1. Database Overview
-
-
1.1 Database Introduction
IO-LncLand is a comprehensive pan-cancer single-cell atlas of immune-associated long noncoding RNAs across the tumor microenvironment. Built from more than 370,000 immune cells spanning 18 cancer types, the atlas systematically maps lncRNA expression programs, regulatory circuits, and functional states across major lymphoid and myeloid lineages. By integrating single-cell transcriptomics with large-scale CRISPR functional screens and clinical outcomes, IO-LncLand prioritizes immune-related lncRNAs that govern immune cell fate while sustaining tumor fitness, providing a powerful resource for biomarker discovery and precision immunotherapy.
-
1.2 Main Function Overview
IO-LncLand offers the following core functions:
Gene Search: Search for lncRNA expression and function across different immune cell types by gene name
Cell Type Search: Search for lncRNA expression and function across different immune cell types by gene name
Multi-dimensional Analysis: Analyze lncRNA functions from multiple perspectives including LncRNA cell marker, Tumor-reactive lncRNA, Cell state-associated lncRNA, Cell-type-specific lncRNA network, TAM-associated lncRNA, Cell-type-specific prognostic lncRNA and CRISPR-validated lncRNA
Data Visualization: Provide intuitive charts to display lncRNA expression patterns and regulatory relationships


-
2. Detailed Explanation of Homepage Functional Modules
2.1 LncRNA cell marker
This module showcases lncRNA markers with cell type-specific expression patterns, which can be used to identify and characterize T cells or myeloid cells.
Key Metrics:
Smart-seq platform: log2FC > 1, expression ratio > 20%, FDR < 0.05
10x platform: log2FC > 0.1, expression ratio > 20%, FDR < 0.05
Main Functions:
Filter lncRNA markers across different cancer types and immune cell subsets
View lncRNA expression patterns and statistical significance

2.2 Tumor-reactive lncRNA
This module presents lncRNAs specifically expressed in tumor-reactive T cells, which may be serve as tumor-reactivity markers.

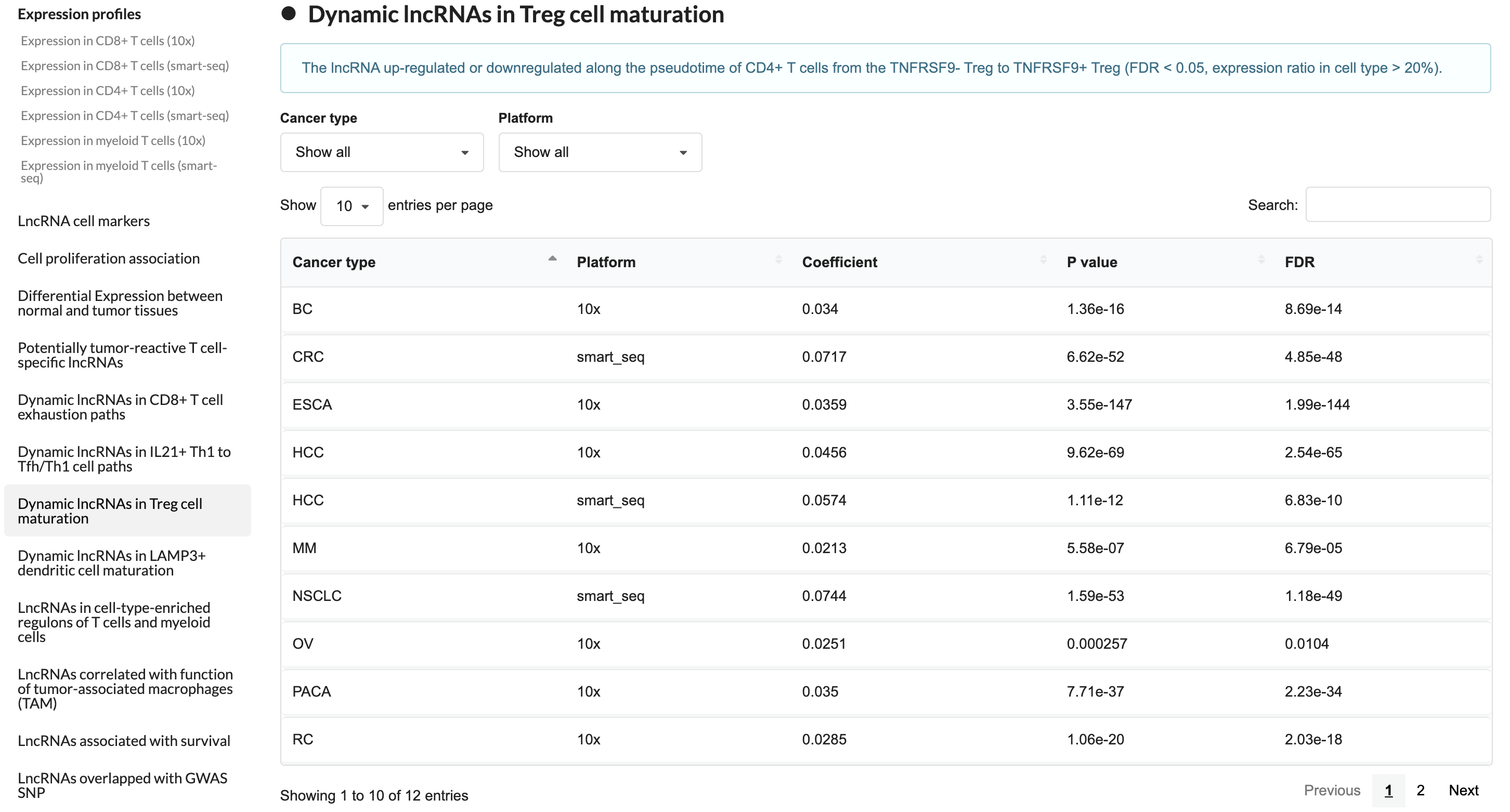
2.3 Cell-state associated lncRNA
This module displays lncRNAs associated with immune cell state transitions, including processes such as CD8+ T cell exhaustion, Treg activation, and LAMP3+ dendritic cell maturation.


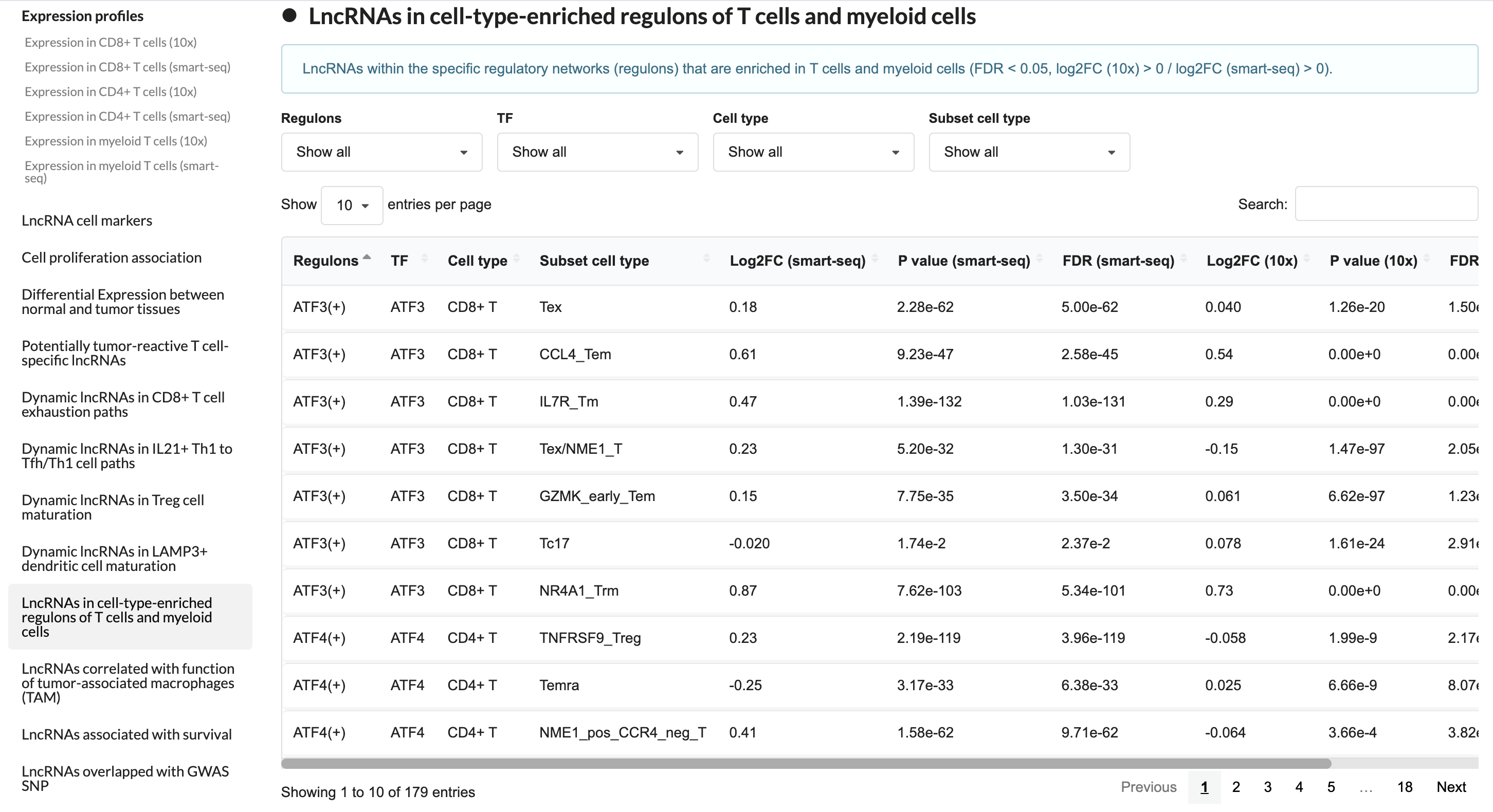
2.4 Cell-type-specific lncRNA network
This module showcases cell type-specific lncRNA regulatory networks, including transcription factor-lncRNA regulatory relationships and signaling pathways involving lncRNAs.

2.5 TAM-associated lncRNA
This module presents lncRNAs associated with tumor-associated macrophage (TAM) functions, particularly those involved in angiogenesis regulation.

2.6 Cell-type-specific prognostic lncRNA
This module displays cell type-specific lncRNAs associated with clinical prognosis.
2.7 CRISPR-validated lncRNA
This module showcases lncRNAs validated through CRISPR functional screens, which are essential for tumor cell survival or proliferation.
3. Detailed Explanation of Gene Page Functions
Users can search for detailed information on specific lncRNAs by gene name. The search results page includes the following sections:
Basic gene information (Chromosomal location, transcript structure, etc.)
Expression profiles across different immune cell types
Immune cell-specific lncRNA markers
Participation in immune cell-specific regulatory networks
Association with tumor-reactive T cell states
Association with angiogenic functions of tumor-associated macrophages (TAMs)
Correlation with immune cell differentiation states (T cell exhaustion, Tfh/Th1 state transitions, Treg and LAMP3+ cDCs differentiation)
Association with clinical prognosis


4. Detailed Explanation of Cell Type Page Functions
4.1 Statistical Dimension Description
The Cell Type page summarizes functional modules associated with each gene across different cell types, including:
Cell marker
DE tumor normal
Potentially tumor-reactive T marker
Cell-state-transition marker
Cell-type-enriched regulon target
Angiogenesis association
Survival
T cell proliferation
4.2 Functional Module Association Display
The page presents the participation of each gene in different functional modules in tabular form, with checkmarks indicating involvement. Users can quickly find combinations of genes and functional modules of interest through filtering functions.


5. Contact us
If you have any questions or comments, please feel free to contact us via email (jiangs@cqmu.edu.cn, wangshiting@big.ac.cn, qianqiheng2018m@big.ac.cn).
6. Licenses
IO-LncLand is free for academic use only. For any commercial use, please contact us for commercial licensing terms.