|
|
| Line 34: |
Line 34: |
| | ===Characteristics === | | ===Characteristics === |
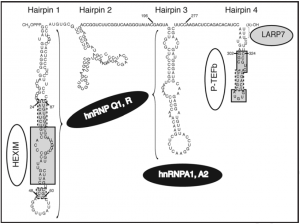
| | [[File:Functional domains of 7SK RNA..png|right|thumb|Functional domains of 7SK RNA([https://www.ncbi.nlm.nih.gov/pubmed/19246988 (Diribarne 2009)])]] | | [[File:Functional domains of 7SK RNA..png|right|thumb|Functional domains of 7SK RNA([https://www.ncbi.nlm.nih.gov/pubmed/19246988 (Diribarne 2009)])]] |
| − |
| |
| − | ~330 nt in vertebrates*. Transcribed by RNAP III, GC-rich sequence forming conserved secondary structures (especially 3' and 5' stem-loop motifs).
| |
| − |
| |
| − | The 7SK gene is located on chromosome 6, and chromosome 6 is the sole human chromosome that produces 7SK RNA ([https://www.ncbi.nlm.nih.gov/pubmed/8139910 (Driscoll 1994)])
| |
| − |
| |
| − | 7SK RNA is capped at its 5' end by BCDIN3, a specific methylase methylphosphate capping enzyme (MePCE) ([http://www.ncbi.nlm.nih.gov/pubmed/17643375 (Jeronimo 2007)]).
| |
| − |
| |
| − | RNAP II was recently found to bind near 7SK promoter, as well as many other known Pol III genes, suggesting that RNAP II may also play a role in regulating their transcription ([http://www.ncbi.nlm.nih.gov/pubmed/20139302 (Raha 2010)]).
| |
| − |
| |
| − | In invertebrates, 7SK homologs may have different sizes (such as >400 nt and ~130 nt in drosophilids and nematodes, respectively). ([http://www.ncbi.nlm.nih.gov/pubmed/20139302 (Gruber 2008)]) ([http://www.ncbi.nlm.nih.gov/pubmed/18566019 (Marz 2009)])
| |
| | | | |
| | ===Expression=== | | ===Expression=== |
| − | Nuclear, highly abundant (one of the most abundant small RNAs in vertebrate cells), first isolated from HeLa nuclear extracts, but ubiquitously expressed.
| |
| − |
| |
| − | RNA sequencing from 11 humans tissues confirmed ubiquitous high expression of 7SK with expression in some tissues being higher than any mRNA ([http://www.ncbi.nlm.nih.gov/pubmed/20668672 (Castle 2010)]).
| |
| | | | |
| | ===Regulation=== | | ===Regulation=== |
| − | In the 7SK ribonucleoprotein, Larp7 binds directly to 3′ terminus of 7SK RNA ([https://www.ncbi.nlm.nih.gov/pubmed/18281698 (Krueger 2008)]) ([https://www.ncbi.nlm.nih.gov/pubmed/18483487 (Markert 2008)]), and prevents degradation of 7SK in vivo ([https://www.ncbi.nlm.nih.gov/pubmed/18281698 (Krueger 2008)]).
| |
| | | | |
| | ===Function=== | | ===Function=== |
| | [[File:Model of hLarp7 recognition of the 7SK.png|right|thumb|Model of hLarp7 recognition of the 7SK 3′end and mechanism of assembly of core 7SK RNP([https://www.ncbi.nlm.nih.gov/pubmed/29946027 (Eichhorn 2018)])]] | | [[File:Model of hLarp7 recognition of the 7SK.png|right|thumb|Model of hLarp7 recognition of the 7SK 3′end and mechanism of assembly of core 7SK RNP([https://www.ncbi.nlm.nih.gov/pubmed/29946027 (Eichhorn 2018)])]] |
| − |
| |
| − | 7SK snRNA functions in transcriptional regulation by interacting with PTEF-B complex ([http://www.ncbi.nlm.nih.gov/pubmed/11713533 (Nguyen 2001)]) ([http://www.ncbi.nlm.nih.gov/pubmed/11713532 (Yang 2001)]), BAF chromatin-remodeling complex ([https://www.ncbi.nlm.nih.gov/pubmed/26878240 (Flynn 2016)]), or hnRNP R ([https://www.ncbi.nlm.nih.gov/pubmed/29507242 (Briese 2018)]). Consistently, it has been found highly enriched in isolated chromatin fractions, which may be related to its role in transcriptional regulation ([http://www.ncbi.nlm.nih.gov/pubmed/20404130 (Mondal 2010)]). In addition to its critical role for controlling transcription, 7SK snRNA is also involved in alternative splicing ([http://www.ncbi.nlm.nih.gov/pubmed/19416841 (Barboric 2009)]) and the localization of protein in nucleolus ([http://www.ncbi.nlm.nih.gov/pubmed/17381310 (He 2007)]). Therefore, 7SK snRNA has a variety of functions in the nuclear, playing important roles in cell growth and differentiation ([http://www.ncbi.nlm.nih.gov/pubmed/11713533 (Nguyen 2001)]) ([http://www.ncbi.nlm.nih.gov/pubmed/11713532 (Yang 2001)]), axon maintenance ([https://www.ncbi.nlm.nih.gov/pubmed/29507242 (Briese 2018)]) and vertebrate development ([http://www.ncbi.nlm.nih.gov/pubmed/19416841 (Barboric 2009)]).
| |
| − |
| |
| − | 7SK snRNA controls RNAP II activity by inhibiting P-TEFb elongation factor, which is a cdk-cyclin kinase that functions as both a general and an HIV-1 Tat-specific transcription factor ([http://www.ncbi.nlm.nih.gov/pubmed/11713533 (Nguyen 2001)]) ([http://www.ncbi.nlm.nih.gov/pubmed/11713532 (Yang 2001)]), with an impact on cell growth and differentiation. Specifically, 7SK snRNA functions as the central scaffold that coordinates protein-protein interactions and, by inhibiting P-TEFb kinase-mediated CTD phosphorylation, regulates RNAP II elongation ([http://www.ncbi.nlm.nih.gov/pubmed/11713533 (Nguyen 2001)]).
| |
| − |
| |
| − | At an early stage of the HIV transcription cycle, elongation is prevented as P-TEFb is recruited to the HIV-1 promoter in a catalytically inactive state bound to the 7SK snRNP and also the Tat trans-activator of transcription protein. The inhibitory 7SK snRNP may be displaced by the nascent TAR HIV RNA that also binds Tat protein, activating P-TEFb kinase and transcriptional elongation ([http://www.ncbi.nlm.nih.gov/pubmed/20562857 (D'Orso 2010)]). Displacement of 7SK may also be performed by cellular RNAs, as indicated by the 3'-untranslated region (~300-nt) of HIC mRNA, which forms complexes with P-TEFb and is necessary and sufficient for stimulation of P-TEFb-dependent transcription of the HIV promoter ([http://www.ncbi.nlm.nih.gov/pubmed/17925858 (Young 2007)]).
| |
| − |
| |
| − | 7SK snRNA inhibits enhancer transcription by modulating nucleosome position. 7SK physically interacts with the BAF chromatin-remodeling complex, recruits BAF to enhancers and inhibits enhancer transcription by modulating chromatin structure ([https://www.ncbi.nlm.nih.gov/pubmed/26878240 (Flynn 2016)]).
| |
| − |
| |
| − | In axons, 7SK snRNA interacts with hnRNP R to regulate its function in axon maintenance ([https://www.ncbi.nlm.nih.gov/pubmed/29507242 (Briese 2018)]).
| |
| − |
| |
| − | 7SK snRNP (composed of 7SK snRNA, Hexim1, Larp7/Pip7S, and the P-TEFb subunits CycT1 and Cdk9) is not only critical for controlling transcription, but also for regulating alternative splicing coupled to transcription elongation ([http://www.ncbi.nlm.nih.gov/pubmed/19416841 (Barboric 2009)]). 7SK snRNP disintegration promotes inclusion of an alternative exon via the increased occupancy of P-TEFb, Ser2-phosphorylated (Ser2-P) RNAPII, and the splicing factor SF2/ASF at the minigene ([http://www.ncbi.nlm.nih.gov/pubmed/19416841 (Barboric 2009)]).
| |
| − |
| |
| − | 7SK snRNA also inhibits APOBEC3C deaminase activity and sequesters it to the nucleolus, suggesting broader role for 7SK RNA in regulating key nuclear functions ([http://www.ncbi.nlm.nih.gov/pubmed/17381310 (He 2007)]).
| |
| | | | |
| | ===Disease=== | | ===Disease=== |
| Line 77: |
Line 49: |
| | | | |
| | ==Labs working on this lncRNA== | | ==Labs working on this lncRNA== |
| − | Please input related labs here.
| + | * College of Veterinary Medicine, Northwest A&F University, Yangling, Shaanxi 712100, China.<ref name="ref1" /> |
| | | | |
| | ==References== | | ==References== |
Annotation
Name
Approved symbol: LINC00319
Approved name: long intergenic non-protein coding RNA 319
HGNC ID: HGNC:19730
Previous symbols: C21orf125; NCRNA00319
Previous names: chromosome 21 open reading frame 125; non-protein coding RNA 319
Alias symbols: PRED49; FLJ38036
RefSeq ID: NR_026960
LncBook ID: HSALNT0289323
Disease
Disease: lung cancer (PMID:28800794)
Dysfunction type: regulation
Description: Linc00319 promotes cell proliferation and invasion in lung cancer cells by directly binding with and downregulating the tumor suppressor miR-32.
Function
Function Mechanism: ceRNA (PMID:28800794)
Biological Process: pathogenic process
Description: NA
Characteristics
Expression
Regulation
Function
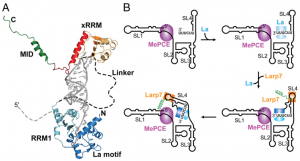
Model of hLarp7 recognition of the 7SK 3′end and mechanism of assembly of core 7SK RNP(
(Eichhorn 2018))
Disease
colon adenocarcinoma [1]
Evolution
Please input evolution information here.
Labs working on this lncRNA
- College of Veterinary Medicine, Northwest A&F University, Yangling, Shaanxi 712100, China.[1]
References
- ↑ 1.0 1.1 Zhou B, Yuan W and Li X. Long Intergenic Noncoding RNA 319 (linc00319) Promotes Cell Proliferation and Invasion in Lung Cancer Cells by Directly Downregulating the Tumor Suppressor MiR-32[J]. Oncol Res, 2017.