Os01g0919900
Gene Os01g0919900,namely OsSSI2,meaning fatty-acid desaturase gene, is involved in the negative regulation of defense responses in rice, as are itsArabidopsis and soybean counterparts. [1]
Contents
Annotated Information
Function
- Fatty acids and their derivatives play important signaling roles in plant defense responses. It has been shown that suppressing a gene for stearoyl acyl carrier protein fatty-acid desaturase (SACPD) enhances the resistance of Arabidopsis (SSI2) and soybean to multiple pathogens. we present functional analyses of a rice homolog of SSI2(OsSSI2) in disease resistance of rice plants.[1]
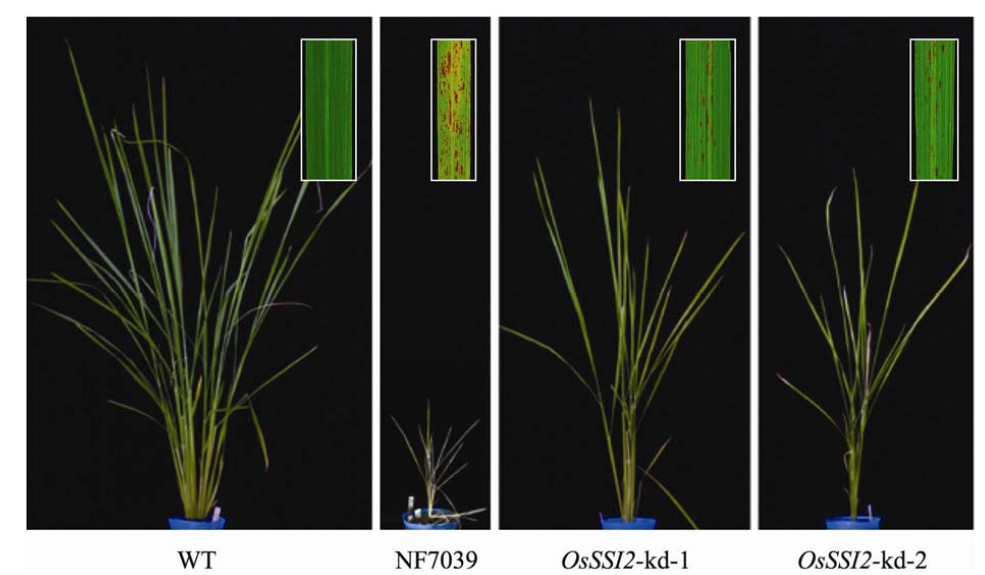
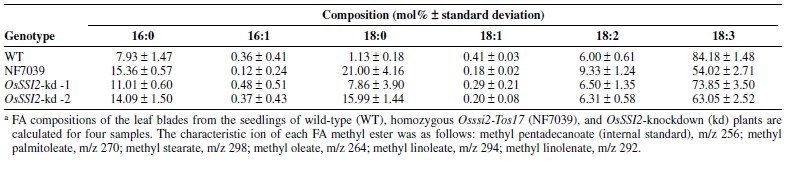
- Downregulation of OsSSI2 affects plant growth and FA profiles.To characterize the loss-of-function effects of OsSSI2, we identified and obtained two lines of Tos17 insertion mutants for OsSSI2 (Osssi2-Tos17: NF7039 and NF8001) from the Rice Genome Resource Center, Japan [2][3]. The NF7039 and NF8001 lines had Tos17 insertions in exon 2 and intron 1 of OsSSI2 , respectively.OsSSI2 transcripts of normal size were not observed on an RNA blot in homozygous NF7039 plants. Instead, a band for large-sized fusion transcripts that contained OsSSI2 with Tos17 sequences was detected, suggesting that this line is a null mutant. On the other hand, nearly normal levels andsizes of OsSSI2 transcripts were detected in homozygous NF8001 plants (data not shown); therefore, this line was not characterized further.The NF7039 plants exhibited spontaneous lesion formation in leaves and severely stunted plant growth (Fig. 2). Most mature leaves died of precocious senescence, leaving only one or two youngest expanded leaves alive (Fig. 2). They had very low fertility; consequently, it was difficult to use these plants for functional analyses of OsSSI2 in rice disease resistance. As an alternative, we generated transgenic ricelines for RNAi-mediated OsSSI2-kd. We obtained 14 independent lines of OsSSI2-kd rice with barely detectable OsSSI2 transcripts in the RNA blot analysis (Fig. 3). Most of these lines showed growth and developmental phenotypes very similar to those of NF7039 (data not shown); however, two lines (i.e., OsSSI2-kd-1 and OsSSI2-kd-2) showed moderate phenotypes with grain sets approximately 60 to 70% of wild type (Fig. 2). Therefore, these OsSSI2-kd lineswere mainly used for the following experiments. Expression of the two closest OsSSI2 homologs was unaffected in both the NF7039 and OsSSI2-kd lines (Fig. 3), indicating that the phenotypes observed in these lines are specifically due to OsSSI2 downregulation.Determination of the FA composition revealed a significant reduction in the 18:1 level accompanied by a large increase in18:0 accumulation in the OsSSI2-Tos17 (NF7039) and OsSSI2-kd lines compared with the wild-type plants (Table 1). Some changes were observed in the levels of C16 FA, particularly in the NF7039 line, but the changes in the 16:0 levels were rather smaller than those in C18 FA. These results demonstrate that OsSSI2 is mainly responsible for desaturase activity toward C18 FA.[1]
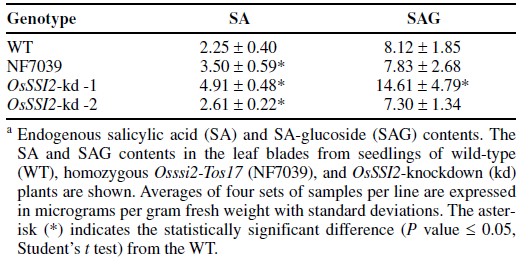
- Suppression of OsSSI2 upregulates defense-related gene expression and slightly increased free SA levels.To investigate the potential functions of OsSSI2 in the defense response of rice, we analyzed the expression of defenserelated genes in NF7039 and OsSSI2-kd plants. OsPR1b is an SA/JA-responsive gene often used as a marker gene for defense responses in rice [4]. OsPR1b was expressed at high levels in NF7039 and OsSSI2-kd plants (Fig.3), indicating that defense signaling is activated in these plants. The gene for a transcription factor, WRKY45, was also upregulated to high levels in the NF7039 and OsSSI2-kd plants (Fig. 3). Previously, we showed that WRKY45 was highly specifically induced by SA and BTH and encodes a transcription factor that plays a key role in SA/BTH-induced resistance to rice blast disease [5]. This observation suggests that the SA signaling pathway is activated in these plants.In Arabidopsis, application of 18:1 rescues ssi2 phenotypes.To examine the effects of 18:1 application on the phenotype of constitutive WRKY45 and PR1b expression in NF7039 and OsSSI2-kd plants (Fig. 3), we applied 18:1 to rice plants. We used leaf discs to infiltrate the FA from their cut ends, thereby circumventing its poor penetration through the leaf surface of rice. Gene expression analysis revealed that applying 18:1 significantly reduced the expression of WRKY45 and PR1b in NF7039 and OsSSI2-kd plants, while applying 18:0 caused no effects on the expression of both the genes (Fig. 4). These results demonstrate that the lowered level of 18:1 is the causal factor for the activation of defense responses in NF7039 and OsSSI2-kd plants. Determination of the endogenous SA content revealed slight but statistically significant (t test, P ≤ 0.05) increases in the free SA levels in NF7039 and OsSSI2-kd plants compared with the wild-type plants (Table 2). In contrast, no alterations were observed in the SA-β-glucoside (SAG) levels, with the exception of an approximately 80% increase in OsSSI2-kd-1 plants(Table 2). These results contrast with the observations made with Arabidopsis and soybean, in which mutating SSI2 or silencing of GmSACPD-A or GmSACPD-B resulted in the accumulation of several-fold higher levels of SA and SAG [6][7]. Smallness of the changes in SA levels has been reported in rice for various treatments, which is probably related to the high basal SA levels in rice plants.
- Disease resistance is enhanced in OsSSI2-kd rice.Mutation of the SSI2 gene in Arabidopsis (ssi2) enhances the resistance to multiple pathogens, including Hyaloperonospora parasitica, P. syringae pv. tomato DC3000, and Cucumber mosaic virus [7][8][9][10]. In soybean, silencing of GmSACPD-A/-B genes enhanced the resistance to P. syringae pv. glycinea and Phytophthora sojae [6].These results, taken together with the upregulation of defense genes in OsSSI2-downregulated rice, prompted us to test whether OsSSI2 downregulation affects the resistance of rice to M. grisea and X. oryzae pv. oryzae, which are the causal pathogens of rice blast and rice leaf-blight diseases, respectively.The number of blast lesions (Fig. 5A) and the length of blight lesions (Fig. 5B) were markedly reduced in OsSSI2-kd plants compared with wild-type plants. Thus, OsSSI2down regulation enhanced the resistance of rice to these two different pathogens.[1]
- Glycerol application enhances resistance in rice.Exogenous application of glycerol lowers the 18:1 levels and enhances disease resistance in Arabidopsis [11][12] and in soybean plants [6]. To examine whether glycerol causes similar effects in rice, we spray treated rice plants with 1% glycerol and examined the expression of defense-related genes and blast and leaf-blight resistances. The results showed that glycerol application induced WRKY45 and PR1b expression (Fig. 6A) and significantly enhanced the resistance to both diseases (Fig. 6B).[1]
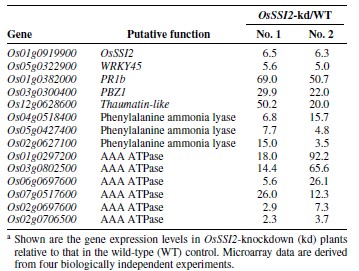
- DNA-microarray profiling of gene expression in OsSSI2-kd plants.To further characterize the genes influenced by OsSSI2 downregulation,we compared the transcript profiles of wild-type and OsSSI2-kd plants by using an oligo DNA microarray for 44,000 rice genes. Only genes that are differentially expressed in both OsSSI2-kd-1 and OsSSI2-kd-2 plants within the criteria of statistical significance (i.e., P ≤ 0.05 and false discovery rate [FDR] ≤ 5%) were selected for data analyses. More stringent filtering (e.g., P ≤ 0.01 or FDR ≤ 0.01) failed to recover some of the genes whose differential expression was detected in RNA blotting (e.g., PR1b). In total, 406 genes were found to be differentially expressed between the wild-type and OsSSI2-kd plants by a factor of more than twofold (Table 3; Supplementary Table S1). Among these, 74% (299 genes) and 26%(107 genes) were up- and downregulated, respectively, in OsSSI2-kd plants. The differentially expressed genes were classified into different functional groups, using the Kyoto Encyclopedia of Genes and Genomes (KEGG) database. Approximately 42% (169 genes) of the genes were classified as unknown,hypothetical, unclassified proteins, or no hits. OsSSI2 expression was also repressed by approximately sixfold, while no other SACPD-like genes were differentially expressed in OsSSI2-kd plants. The genes involved in “metabolism” accounted for 17.5% (n = 71) of the genes; among these, 3 genes that encoded phenylalanine ammonia lyase (PAL) were upregulated in OsSSI2-kd plants (Table 3). PAL are involved in the SA biosynthesis pathway and implicated in both biotic and abiotic plant responses [13]. Therefore, the increased free SA levels in these plants (Table 2) may be due to the upregulation of these PAL genes.The OsSSI2-regulated genes also included a large number of transcription factors (7.9%, n = 32), protein kinases (6.9%, n =28), and defense-related genes (3.9%, n = 16). WRKY45,PR1b, PBZ1, and a thaumatin-like gene were upregulated by 5-, >50-, >22-, and >20-fold, respectively, in OsSSI2-kd plants(Table 3). Among the defense-related genes, there were five genes for harpin-induced 1 domain-containing proteins and four genes for disease-resistance protein family proteins. In addition, six genes for AAA ATPase, which compose a distinct class of ATPases, were highly upregulated in OsSSI2-kd plants(Table 3).Recently, we used microarray analyses to identify approximately 2,000 BTH-responsive genes (unpublished data). In these assays, BTH was applied to the basal cut surface of the shoot instead of being sprayed on the leaves, as reported in our previous study [5]. This led to a more efficient BTH response. Comparison of these genes with the list of OsSSI2-regulated genes revealed that approximately 39%(n = 156) of the OsSSI2-regulated genes were common to both lists, and all but 7 genes showed expression changes in the same direction. These results support the notion that OsSSI2 negatively regulates the SA-signaling pathway in rice. However,a number of OsSSI2-regulated genes were BTH unresponsive,suggesting that OsSSI2 also regulates a signaling pathway or pathways other than the SA signaling pathway. In summary, we have shown that OsSSI2 encodes an FA desaturase and negatively regulates the defense responses in rice partly through suppressing SA-responsive genes. Transcript profiling suggested that OsSSI2 also regulates SA-independent signaling pathways, which may also play a role in the defense mechanism (Fig. 7). Taken together with the results obtained from Arabidopsis and soybean, our results suggest that SSI2 is functionally well conserved in defense signaling pathways across plant species.[1]
Expression
- A transposon insertion mutation (Osssi2-Tos17) and RNAi-mediated knockdown of OsSSI2 (OsSSI2-kd) reduced the oleic acid (18:1) level and increased that of stearic acid (18:0), indicating that OsSSI2 is responsible for fatty-acid desaturase activity. These plants displayed spontaneous lesion formation in leaf blades, retarded growth, slight increase in endogenous free salicylic acid (SA) levels, and SA/benzothiadiazole (BTH)-specific inducible genes, includingWRKY45, a key regulator of SA/BTH-induced resistance, in rice. Moreover, the OsSSI2-kd plants showed markedly enhanced resistance to the blast fungus Magnaporthe grisea and leaf-blight bacteria Xanthomonas oryzae pv. oryzae. These results suggest that OsSSI2 is involved in the negative regulation of defense responses in rice, as are itsArabidopsis and soybean counterparts. [1]
Evolution
- Microarray analyses identified 406 genes that were differentially expressed (≥2-fold) in OsSSI2-kd rice plants compared with wild-type rice and, of these, approximately 39% were BTH responsive. Taken together, our results suggest that induction of SA-responsive genes, including WRKY45, is likely responsible for enhanced disease resistance in OsSSI2-kd rice plants.[1]

Fig. 1. Phylogenetic tree of stearoyl acyl carrier protein (ACP) fatty-acid desaturase (SACPD) family proteins in rice, Arabidopsis, and soybean. The protein sequences were aligned with Clustal-X software and the tree was constructed using TreeView software. The Rice Annotation Project (RAP) codes and the Arabidopsis Genome Initiative (AGI) codes are shown in the figure. The GenBank accession numbers for soybean proteins are as follows: GmSACPD-A, AAX86050; GmSACPD-B, AAX86049; and GmSACPD-C, ABM45911.
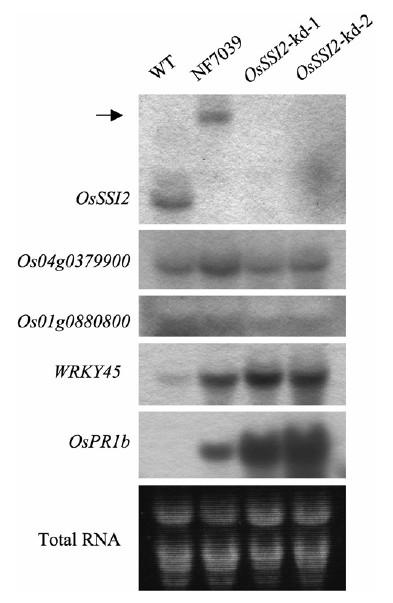
Fig. 3. RNA blot analysis. Transcripts of OsSSI2, two OsSSI2 homologs, WRKY45, and PR1b were analyzed in wild-type (WT) and OsSSI2 mutant plants. OsSSI2 transcripts were not detected in homozygous Osssi2-Tos17 (NF7039) and OsSSI2-knockdown (kd) plants, whereas the expression of the two closest homologs of OsSSI2 (i.e., Os04g0379900 and Os01g0880800) was unaffected in these plants. The arrow indicates a band of fusion transcripts encoding OsSSI2 with the Tos17 insertion. The salicylic acid/benzothiadiazole-induced gene WRKY45 and the pathogenesis-related gene OsPR1b were constitutively expressed in the NF7039 and OsSSI2-kd plants. Leaf blades from the fourth leaves of seedlings were used.

Fig. 4. Restoration of the expression of WRKY45 and PR1b after 18:1- application. Segments of leaf blades from seedlings of wild-type (WT), homozygous Osssi2-Tos17 (NF7039), and OsSSI2-knockdown (kd) plants were incubated in mock, solution containing 18:0, or solution containing 18:1 under light for 12 h at 30°C. Transcripts of WRKY45 and PR1b were analyzed by quantitative reverse-transcription polymerase chain reaction. Averages of three determinations relative to those of Rubq1 are shown with standard deviations (SD).

Fig. 5. Enhanced disease resistance in OsSSI2-knockdown (kd) plants. Wild-type (WT) and OsSSI2-kd plants were inoculated with A, the rice blast fungus Magnaporthe grisea and B, blight bacteria Xanthomonas oryzae pv. oryzae. Numbers on the y axes of the graph indicate A, the number of fungal blast lesions per 10-cm middle region of leaf blades and B, the lesion lengths of bacterial leaf blight. Each value represents the average of 15 to 20 plants. The bars indicate standard deviations (SD).

Fig. 6. Defense response induced by glycerol application. The seedlings of wild-type rice plants were sprayed with mock or glycerol (Gly). A, Induction of WRKY45 and PR1b expression by glycerol application. Transcript levels were analyzed by quantitative reverse-transcription polymerase chain reaction. Averages of three determinations relative to those of Rubq1 are shown with standard deviations (SD). B, Enhanced resistance to blast fungus Magnaporthe grisea (shown on left axis) and blight bacteria Xanthomonas oryzae pv. oryzae (shown on right axis) after glycerol application. The values on the y axes of the graph indicate A, the number of fungal blast lesions per 10-cm middle region of leaf blades and B, the lesion lengths of bacterial leaf blight. Each value represents the average of 15 to 20 plants.

Fig. 7. Proposed model for the functioning of OsSSI2 in the rice defense pathway. OsSSI2 negatively regulates the defense responses in rice partly through suppressing salicylic acid (SA)-responsive genes. OsSSI2 is also likely to regulate an SA-independent defense signaling pathway mediated by an unknown factor (X).
Labs working on this gene
- Plant Disease Resistance Research Unit, Division of Plant Science, National Institute of Agrobiological Sciences,Kannondai 2-1-2, Tsukuba, 305-8602 Japan
- College of Agriculture, Ibaraki University, Ami 300-0393, Japan
- Department of Molecular Genetics, National Institute of Agrobiological Resources, Tsukuba, 305-8602, Ibaraki, Japan
- Molecular Genetics Department, National Institute of Agrobiological Sciences, Tsukuba City, Ibaraki 305-8602, Japan
- Department of Molecular Biology, College of Natural Science, Sejong University, Seoul 143-747, Republic of Korea
- Plant Disease Resistance Research Unit, National Institute of Agrobiological Sciences, Ibaraki, Japan
- Department of Plant Pathology, University of Kentucky, Lexington, Kentucky 40546, USA
- Boyce Thompson Institute for Plant Research, Tower Road, Ithaca, NY 14853, USA
- Waksman Institute and Department of Molecular Biology and Biochemistry, Rutgers, The State University of New Jersey, 190 Frelinghuysen Road, Piscataway, NJ 08855-8020, USA
- Molecular, Cellular and Developmental Biology Program, Division of Biology, Kansas State University, 303 Ackert Hall, Manhattan, KS 66506-4901, USA
- Department of Life Science, Graduate School of Agricultural Science, Tohoku University, 1-1 Tsutsumidori-Amamiyamachi, Aoba-ku, Sendai 981-8555, Japan
- Division of Biology, and The Molecular, Cellular and Developmental Biology Program, 303 Ackert Hall, Kansas State University, 66506-4901, Manhattan, Kansas, USA
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 1.6 1.7 Chang-Jie Jiang,Masaki Shimono,Satoru Maeda,Haruhiko Inoue,Masaki Mori,Morifumi Hasegawa,Shoji Sugano,and Hiroshi Takatsuji.2009.Suppression of the Rice Fatty-Acid Desaturase Gene OsSSI2 Enhances Resistance to Blast and Leaf Blight Diseases in Rice.Molecular Plant-Microbe Interactions.22:820-829.
- ↑ Hirochika, H. 2001. Contribution of the Tos17 retrotransposon to rice functional genomics. Curr. Opin. Plant Biol. 4:118-122.
- ↑ Hirochika, H., Guiderdoni, E., An, G., Hsing, Y. I., Eun, M. Y., Han, C. D.,Upadhyaya, N., Ramachandran, S., Zhang, Q., Pereira, A., Sundaresan,V. and Leung, H. 2004. Rice mutant resources for gene discovery. Plant Mol. Biol. 54:325-334.
- ↑ Jwa, N. S., Agrawal, G. K., Tamogami, S., Yonekura, M., Han, O.,Iwahashi, H., and Rakwal, R. 2006. Role of defense/stress-related marker genes, proteins and secondary metabolites in defining rice selfdefense mechanisms. Plant Physiol. Biochem. 44:261-273.
- ↑ 5.0 5.1 Shimono, M., Sugano, S., Nakayama, A., Jiang, C. J., Ono, K., Toki, S.,and Takatsuji, H. 2007. Rice WRKY45 plays a crucial role in benzothiadiazole-inducible blast resistance. Plant Cell 19:2064-2076.
- ↑ 6.0 6.1 6.2 Kachroo, A., Fu, D. Q., Havens, W., Navarre, D., Kachroo, P., and Ghabrial, S. A. 2008. An oleic acid-mediated pathway induces constitutive defense signaling and enhanced resistance to multiple pathogens in soybean. Mol. Plant-Microbe Interact. 21:564-575.
- ↑ 7.0 7.1 Shah, J., Kachroo, P., Nandi, A., and Klessig, D. F. 2001. A recessive mutation in the Arabidopsis SSI2 gene confers SA- and NPR1-independent expression of PR genes and resistance against bacterial and oomycete pathogens. Plant J. 25:563-574.
- ↑ Kachroo, P., Shanklin, J., Shah, J., Whittle, E. J., and Klessig, D. F. 2001.A fatty acid desaturase modulates the activation of defense signaling pathways in plants. Proc. Natl. Acad. Sci. U.S.A. 98:9448-9453.
- ↑ Kachroo, A., Lapchyk, L., Fukushige, H., Hildebrand, D., Klessig, D., and Kachroo, P. 2003a. Plastidial fatty acid signaling modulates salicylic acid- and jasmonic acid-mediated defense pathways in the Arabidopsi ssi2 mutant. Plant Cell 15:2952-2965.
- ↑ Sekine, K. T., Nandi, A., Ishihara, T., Hase, S., Ikegami, M., Shah, J., and Takahashi, H. 2004. Enhanced resistance to Cucumber mosaic virus the Arabidopsis thaliana ssi2 mutant is mediated via an SA-independent mechanism. Mol. Plant-Microbe Interact. 17:623-632.
- ↑ Kachroo, A., Venugopal, S. C., Lapchyk, L., Falcone, D., Hildebrand, D.,and Kachroo, P. 2004. Oleic acid levels regulated by glycerolipid metabolism modulate defense gene expression in Arabidopsis. Proc. Natl. Acad. Sci. U.S.A. 101:5152-5157.
- ↑ Kachroo, P., Venugopal, S. C., Navarre, D. A., Lapchyk, L., and Kachroo, A. 2005. Role of salicylic acid and fatty acid desaturation pathways in ssi2-mediated signaling. Plant Physiol. 139:1717-1735.
- ↑ Shah, J. 2003. The salicylic acid loop in plant defense. Curr. Opin. Plant Biol. 6:365-371.
- SACPD-like gene family in rice.A genomic database search revealed that the rice genome encodes seven SACPD-like proteins that have 46 to 86% amino acid sequence identity with Arabidopsis SSI2 (Fig. 1).Phylogenetically, these proteins fall into two subgroups. One includes Os01g0919900 and Os04g0379900, which share 86 and 82% amino acid sequence identity, respectively, with SSI2.The other contains the five remaining members (Os08g0200100,Os03g0423300, Os02g0504800, Os08g0199400, and Os01g-0880800), which share 46 to 71% amino acid sequence identity with SSI2 and are more closely related to Arabidopsis At1g43800 (SACPD6) and soybean GmSACPD-C. Prediction of subcellular targeting by using the TargetP and WoLF PSORT prediction programs suggested that Os01g0919900,Os01g0880800, and Os08g0199400 are localized in the chloplast or mitochondrion, whereas the others are present in chloroplasts. These predictions are consistent with findings that FA synthesis in plants occurs mainly in plastids.[1]However, cell biological studies are necessary for the conclusive localization of these proteins within living cells.The protein encoded by Os01g0919900 shared the highest sequence identity with Arabidopsis SSI2 (86%), referred to as OsSSI2, and it was then further characterized with a focus on its role in disease resistance in rice.
Structured Information
- ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedref2