Os02g0630300
Please input one-sentence summary here.
Contents
Annotated Information
Function
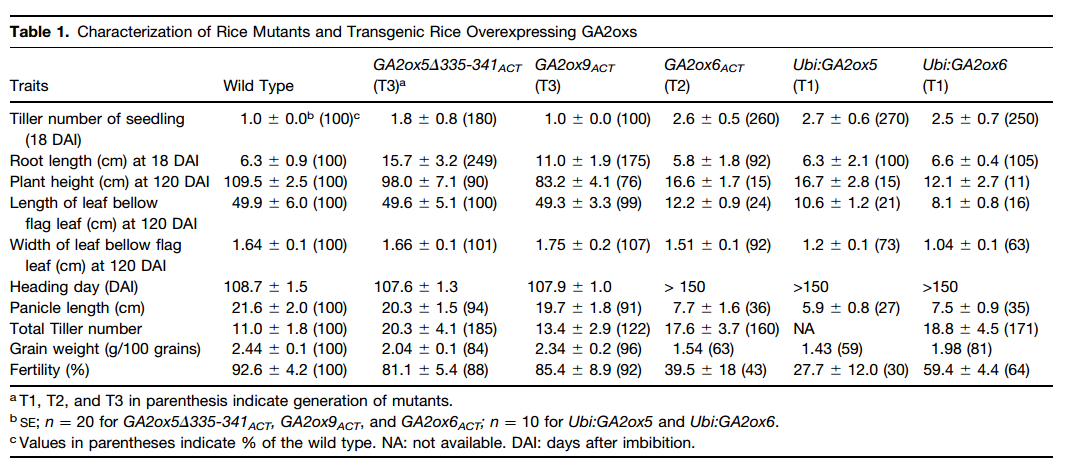
The semidwarf mutant M58817, designated as GA2ox9ACT,carries a T-DNA insertion at a position 2.4 kb upstream of the translation start codon of GA2ox9 (Figure 5D). Accumulation of GA2ox9 mRNA was significantly enhanced by T-DNA activation tagging, and the semidwarf phenotype was observed in both the homozygous and heterozygous mutants. The GA2ox9ACT homozygous mutant had an average plant height of 76% and produced seeds with an average fertility of 92% of the wild type(Table 1)GA2ox9ACT displayed a normal height but had longer roots and higher tiller numbers than the wild type (Table 1).
Expression
Genes GA2ox9 was expressed in leaves, and its expression was also temporally regulated (Figure 3B).GA2ox9 accumulation of its mRNAs in leaves was detected prior to the transition from vegetative to reproductive growth phases.Since expression of most GA2oxs terminated after the active tillering stage, the pattern of tiller growth throughout the rice life cycle was examined. Tiller number increased from 30 to 50 DAI (active tillering), remained constant until 75 DAI, and then increased again until 90 DAI (late tillering)when the experiment was terminated (Figure 3C). Expression of each group of GA2oxs paralleled the active and late tillering stages (cf. Figures 3B with 3C). Germination was observed from 1 DAI and reached almost 100% at 2 DAI (Figure 4A). The accumulation of most GA2ox mRNAs was detectable starting from 0 to 1 DAI and maintained at similar levels afterward, except that of GA2ox5 and GA2ox9 was moderately reduced at 2 DAI (Figure 4B).

Evolution

Labs working on this gene
- Institute of Molecular Biology, National Chung-Hsing University, Taichung 402, Taiwan, Republic of China
- Institute of Molecular Biology, Academia Sinica, Taipei 115, Taiwan, Republic of China
- Institute of Plant and Microbial Biology, Academia Sinica, Taipei 115, Taiwan, Republic of China
- Department of Energy Plant Research Laboratory and Department of Plant Biology, Michigan State University,East Lansing, Michigan 48824-1312
References
Please input cited references here.