Os04g0415000
Please input one-sentence summary here.
Contents
Annotated Information
Function
To isolate a rice transferred DNA (T-DNA) insertion mutant, BY240, which exhibited an abaxial leaf curling phenotype that co-segregated with the inserted T-DNA. The T-DNA was inserted in the promoter of a novel gene, ACL1 (Abaxially Curled Leaf 1), and led to overexpression of this gene in BY240. Overexpression of ACL1 in wild-type rice also resulted in abaxial leaf curling. ACL1 encodes a protein of 116 amino acids with no known conserved functional domains. Overexpression of ACL2, the only homolog of ACL1 in rice, also induced abaxial leaf curling. RT–PCR analysis revealed high expressions of ACLs in leaf sheaths and leaf blades, suggesting a role for these genes in leaf development. In situ hybridization revealed non-tissue-specific expression of the ACLs in the shoot apical meristem, leaf primordium, and young leaf. Histological analysis showed increased number and exaggeration of bulliform cells and expansion of epidermal cells in the leaves of BY240, which caused developmental discoordination of the abaxial and adaxial sides, resulting in abaxially curled leaves.
Expression
Expression Patterns of ACLs in the Wild-Type and BY240 Implied their Function in Leaf Development
The expression patterns of the ACLs were analyzed to test whether they function in leaf development. In the wild-type, the expression of ACL1 was barely detectable, while in BY240, it was greatly increased. In contrast, the expression of ACL2 showed no difference in BY240 and the wild-type (Figure 4A). The expression profiles of the ACLs in the wild-type and BY240 was examined by RT–PCR using various tissues, including 14-day-old green seedling, mature leaf, sheath, stem, spike, and root. In the wild-type, although ACL1 and ACL2 expressed in almost all the tissues tested, the highest expression was in the leaf blade and the leaf sheaths, indicating that they might function in leaf development. In addition, in all the tissues examined, ACL1 was significantly up-regulated in BY240, while ACL2 was not (Figure 4B).
Furthermore, we characterized the expression patterns of ACLs during leaf development. Single leaf development can be divided into seven stages (Itoh et al., 2005), among which the P4 stage indicates elongation of the leaf blade, when the leaf is still enwrapped in the previous leaf sheath. At the early P4 stage, the leaf blade elongates to half its length and at the late P4 stage, the leaf blade is totally elongated but is still enwrapped in the previous leaf sheath. The P5 stage indicates elongation of the leaf sheath; at the early P5 stage, the leaf blade begins to emerge from the sheath of the previous leaf, and at the late P5 stage, the leaf blade totally emerges. The P6 stage represents the mature leaf. We examined early P4 leaves, l ate P4 leaves, 1/4 P5 leaves, 1/2 P5 leaves, and P6 leaves of the wild-type to examine expression of ACL1 and ACL2. Expressions of both ACL1 and ACL2 increased as development progressed in the leaves. At the 1/2 P5 stage, expressions of ACL1 and ACL2 reached their peaks, and these peaks were maintained in the following stages. Furthermore, we examined expressions of ACL1 and ACL2 in leaves of different developmental stages in BY240. In addition to the gradually elevated expression trends as development progressed, ACL1 was overexpressed in all the corresponding stages in BY240, while ACL2 was not (Figure 4C).
To characterize the expression pattern of ACL1 and ACL2, we carried out in situ hybridization in the areas around the shoot apical meristem (SAM) of five-leaf-stage wild-type ZH11. ACL1 expressed in the SAM, leaf primordium, and young leaf (Figure 4D); no difference in expression was detected in the abaxial and adaxial leaf blades. ACL2 showed a similar non-tissue-specific and non-polarized expression pattern to that of ACL1 (Figure 4F).
In summary, the T-DNA inserted gene ACL1 was overexpressed in mutant BY240. Furthermore, the expression patterns of the ACL1 and ACL2 genes in the wild-type were very similar, showing neither tissue specificity nor polarity, and were mainly expressed in the SAM, leaf primordium, and young leaf. Their highest expressions were in the leaf blade and leaf sheath, and their expressions increased as development progressed, which indicated that they might function in leaf development.

Figure 4. ACL1 Was Overexpressed in BY240, and Expression Pattern of ACL1 and ACL2.
(A) Expression of ACL1 and ACL2 in mature leaves of ZH11 and BY240 by RNA gel blot analysis. ACL1 was significantly overexpressed in BY240, while ACL2 was not.
(B) Expression profile of ACLs in ZH11 and BY240. S, seedling; LB, mature leaf blade; LS, mature leaf sheath; C, culm; P, panicle; R, root. The PCR process was carried out 25 cycles for actin, 35 cycles for ACL1, and 28 cycles for ACL2 (sampling was repeated three times and the RT–PCR processes were repeated at least three times).
(C) Gradually increased expression of ACL1 and ACL2 during leaf development. 1–5, expression of ACL1 and ACL2 in leaves of different developmental stages of ZH11, 6–10, expression of ACL1 and ACL2 in leaves of different developmental stages of BY240. 1 and 6: early P4 leaf (still enwrapped in the sheath, with the tip reaching 1/2 height of the sheath), 2 and 7: late P4 leaf (still enwrapped in the sheath, only the tip visible), 3 and 8: 1/4 P5 leaf, 4 and 9: 1/2 P5 leaf, and 5 and 10: P6 leaves (mature leaves). The PCR process was carried out 25 cycles for actin, 35 cycles for ACL1, and 28 cycles for ACL2 (sampling was repeated three times and RT–PCR processes were repeated at least three times).
(D) In situ hybridization of the ACL1 gene in SAMs of ZH11, antisense probe.
(E) In situ hybridization of the ACL1 gene in SAMs of ZH11, sense probe.
(F) In situ hybridization of the ACL2 gene in SAMs of ZH11, antisense probe.
(G) In situ hybridization of the ACL2 gene in SAMs of ZH11, sense probe. Bar in (G) = 100 μm and bars were the same for (D)–(G).
Down-Regulation of ACLs Caused No Abnormality in the Leaf
To further study the function of ACL1 and ACL2, we carried out RNAi analysis. First, an anti-ACL1 construct was transformed into the wild-type to knock down the expression of ACL1 (Figure 2C). Among the 15 T0 transformants obtained, three showed successful down-regulation of ACL1 (Figure 7A). However, no obvious phenotypic change was observed in the T0 or the succeeding generations of these transformants. An anti-ACL2 vector was constructed and transformed into the wild-type; 16 T0 transformants were obtained, although ACL2 was evidently knocked down in some of the transformants (such as No. 3 and 8 in Figure 7B). No defects in the leaves were observed either (Figure 7B).
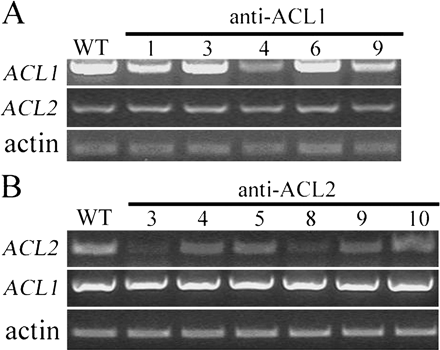
Figure 7. Examination of Antisense Transformants of ACLs.
(A) RT–PCR analysis of ACL1 and ACL2 expression in five anti-sense ACL1 transformants. RT–PCR was processed 35 cycles for ACL1, 28 cycles for ACL2, and 25 cycles for actin.
(B) RT–PCR analysis of ACL2 and ACL1 expression in six antisense ACL2 transformants. RT–PCR was processed 28 cycles for ACL2, 35 cycles for ACL1, and 25 cycles for actin.
Evolution
To clone the flanking sequence of the T-DNA in BY240, TAIL–PCR was conducted. To verify the inserting region, primers B4LI and B4RI spanning this region were designed to respectively pair with primers on the T-DNA, namely B4LI coupling with TL and B4RI with TR. In BY240, T-DNA was integrated at position 26 683 bp of PAC clone OJ991214_12 on rice chromosome 4, and the integration resulted in the deletion of a short sequence from the T-DNA region and an 8-bp deletion from the BY240 genome. Furthermore, the inserting region was in the promoter of the predicted gene Os04g33860 (www.tigr.org), 1798 bp upstream of the ATG start codon. We named this putative gene as ACL1. ACL1 comprises two exons and one intron with a full-length cDNA of 1023 bp (Accession: CT831358, Clone Name: OSIGCSN071D21, www.ncgr.ac.cn/ricd/); the first exon is non-coding and the second exon encodes the ACL1 protein of 116 amino acids. Bioinformatics analysis indicated that Os02g33330 (www.tigr.org) in the rice genome was homologous with ACL1 (57.8% protein identity), and was thereby named as ACL2. Similarly, ACL2 is composed of two exons and one intron, and its putative protein of 123 amino acids is encoded by the second exon. In the Arabidopsis genome, two homologous genes, At5g06270 and At3g11600, were found, sharing 48.4 and 38.5% identity with ACL1, respectively. In Vitis vinifera, two homolog proteins, CAO15841 and CAN79170, were identified (Figure 3). However, the function of these ACL proteins remains unknown.
Figure 3. Alignment of Amino Acid Sequences of ACL1 and its Homologies.
The amino acids showing identity were shaded black, whereas similar amino acids were shaded grey. * indicates that region of similarity/identity were from an alignment of 141–261 from CAO15841 and 161–280 from CAN79170 with the corresponding region of ACL1, respectively.
Labs working on this gene
Ling Lia,b, Zhen-Ying Shia, Lin Lia, Ge-Zhi Shenc, Xin-Qi Wangc, Lin-Sheng Ana and Jing-Liu Zhanga,1
Overexpression of ACL1 (abaxially curled leaf 1) Increased Bulliform Cells and Induced Abaxial Curling of Leaf Blades in Rice
a National Key Laboratory of Plant Molecular Genetics, Institute of Plant Physiology and Ecology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, 300 Fenglin Road, Shanghai 200032, China
b Graduate School of the Chinese Academy of Sciences, 19 Yuquan Road, Beijing 100039, China
c The Plant Breeding and Cultivation Research Institute, Shanghai Academy of Agriculture Sciences, 2900 Beidi Road, Shanghai 201106, China
To whom correspondence should be addressed. E-mail jlzhang@sippe.ac.cn, fax +86-021-54924015, tel. +86-021-54924079.
References
- ↑ Li L1, Shi ZY, Li L, Shen GZ, Wang XQ, An LS, Zhang JL. Overexpression of ACL1 (abaxially curled leaf 1) increased Bulliform cells and induced Abaxial curling of leaf blades in rice. Mol Plant. 2010 Sep;3(5):807-17.
- ↑ Alvarez JM, Rocha JF, Machado SR. Bulliform cells in Loudetiopsis chrysothrix (Nees) Conert and Tristachya leiostachya Nees (Poaceae): structure in relation to function. Braz. Arch. Bio. Technol 2008;51:113-119.
- ↑ Itoh J, Nonomura K, Ikeda K, Yamaki S, Inukai Y, Yamagishi H, Kitano H, Nagato Y. Rice plant development: from zygote to spikelet. Plant Cell Physiol 2005;46:23-47.