Difference between revisions of "Os11g0685700"
Tangmingxing (talk | contribs) (→Expression) |
(→Structured Information) |
||
| Line 24: | Line 24: | ||
The precise mechanism of WRKY-mediated gene regulation remains unclear. WRKY mRNA accumulates rapidly after induction and seemingly does not require the synthesis of new transcription factors. Some studies indicate that a few of WRKY proteins may be the target of mitogen-activated protein kinases (MAPK) that alter their activity.Both WIPK and SIPK, which are two well-studied stress-responsive kinases and can be phosphorylated by MAPK NtMEK2 (in tobacco) in response to infection and an unknown upstream MAPK,appear to be upstream of NtWRKY1 and NtWRKY3 in a defense-signaling cascade in Nicotiana. Besides,AtWRKY22 and AtWRKY29 have also been identified as being downstream of MAPK signaling cascades. A hypothetical model to explain PcPR10 gene activation, which which may be similarly applicable to other WRKY factors in other plants, demonstrates that W-box elements are generally occupied by WRKY factors that are either inactive or promoting basal RNA expression.Following recognition of an elicitor molecule by a receptor, an MAPK cascade is activated, with a protein kinase being translocated into the nucleus of the cell where it can hypothetically modify bound WRKY factors directly.This allosteric interaction causes them to release from their cognate W-box elements, thereby derepressing PcPR10 and PcWRKY1, which interacts with additional target promoters and also autoregulates its own expression. | The precise mechanism of WRKY-mediated gene regulation remains unclear. WRKY mRNA accumulates rapidly after induction and seemingly does not require the synthesis of new transcription factors. Some studies indicate that a few of WRKY proteins may be the target of mitogen-activated protein kinases (MAPK) that alter their activity.Both WIPK and SIPK, which are two well-studied stress-responsive kinases and can be phosphorylated by MAPK NtMEK2 (in tobacco) in response to infection and an unknown upstream MAPK,appear to be upstream of NtWRKY1 and NtWRKY3 in a defense-signaling cascade in Nicotiana. Besides,AtWRKY22 and AtWRKY29 have also been identified as being downstream of MAPK signaling cascades. A hypothetical model to explain PcPR10 gene activation, which which may be similarly applicable to other WRKY factors in other plants, demonstrates that W-box elements are generally occupied by WRKY factors that are either inactive or promoting basal RNA expression.Following recognition of an elicitor molecule by a receptor, an MAPK cascade is activated, with a protein kinase being translocated into the nucleus of the cell where it can hypothetically modify bound WRKY factors directly.This allosteric interaction causes them to release from their cognate W-box elements, thereby derepressing PcPR10 and PcWRKY1, which interacts with additional target promoters and also autoregulates its own expression. | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
[[Category:Genes]] | [[Category:Genes]] | ||
[[Category:Japonica mRNA]] | [[Category:Japonica mRNA]] | ||
[[Category:Oryza Sativa Japonica Group]] | [[Category:Oryza Sativa Japonica Group]] | ||
[[Category:Japonica Genes]] | [[Category:Japonica Genes]] | ||
Revision as of 04:29, 12 June 2015
Annotated Information
Function
1,Roles in disease resistance and responses to salicylic acid: The major role of WRKY genes in flowering plants is to mediate defense responses.WRKY18, WRKY40, and WRKY60 form both homocomplexes and heterocomplexes to regulate Arabidopsis responses to the hemibiotrophic bacterial pathogen Pseudomonas syringae and the necrotrophic fungal pathogen Botrytis cinerea.An indica rice WRKY gene, which is identical to OsWRKY12,is induced by the bacterial pathogen Xanthomonas oryzae pv. oryzae.For japonica, an analysis of OsWRKY genes revealed that the expression level of 15 WRKY genes is increased in an interaction between rice and Magnaporthe frisea, a fungal pathogen that causes devastating rice blast disease. Besides,the expression of two OsWRKY genes was increased in SA-treated leaves and that of three OsWRKY genes was increased by jasmonic acid (JA) treatment.
2.Roles in seed development and germination: MINI3 encodes WRKY10, a WRKY class transcription factor, which is responsible for seed growth and development. Besides, two WRKY genes are induced during somatic embryo genesis of orchardgrass. The products of WRKY genes, which named SUSIBA2,is a regulatory transcription factor in starch synthesis. OsWRKY51 and OsWRKY71 are two ABA-inducible and gibberellin (GA)-repressible rice WRKY genes, which mediate the cross-talk of GA and ABA signaling,can repress the expression of α- amylase gene, as the figure shows. TTG2,a group I WRKY protein is also responsible for trichome development and mucilage production in the seed coat of rice. What's more,WRKY genes is also involved in the process of senescence.Arabidopsis AtWRKY53 is expressed
in an age-dependent manner and, when overexpressed, leads to early flowering and senescence.

3,Roles in responses to abiotic stresses and ABA: In the desert plant Retama raetam, a WRKY gene was only induced when the plant was exposed to a combination of heat shock and drought.Several rice WRKY genes to be capable of regulating the ABA-inducible HVA22 promoter in a positive(OsWRKY72 and -77) or negative (OsWRKY24 and -45) manner.A dozen rice WRKY genes are induced by heat shock, cold stress, high salinity, and polyethylene glycol.Hv-WRKY38,the ortholog of OsWRKY51 and -71 ,is upregulated in response to drought and cold in vegetative tissues. Moreover,OsWRKY71 was upregulated by SA,JA, 1-aminocyclo-propane-1-carboxylic acid (ACC), wounding, and pathogen infection. Overexpression of OsWRKY71 in rice resulted in enhanced resistance to virulent Xoo 13751.
4,Roles in the biosynthesis of secondary metabolites: WRKY genes regulate the biosynthesis of sesquiterpene and benzylisoquinoline alkaloid.CjWRKY1 in C. japonica protoplasts was found to be responsible for the level of transcripts of berberine biosynthetic genes.
Evolution
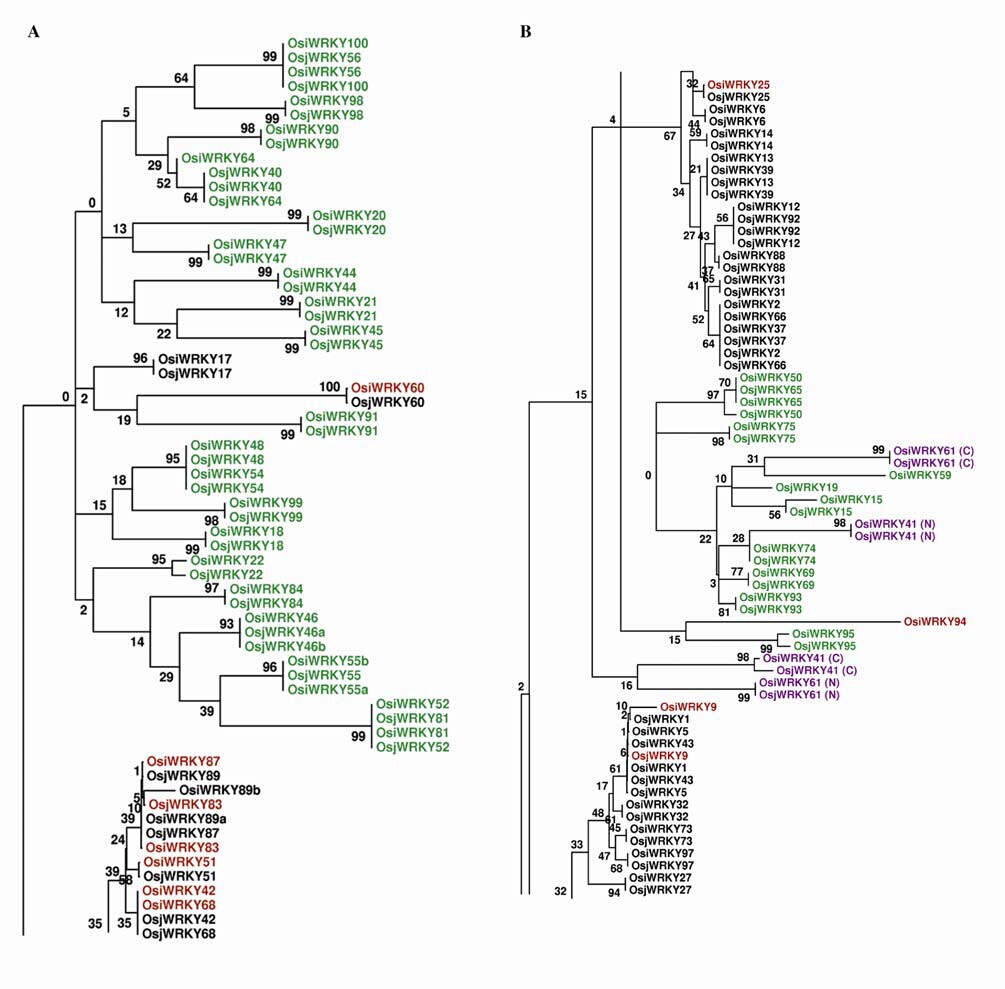
WRKY domains were initially defined by a nearly 60 amino acid long motif possessing the well-conserved amino acid signature WRKYGQK at their N-terminus and containing a novel zinc finger-like structure of the form CX4–5CX22–23HXH.WRKY proteins containing a single WRKY domain are group II proteins; those with two are group I.The group II WRKY proteins are further subdivided into subgroups a–e based on the presence of short conserved structural motifs.The group IId WRKY proteins have been shown to bind calmodulin through the defining non-classical calmodulin-binding domain (CaMBD) VSSFK(K/R) VISLL.A third group, group III, contains a single WRKY domain with a variant zinc-finger CX7CX23HXC.The terminal histidine of the zinc-finger of groups Ib and III is replaced with cysteine and the spacing is altered within group III.Group IV WRKY proteins contain the WRKY motif, but lack a complete zinc-finger. Multiple domain acquisition and loss events appear to have shaped the WRKY family. Numerous WRKY gene duplications occurred after the divergence of the monocotyledons from dicotyledons some 50–80 million years ago. The WRKY domains are found in expressed sequence tags (ESTs) from over 40 species of land plants.There is no evidence for WRKY genes in the archaea,eubacteria, the eukaryotic fungi, or animal lineages. However,a single WRKY gene with two WRKY domains exists in thegenomes of the protist Giardia lamblia and the slime mold Dictyostelium discoideum.the WRKY family may occur in primitive eukaryotes, before the emergence of the plant phyla, and its gross expansion during the course of plant evolutionary radiation, likely because of selective pressures favoring greater adaptability.The presence of a group I WRKY protein in these ancient organisms suggests that group I WRKY genes represent the ancestral form, with other groups arising later through losses and gains of a WRKY domain, and that this family originated some 1.8–2 billion years ago.Only group Ia genes, but not group Ib genes, appear to be derived from ancestral forms of the WRKY genes in plants. Researchers find evidence of loss of the N-terminus in the evolution of single-domain group II and group IV genes from group Ia genes and independent loss of the C-terminus in the evolution of a single-domain group II gene. Other group II genes may have evolved by this process.The group Ib OsWRKY genes are likely to have evolved by intramolecular duplication of a group III WRKY domain that had already evolved the C2HC type zinc-finger.Figure 2 shows that there is a difference between N- and C-terminal WRKY domains for those WRKY proteins with two WRKY domains. These domains tend to cluster into discrete groupings and there is more variability among the N- than the C-terminal WRKY domains. In the WRKY protein,the C-terminal domains are required for DNA-binding activity, thus are constrained in their ability to mutate without losing function, whereas the N-terminal domains mediate protein-protein interactions and may be less functionally constrained. The group II rice WRKY genes arose from the two-domain group Ia WRKY genes through a single domain loss event. The group Ib genes were derived from group III by domain duplication. The C- and N-terminal domains of group Ib genes are exclusively clustered with those of group 3 genes. The group Ia genes arose from a fusion of two group II WRKY genes evolved at an early time.The WRKY80 domain may represent the extant descendent of the group Ia C-terminal progenitor and WRKY57 a descendent of the N-terminal progenitor.
Expression
The precise mechanism of WRKY-mediated gene regulation remains unclear. WRKY mRNA accumulates rapidly after induction and seemingly does not require the synthesis of new transcription factors. Some studies indicate that a few of WRKY proteins may be the target of mitogen-activated protein kinases (MAPK) that alter their activity.Both WIPK and SIPK, which are two well-studied stress-responsive kinases and can be phosphorylated by MAPK NtMEK2 (in tobacco) in response to infection and an unknown upstream MAPK,appear to be upstream of NtWRKY1 and NtWRKY3 in a defense-signaling cascade in Nicotiana. Besides,AtWRKY22 and AtWRKY29 have also been identified as being downstream of MAPK signaling cascades. A hypothetical model to explain PcPR10 gene activation, which which may be similarly applicable to other WRKY factors in other plants, demonstrates that W-box elements are generally occupied by WRKY factors that are either inactive or promoting basal RNA expression.Following recognition of an elicitor molecule by a receptor, an MAPK cascade is activated, with a protein kinase being translocated into the nucleus of the cell where it can hypothetically modify bound WRKY factors directly.This allosteric interaction causes them to release from their cognate W-box elements, thereby derepressing PcPR10 and PcWRKY1, which interacts with additional target promoters and also autoregulates its own expression.