Os07g0182900
Please input one-sentence summary here.
Contents
Annotated Information
Function
DNA methylation is a heritable epigenetic mark that regulates expression of genes involved in developmental processes [1]. In mammals, DNA methylation predominantly occurs in CG dinucleotides and is maintained by DNA methyltransferase1, While in plants, CG methylation is maintained by methyltransferase1(MET1) [1]. Methytransferase (OsMET1) genes from rice (Oryza sativa L) have two types, OsMET1a and OsMET1b, on chromosomes 3 and 7, respectively, each encoding a cytosine-5 DNA methyltransferase (MTase), which are mainly responsible for maintaining CG dinucleotide methylation after DNA replication [1] [2].OsMET1 is a plant homolog of mammalian DNA methyltransferase1(DNMT1), and MET1b plays a major role in the maintenance DNA methylation compared to MET1a, which is indispensable for the normal development of embryos in rice [1].
Sequence comparison between OsMET1a and OsMET1b
- OsMET1a has an open reading frame of 4,566 nucleotides with 12 exons and 11 introns while OsMET1b has an open reading frame of 4,491; nucleotides with 11 exons and 10 introns. Although OsMET1a and OsMET1b have high sequence similarity overall, they share only 24%; dentity in exon 1, and intron 3 of OsMET1a is absent from OsMET1b.
- A lysine–glycine repeat that is highly conserved in DNA MTases is present two-thirds of the way through the protein, separating the catalytic and regulatory domains in both OsMET1a and OsMET1b.
- The amino acid sequence downstream of the lysine–glycine repeat motif in OsMET1a showed a higher conservation (86.5%) with that of OsMET1b than did that (67.7%) for the N terminal domain (residues 1–1059). The inferred amino acid sequences of OsMET1a and OsMET1b contain several functional regions.
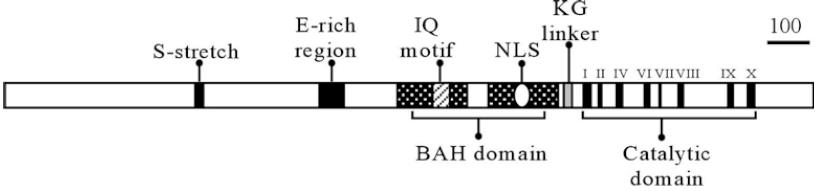
Figure 1:Proportionate diagram of OsMET1-1 showing conserved domains: S-stretch serine-rich region; E-rich region glutamic acid-rich region; BAH bromo adjacent homology domain (black boxes with white dots); IQ motif (calmodulin binding) isoleucine (I) and glutamine (Q) motif (hatched box); NLS putative nuclear localization signal (oval); KG linker the lysine-glycine repeat, KNKGKG (gray box); I to X the catalytic domain with eight highly conserved DNA MTase motifs (black bars).
Mutation assay
- RNAi-mediated knockdown analysis of the rice MET1 genes showed that it resulted in reactivation of a silenced β-glucuronidase (GUS) transgene in rice callus.
- A Osmet1a null mutant has no obvious phenotypes, while Osmet1b null mutant exhibited abnormal seed phenotypes, such as early embryonic lethality, decreased levels of DNA methylation at repetitive CentO sequences and at the FIEI gene locus in the embryos.
Figure 2.Phenotyes of normal type seeds and OsMET1b null mutant type(A to C) seeds
Expression
- Ribonuclease protection assays showed that MET1a and MET1b both express in highly replicating and dividing cells;
- MET1b is more abundantly expressed than MET1a in all of the tissues examined, like callus, root and inflorescence.
- OsMET1b does not express in differentiated tissue (10-day-old leaf), and no expression for either gene was found in mature leaves.
Evolution
This phylogenetic relationships between eukaryotic and prokaryotic DNA methytransferase were derived using the AlignX program of the VectorNTI suite, suggesting that the rice DNA methytransferase are related to DNA methytransferase of Dnmt1/MET1 class in diverse organisms from several kingdom.
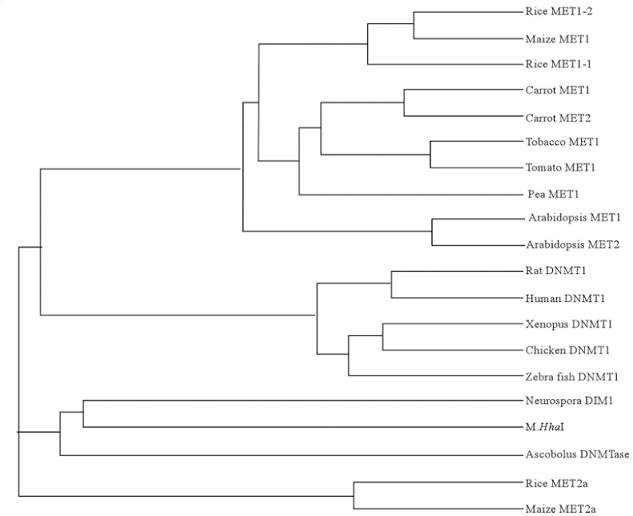
Figure 3. Phylogenetic relationships between eukaryotic and prokaryotic DNA methytransferase Phylogenetic relationships between eukaryotic and prokaryotic MTases were derived using the AlignX program of the VectorNTI suite with the following GenBank accessions: maize MET1 (AF063403), rice MET1-1 (OsMET1-1, AF462029), rice MET1-2 (OsMET1-2, TPA BK001405), Arabidopsis MET1(AtMET1, AT5G49160) and MET2 (MET2-type C-5 DNA MTase, AT4G08990), pea MET1 (putative C-5 DNA MTase, AF034419), carrot MET1 (carrot C-5 DNA MTase, AF007807) and MET2 (MET2-type C-5 DNA MTase, AF007808), tobacco MET1 (NtMET1, AB030726), tomato (putative C-5 DNA MTase AJ002140), rat DNMT1 (AB012214), human DNMT1 (AF180682), Xenopus DNMT1(D78638), zebra fish DNMT1 (AF483203), chicken DNMT1 (2112268A), Neurospora DIM2 (AF348971), Ascobolus DNMTase (Z96933), rice MET2a (AC069324), maize MET2a (AF243043) and Haemophilus haemolyticus M.HhaI (XYH1H1)
Knowledge extension
Apart from the main information listed above about function, expression and evolution about OsMET1 gene, there are also some advanced researches about this gene being conducted in the past few years. Takaki Yamauchi et al. reported in 2007 that the alternative splicing mechanisms of OsMET1a transcript and OsMET1b transcript differ, which is key to the regulation of active OsMET1 protein generation in different tissues. Daisuke Miki et al. reported in 2008 that de novo DNA methylation can be induced by siRNA targeted to endogenous transcribed sequences which is gene-specific and OsMET1-indepent in rice, which is also the first time to observe transcriptional gene silencing by siRNAs. Takaki Yamauchi et al. reported in 2009 that they successfully realized homologous recombination-promoted knock-in targeting to monitor gene expression by fusing a reporter gene named GUS codingβ-glucuronidase with its promoter in rice, in principle, this method can be applied to modify any endogenous gene which would promote basic and applied plant research.
Labs working on this gene
- Institute of Developmental and Molecular Biology, and Department of Biology, Texas A&M University, College Station, TX, USA
- National Institute for Basic Biology, Okazaki, Japan
- Graduate School of Science and Technology, Chiba University, Matsudo, Japan
- Graduate School of Bioagricultural Sciences, Nagoya University, Nagoya, Japan
- Graduate School of Nutritional and Environmental Sciences, University of Shizuoka, Shizuoka, Japan
- Faculty of Agriculture, Meijo University, Nagoya, Japan
- Graduate School of Horticulture, Chiba University, Matsudo, Japan
- Laboratory of Plant Molecular Genetics, Nara Institute of Science and Technology (NAIST), Ikoma, Japan
- Graduate School of Agriculture and Life Sciences, University of Tokyo, Tokyo, Japan
- Department of Basic Biology, School of Life Science, The Graduate University for Advanced Studies
References
Please input cited references here.
Structured Information
| Gene Name |
Os07g0182900 |
|---|---|
| Description |
Similar to Cytosine-5 DNA methyltransferase MET1 (Fragment) |
| Version |
NM_001065587.1 GI:115470906 GeneID:4342575 |
| Length |
2298 bp |
| Definition |
Oryza sativa Japonica Group Os07g0182900, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 7:4421779..4424076 |
| Sequence Coding Region |
4421780..4421864,4421955..4422066,4422154..4422301,4422417..4422614,4422710..4422988 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008400:4421779..4424076 source=RiceChromosome07 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008400:4421779..4424076 source=RiceChromosome07 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>acaaaggtgtccaaagagaaccgtcttgcaactcttgacatttttgctggctgtggaggtttatcagaaggactgcagcaagctggtgtatcgtttacaaagtgggcgattgaatatgaggaaccagctggtgaagcatttaccaaaaatcatccagaagctgcggtgtttgtggataactgcaatgtgattttgaaggcaattatggacaaatgtggagatgctgatgattgcatttcaacttctgaggctgctgaacaagcagctaaattttctcaggacaatattatgaaccttcctgtccctggtgaagtagaattcataaatggtggtcctccatgtcagggcttttctgggatgaacagatttaaccaaagcccatggagtaaagttcaatgtgagatgattttagcattcctgtcttttgctgaatatttccgccccagattctttcttttagaaaatgttaggaactttgtttcattcaacaaaggacagacatttcgactgacagttgcatcccttctggagatgggataccaggtccggtttggaattttagaggcagggacttttggtgttgctcagtccaggaaaagagcattcatttgggctgctgcacctggagagactctgcctgattggccagaaccaatgcatgtgtttgctagccctgagctgaaaataaatttgcctgatggtaaatactatgcagctgcaaaaagcactgctggtggtgctcctttccgtgcaataacagttagagatacaataggcgatttaccaaaggtggagaatggcgccagtaaactcctacttgagtacggtggcgaacccatctcctggttccagaagaagattagagggaacacgatcgcgctgaatgatcacatatctaaggagatgaatgaactgaacctcatcagatgccaacgcattccaaagcggcctggttgtgattggcatgacctaccagatgagaaggtgaaactatcatctggccaactggtggacctgatcccttggtgcctgcctaacacagctaaaaggcacaaccagtggaaagggctttatggtaggctggattgggagggcaatttccctacttctgtgacagacccccagccaatgggcaaggttggcatgtgcttccaccctgaccaggataggatcatcacagtccgtgaatgtgcgcgatctcagggctttcctgacaactaccagttcgcgggcaacatccagagcaagcacaggcagattggcaatgcggtgcctccacctcttgccttcgccctcgggaggaaactgaaggaagctgttgatgcaaagcgtcagtag</cdnaseq> |
| Protein Sequence |
<aaseq>TKVSKENRLATLDIFAGCGGLSEGLQQAGVSFTKWAIEYEEPAG EAFTKNHPEAAVFVDNCNVILKAIMDKCGDADDCISTSEAAEQAAKFSQDNIMNLPVP GEVEFINGGPPCQGFSGMNRFNQSPWSKVQCEMILAFLSFAEYFRPRFFLLENVRNFV SFNKGQTFRLTVASLLEMGYQVRFGILEAGTFGVAQSRKRAFIWAAAPGETLPDWPEP MHVFASPELKINLPDGKYYAAAKSTAGGAPFRAITVRDTIGDLPKVENGASKLLLEYG GEPISWFQKKIRGNTIALNDHISKEMNELNLIRCQRIPKRPGCDWHDLPDEKVKLSSG QLVDLIPWCLPNTAKRHNQWKGLYGRLDWEGNFPTSVTDPQPMGKVGMCFHPDQDRII TVRECARSQGFPDNYQFAGNIQSKHRQIGNAVPPPLAFALGRKLKEAVDAKRQ</aaseq> |
| Gene Sequence |
<dnaseqindica>2..86#177..288#376..523#639..836#932..1210#1306..1467#1566..1784#1898..2032#tacaaaggtgtccaaagagaaccgtcttgcaactcttgacatttttgctggctgtggaggtttatcagaaggactgcagcaagctggtatgcactttccttcaataatggtgtttagctttatcatcaacatgtcggccatcagactgatattttattaccatttaaattctgtaggtgtatcgtttacaaagtgggcgattgaatatgaggaaccagctggtgaagcatttaccaaaaatcatccagaagctgcggtgtttgtggataactgcaatgtgattttgaagtaggtgcaccgtgcacttccttgattttttctgtggttttctttttacctatatagctcaatagggactgttatatgattttgcagggcaattatggacaaatgtggagatgctgatgattgcatttcaacttctgaggctgctgaacaagcagctaaattttctcaggacaatattatgaaccttcctgtccctggtgaagtagaattcataaatggtggtcctccatgtcaggtatattattttgctatacttgatggaattttctttgcacctaggcaagcaaattgtttgcacttcaatacttgtgataaatgacttcaattttgtaatttttttctcaactcagggcttttctgggatgaacagatttaaccaaagcccatggagtaaagttcaatgtgagatgattttagcattcctgtcttttgctgaatatttccgccccagattctttcttttagaaaatgttaggaactttgtttcattcaacaaaggacagacatttcgactgacagttgcatcccttctggagatgggataccaggtaattattcttctggttatacatacttaaaatgcctatgtacagcatttgttttgaatcttataataaccagaagctgtttacttgttgcttaggtccggtttggaattttagaggcagggacttttggtgttgctcagtccaggaaaagagcattcatttgggctgctgcacctggagagactctgcctgattggccagaaccaatgcatgtgtttgctagccctgagctgaaaataaatttgcctgatggtaaatactatgcagctgcaaaaagcactgctggtggtgctcctttccgtgcaataacagttagagatacaataggcgatttaccaaaggtggagaatggcgccagtaaactcctacttgaggtaaaattgcttctcctatatggctatccgctccattttatcctgtttcttctcttgatttatatgctgaatgaatcccctcttttgtaatgcagtacggtggcgaacccatctcctggttccagaagaagattagagggaacacgatcgcgctgaatgatcacatatctaaggagatgaatgaactgaacctcatcagatgccaacgcattccaaagcggcctggttgtgattggcatgacctaccagatgagaaggtaaatgccatctaccttgcggttgcattcacttccttttgtgctcttccatacattccttgcatcagcggaatgttaaccattatgagcgtgtgcaggtgaaactatcatctggccaactggtggacctgatcccttggtgcctgcctaacacagctaaaaggcacaaccagtggaaagggctttatggtaggctggattgggagggcaatttccctacttctgtgacagacccccagccaatgggcaaggttggcatgtgcttccaccctgaccaggataggatcatcacagtccgtgaatgtgcgcgatctcaggtaagctgctattgctatccatccattcaacattctcttcctgtcttctaagatattgtgaatttggaggggagtcagtactgaccgtttaactaacctattcttgtctgcagggctttcctgacaactaccagttcgcgggcaacatccagagcaagcacaggcagattggcaatgcggtgcctccacctcttgccttcgccctcgggaggaaactgaaggaagctgttgatgcaaagcgtcagtaggtgctcagagtccttctccgatgatcgagggagatgattctcccctgaaaagaaacaaacaaccaaacatatgagtgtacttattttaattgtgcgctgcatttaactttgtgtgttgattaagattttagtcatatagatccttggtaaccttgtacattttagaggttgtgttgtgattgatacccttgattcaggggtatgatttagtggctgtattcagatatttaaacaaaataatactagttagtgatggttgaattgtt</dnaseqindica> |
| External Link(s) |