DEP2
Contents
Annotated Information
Function
- DEP2 gene reduced the development in the longitudinal direction but enhanced the development in the lateral direction of plural organs in rice. In the DEP2 mutants,the panicle length was reduced and panicle diameter was increased[1].
- Besides panicle erectness, the DEP2 mutants also showed a slight reduction in plant height , an obvious decrease in panicle length, and a significant increase in both rachis and stem diameter. The leaves of DEP2 are short, wide, and erect, and the overall appearance of the mutant is more compact. The grain density is increased due to the decreased panicle length but not the change of grain number. The grains of the DEP2 mutant are wider and shorter than the wild type[1].
- The mutation in DEP2 has pleiotropic effects on plant architecture,and that increased diameter of the rachis and decreased panicle length altogether contributed to the dense and erect panicle phenotype[1].
- The DEP2 gene may regulate GA synthesis, and mutation of DEP2 leads to a decrease in plant height[1].

Map-based cloning of the DEP2 gene(from reference.[1].)
Mutation
- The DEP2 locus was mapped to the long arm of rice chromosome 7[1].
dep2-1:The 31-bp deletion in dep2-1 starts at 2184bp from the initiation codon ATG and caused a frameshift; dep2-2:the G/A substitution in the second intron of dep2-2 caused an altered splicing site of the second intron and also led to a frameshift. LOC_Os07g42410 was considered a candidate for the DEP2 gene[1].
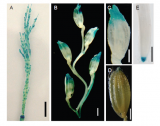
DEP2 expression pattern[1]
Expression
- DEP2 gene expression in the dividing zones of rachis, branches, and florets was higher than in other parts of the rice plant.
- This gene is highly expressed in young panicles ranging from 1 to 15 cm in length;low level expression was also detected in the other organs, including roots,stems,leaves and leaf sheathes.
Evolution
- DEP2 may be a plant-specific protein. The only existing low similarity to CIP7 in Arabidopsis is found at the N terminal part of the protein, while the COP1-interacting motif, transcription activation domain, and the nuclear localization signal domain are all missing in DEP2.
Knowledge extension
- Except dep2,cl7(t) was a novel allele of dep2. Sequence analysis showed that cl7(t) had a single nucleotide substitution (C to A) in the third exon that leads to a Ser substitution with a stop codon, giving a truncated DEP2 protein.[2]
- The leaves of the cl7(t) mutant were shorter with a smaller angle between the leaves and stem, and a higher chlorophyll content than that of wild type. Compared with the wild type, the cl7(t) mutant displayed a 30% reduction in panicle length and a marked decrease in the number of grains per panicle. the grains of the mutant were wider and shorter than those of the wild type.[2]
- The cl7(t) was created by ethyl methanesulfonate mutagenesis.The mutant exhibited cleistogamy,and had closed spikelets, reduced plant height, and altered morphology of the leaves,panicle,and seeds. Further studies demonstrated that the force required to open the lemma and palea was higher in the cl7(t) mutant, and there was weak swelling ability in the lodicules, which leads to cleistogamy.[2]
- Morphology observation showed no difference between the cl7(t) mutant and wild-type plants from the vegetative developmental stage to the early reproductive stage. However, the cl7(t) mutant exhibited a remarkable phenotype change in terms of flowering. During the heading stage, the wild-type spikelets opened and the anthers protruded outside, whereas the cl7(t) spikelets remained closed. Other than cleistogamy, the architecture of the cl7(t) mutant was also significantly different from that of wild type. The plant height of the cl7(t) mutant was reduced about 20% compared with wild type. The panicles of the cl7(t) mutant remained upright at seed maturity, whereas the panicles of the wild type began to bend as the seed filling caused an increase in grain weight after flowering. The leaves of the cl7(t) mutant were shorter with a smaller angle between the leaves and stem, and a higher chlorophyll content than that of wild type. Its gross plant architecture is similar to the ideal models of japonica rice of Northern China that have high productivity (plant height dwarf, panicle length short, leaf posture erect, leaves wide and short erect panicle.[2]
Labs working on this gene
- Rice Research Institute, Anhui Academy of Agricultural Sciences, Hefei 230031, PR China
- Key Laboratory of Ion Beam Bioengineering, Institute of Technical Biology and Agriculture Engineering, Chinese Academy of Sciences,Hefei 230031, PR China
- Key Laboratory of Resource Plant Biology of Anhui Province, College of Life Sciences, Huaibei Normal University, Huaibei 235000,PR China
- Institute of Agricultural Engineering, Anhui Academy of Agricultural Sciences, Hefei 230031, PR China
- College of Life Sciences, Anhui Agricultural University, Hefei 230031, Anhui, PR China
- State Key Laboratory of Plant Genomics and National Center for Plant Gene Research, Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, Beijing 100101, China
- Graduate School of the Chinese Academy of Sciences, Beijing 100101, China
- Department of Bioscience,Fukui Prefectural University,4-1-1 Matsuoka Kenjyojima,Eiheiji-cho,Yoshida-gun,Fukui 910-1195,Japan
- Institute of Society for Techno-innovation of Agricuture,Forestry and Fisheries.Tsukuba,Ibaraki 305-0854,Japan
- QTL Genomics Research Center,National Institute of Agrobiological Sciences,Tsukuba,Ibaraki 305-8602,Japan
- Bioscience and Biotechnology Center,Nagoya University,Chikusa,Nagoya 464-8604,Japan
- Key Laboratory of Ion Beam Bioengineering, Institute of Technical Biology and Agriculture Engineering, Chinese Academy of Sciences,Hefei 230031, PR China
References
<references>
[2]- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 1.7 1.8 Feng Li,Wenbo Liu,Jiuyou Tang,Jinfeng Chen,et al.(2010)Rice DENSE AND ERECT PANICLE 2 is essential for determining panicle out growth and elongation.Cell Research 20:838-849.
- ↑ 2.0 2.1 2.2 2.3 2.4 Da-Hu Ni1,Juan Li,Yong-Bo Duan1,Ya-Chun Yang,Peng-Cheng Wei,Rong-Fang Xu,Chun-Rong Li,Dan-Dan Liang, Hao Li,Feng-Shun Song,Jin-Long Ni,Li Li and Jian-Bo Yang.(2014)Identification and utilization of cleistogamy gene cl7(t) in rice (Oryza sativa L.).Journal of Experimental Botany, Vol. 65, No. 8, pp. 2107–2117.