Os01g0293100
Please input one-sentence summary here.
Contents
Annotated Information
Function
Please input function information here.
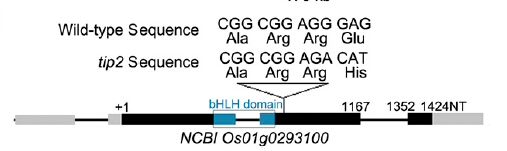
TDR INTERACTING PROTEIN2 (TIP2) is a basic helix-loop-helix (bHLH) protein, functions as a crucial switch in the meristemoid transition and differentiation during early anther development.
- The tip2 mutants display undifferentiated inner three anther wall layers.
- The tip2 anthers have prolonged periclinal cell division activity after the formation of the inner two anther wall layers via periclinal cell division.
- The tip2 displayed undegenerated callose around meiocytes, suggesting defective callose degeneration in the tip2 mutant.
- The tip2 mutants display abort tapetal programmed cell death.
TIP2 acts upstream of TDR and EAT1 and directly regulates the expression of TDR and EAT1, suggested that the bHLH proteins TIP2, TDR, and EAT1 play
a central role in regulating differentiation, morphogenesis, and degradation of anther somatic cell layers.
Mutation and Phenotype
Please input information here.
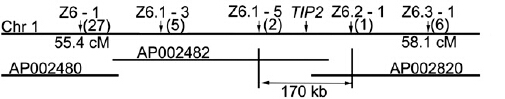
TIP2 locus on chromosome 1 between Z6.1-5 and Z6.2-1, two markers that defined a 170-kb region.
tip2 is a deletion of two continuous G nucleotides in it second exon.
Compared with the wild type, tip2 has normal vegetative growth, inflorescence and flower morphology. However, tip2 has smaller anthers and does not produce mature pollen grains during reproductive stage.
Compared with the wild type,the three inner somatic layers of tip2 anthers displayed no obvious difference in cell shape during stages 6 to 8.Furthermore, DAPI staining showed mutation in TIP2 does not directly affect meiotic initiation but causes the arrest of meiosis.
The rice tapetum marker gene,LIPID TRANSFER PROTEIN45 (LTP45)was preferentially expressed in tapetal cells at stage 7 and stage 8 in wild-type anthers, whereas in tip2 anthers its expression was not detectable.
In the wild type, meiocytes formed tetrads after meiosis, tapetal cells became condensed and initiated PCD-promoted degeneration, and the middle layer also started to be degraded.By contrast, cell morphology of the three inner anther wall layers of tip2 anthers remained similar, and no obvious tapetal PCD ignals were detected.
aniline blue showed tip2 displayed undegenerated callose around meiocytes.
Expression
Please input expression information here.
Using quantitative RT-PCR(qRT-PCR). TIP2 expression was specifically observed in anthers, from the meiosis stage to mitosis, with a maximum expression level at stages 7 and 8.Interestingly, tip2 anthers displayed a lower level of TIP2 than the wild type from stage 6 to stage 8, but a higher level than the wild type after stage 9, when young microspores were released.
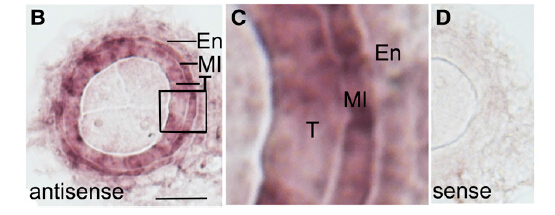
In situ hybridization showed strong expression of TIP2 in the middle layer and tapetum and weak expression in the endothecium in the wild type. By contrast, only background signal was detected in the same anther section with the sense probe.
Evolution
Please input evolution information here.
You can also add sub-section(s) at will.
Labs working on this gene
Please input related labs here.
References
Please input cited references here.
Structured Information
| Gene Name |
Os01g0293100 |
|---|---|
| Description |
Basic helix-loop-helix dimerisation region bHLH domain containing protein |
| Version |
NM_001049330.1 GI:115436073 GeneID:4325164 |
| Length |
2112 bp |
| Definition |
Oryza sativa Japonica Group Os01g0293100, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 1:10712748..10714859 |
| Sequence Coding Region |
10713003..10713074,10713260..10713712,10713812..10714426 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008394:10712748..10714859 source=RiceChromosome01 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008394:10712748..10714859 source=RiceChromosome01 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>atgtatcacccgcagtgcgagctcctgatgccgcttgagagcctggagatggacgtcggccagtcgcacctcgccgccgccgtcgcagcagccatgccgggggagctcaacttccacctcctccactcgctcgacgccgccgcggcggctgcctcctccaccgccgcctcggcctcctcccagcccaccgtcgactacttcttcggcggcgccgaccagcagccgccgccgccggcggcgatgcagtacgaccagctggcggcgccgcaccaccaccagacggtggccatgctgcgcgactactacggcggccactacccgccggcggcggcggcggcggcggccaccgaggcgtacttccgcggcgggccaaggacggccgggtcgtcgtcgctcgtgttcggcccggccgacgacgagtcggccttcatggtcggacccttcgagagctccccgacgccgcggtccggcggcggcaggaagcgtagccgcgccaccgccggcttccacggcggcgggccggccaacggcgtcgagaagaaggagaagcagcgccgcctgcggctcaccgagaagtacaacgccctcatgctcctcatccccaaccgcaccaaggaggatagagcgacggtgatctcagacgcgatcgagtacatccaggagctagggaggacggtggaggagctgacgctgctggtggagaagaagcggcggcggagggagatgcagggggacgtggtggacgcggcgacgtcgtcggtggtggcggggatggatcaggcggcggagagctcggagggcgaggtgatggcggcggcggcgatgggcgcggtggcaccgccgccgcggcaggcgccgatccggagcacgtacatccagcggcggagcaaggagacgttcgtggacgtgcggatcgtggaggacgacgtgaacatcaagctcaccaagcgccgccgcgacggctgtctcgccgccgcgtcgcgcgcgctggacgacctccgcctcgacctcgtccacctctccggcggcaagatcggcgactgccacatctacatgttcaacaccaagattcattcgggatctccagtgtttgcaagtgcagtggccagcaggctgattgaagtggtggatgagtactaa</cdnaseq> |
| Protein Sequence |
<aaseq>MYHPQCELLMPLESLEMDVGQSHLAAAVAAAMPGELNFHLLHSL DAAAAAASSTAASASSQPTVDYFFGGADQQPPPPAAMQYDQLAAPHHHQTVAMLRDYY GGHYPPAAAAAAATEAYFRGGPRTAGSSSLVFGPADDESAFMVGPFESSPTPRSGGGR KRSRATAGFHGGGPANGVEKKEKQRRLRLTEKYNALMLLIPNRTKEDRATVISDAIEY IQELGRTVEELTLLVEKKRRRREMQGDVVDAATSSVVAGMDQAAESSEGEVMAAAAMG AVAPPPRQAPIRSTYIQRRSKETFVDVRIVEDDVNIKLTKRRRDGCLAAASRALDDLR LDLVHLSGGKIGDCHIYMFNTKIHSGSPVFASAVASRLIEVVDEY</aaseq> |
| Gene Sequence |
<dnaseqindica>1786..1857#1148..1600#434..1048#aagaaaccaactgctttctcctacccaatatcacccttgccccttttatatactcttcctctcatcaccttctcgatcggcctctctcctctcctctcatcagctcacacccccaaccaacaaacctagttaatttagctctagttggttcatccctgctgcactgcgagctcaagtaatcgatctgagctctgaagaaaaaggtctgtttcaaattgaaccctttcatctcatcttctcattactgtttgattcattaattctagtaatttagagtttaaaccatcgcttttctttttgtgtgtgttcttgcatatgcttggacgattttgagtttgttctttaaatttgggcttgaacataggttgattaagcgagaagagtttcatggtttggattcttgatttgttgcaggtggtagagtgcgaggaagatgtatcacccgcagtgcgagctcctgatgccgcttgagagcctggagatggacgtcggccagtcgcacctcgccgccgccgtcgcagcagccatgccgggggagctcaacttccacctcctccactcgctcgacgccgccgcggcggctgcctcctccaccgccgcctcggcctcctcccagcccaccgtcgactacttcttcggcggcgccgaccagcagccgccgccgccggcggcgatgcagtacgaccagctggcggcgccgcaccaccaccagacggtggccatgctgcgcgactactacggcggccactacccgccggcggcggcggcggcggcggccaccgaggcgtacttccgcggcgggccaaggacggccgggtcgtcgtcgctcgtgttcggcccggccgacgacgagtcggccttcatggtcggacccttcgagagctccccgacgccgcggtccggcggcggcaggaagcgtagccgcgccaccgccggcttccacggcggcgggccggccaacggcgtcgagaagaaggagaagcagcgccgcctgcggctcaccgagaagtacaacgccctcatgctcctcatccccaaccgcaccaaggtatatcacatgccctagctaccatcgtcttcggtagttcgatctcttcttgtgttcatccatggatcgatcgatgatgattttgtctttctttttcaggaggatagagcgacggtgatctcagacgcgatcgagtacatccaggagctagggaggacggtggaggagctgacgctgctggtggagaagaagcggcggcggagggagatgcagggggacgtggtggacgcggcgacgtcgtcggtggtggcggggatggatcaggcggcggagagctcggagggcgaggtgatggcggcggcggcgatgggcgcggtggcaccgccgccgcggcaggcgccgatccggagcacgtacatccagcggcggagcaaggagacgttcgtggacgtgcggatcgtggaggacgacgtgaacatcaagctcaccaagcgccgccgcgacggctgtctcgccgccgcgtcgcgcgcgctggacgacctccgcctcgacctcgtccacctctccggcggcaagatcggcgactgccacatctacatgttcaacaccaaggtcacaacaaccaactaaaaatatccccgcgtttatctacaaaactaaccaatctcataatagtaattaattctagaacaattttatcgtgttgtgatcaacataaacctatatatgcaaataaattgctgctagttaatctcatcattgatttattattgtttatttggatgcgatgcatgcagattcattcgggatctccagtgtttgcaagtgcagtggccagcaggctgattgaagtggtggatgagtactaactagctcgagctagctaattagccgaccgaccgatcgatatgatgaaagtttctatgttgctagctagctagggttcttggatgcatgagtactgagtagctctttaattaatttccttttaattttagactgtttaatttggattggtaaagactcgtgttagcttttgggagatctttggtatgtcatggtttgcatgtattattttggtctacttggataaataattgatgctctttgagacgttaattaat</dnaseqindica> |
| External Link(s) |