcc atccaaggtg atttcagtgc
5221 caaccatgtg ctgaaagaga ggagaagaag agagaagctc aatgagaagt tcataattct
5281 gcgatctttg gtacctttca tgacaaaggt aattaagtac tccctctatt tctataaagc
5341 cgtatttgac tagttatctt atttagaaag tatgtgcaaa tatgtaaaat ataagtcata
5401 cttaaagaac ttttaatgtt attaaataat aagtcacacc aaaaataaaa catatatatt
5461 tttaataaga caaatgatta aatgtatata taaaaattaa tagcgtcaca tattttaaaa
5521 tagaggggta tttaagtacc cacaggatca tcaaaattca gttatctttt cttaagcctc
5581 taacgaacat tggaagatcc tcactaatgg cagcatgaat ctagggttca ctatttcgga
5641 atgcaaaata tgttttaccg ggcatccgat ttttaaaaaa ttcagaatga agaaaattga
5701 atctttttta tggatttgaa taaatcttga taaattcgaa aaaatttccg aacttttggc
5761 cagaagtgaa tcctacccgt atccaccggt aataaaccta aatttttggg agtaatgaat
5821 taatgttata tataatccat gaattatata gttccaaact actccgtaac aaattttcag
5881 gagtagtgaa attaatatta ttacaatctc agaaaaaaat ggcagaaaca attaatctgt
5941 tttcaattat taattaattt gtttttgtgt ccagatggac aaggcgtcga tactaggcga
6001 cacgatcgag tacgtgaagc agctaaggaa ccgcatacaa gagctcgagt cgtcgtcgtc
6061 gtcgtcacga gcagccgccc gggcgccatc ggcggcggcc gccgggaggc ggaggaagag
6121 atccgccgcc gccgccactg ccacggcggc ggaagggatg agcagcagca atggccgcaa
6181 tggcggcgag gcggcggagg tggtgcaggt gtccatcatc gagagcgacg cgctgctgga
6241 gctccggtgc ggttgcggcg gcggcggcgg cggtgtggtg ctgctccggg tgatgcaggc
6301 gatgcaggag ctccagctgg aggtcaccgc cgtccaggcc tcgtgcgccg gtggcgagct
6361 gctcgccgag ctgcgcgcca aggtcgtcgt tatgatcctg atctgcatga aaatgcaaat
6421 gcaaatgcaa atgcagaatt aa
</dnaseqindica> |
| External Link(s) |