Os04g0645100
FLO2 plays a pivotal regulatory role in rice grain size and starch quality by affecting storage substance accumulation in the endosperm.[1]
Contents
Annotated Information
Function
Rice (Oryza sativa) endosperm accumulates a massive amount of storage starch and storage proteins during seed development. However, little is known about the regulatory system involved in the production of storage substances. The rice flo2 mutation resulted in reduced grain size and starch quality. Map-based cloning identified FLOURY ENDOSPERM2 (FLO2), a member of a novel gene family conserved in plants, as the gene responsible for the rice flo2 mutation. FLO2 harbors a tetratricopeptide repeat motif, considered to mediate a protein-protein interactions. FLO2 was abundantly expressed in developing seeds coincident with production of storage starch and protein, as well as in leaves, while abundant expression of its homologs was observed only in leaves. The flo2 mutation decreased expression of genes involved in production of storage starch and storage proteins in the endosperm. Differences between cultivars in their responsiveness of FLO2 expression during high-temperature stress indicated that FLO2 may be involved in heat tolerance during seed development. Overexpression of FLO2 enlarged the size of grains significantly. These results suggest that FLO2 plays a pivotal regulatory role in rice grain size and starch quality by affecting storage substance accumulation in the endosperm.[1]
Expression and Mutation
The transcript of FLO2 is abundant in immature seeds and mature leaves, but not abundant in panicles before heading, stems, and roots.[1]
The flo2 mutants showed no apparent differences in visible features during the vegetative stage; plant height, the number of leaves, the number of tillers, and the number of panicles were similar to those of wild-type plants. No significant difference was found in the timing of flowering. The number of panicles per plant, the number of spikelets per panicle, and the ratio of ripened grains did not significantly differ from those of the wild-type plant. The flo2 grains had white and floury endosperm (Figure 1A), and scanning electron microscopy images of transverse sections of flo2 grains indicated that this endosperm was filled with loosely packed, small, and spherical starch granules with large air spaces, while the wild-type endosperm consisted of densely packed, large, and irregularly polyhedral starch granules (Figure 1B).[1]

(A) Representative seed of the wild type (Kinmaze), the flo2 mutant (EM37), vector control line (VC), and complemented line (CL). CL and VC are transformants with/without a wild-type FLO2 gene in the flo2 mutant EM37. The top panel shows the grain shapes of the seeds, and the bottom panel shows seeds illuminated with backlight. The floury phenotype (EM37 and VC) is indicated by a dark image. Wild-type and CL grains were more packed and transmitted light, whereas the flo2 mutant and VC grains were chalky and floury, resulting in dark images when illuminated with backlight. Bar = 1 cm. (B) Scanning electron microscopy images of the transverse sections of the wild-type, flo2 mutant, VC, and CL grains. Bars = 50 μm.
Grain size was significantly smaller than that of the wild type (89% on average, n > 40, P < 0.01), as were both grain weight and grain length. In the endosperm of the flo2 mutant, the amylose content was lower than that of the wild type (Figure 2A), and structural changes in amylopectin were also noted, with both the short and the long chains consisting of 9 to 21 degrees of polymerization (DP) and ≥38 DP, respectively, decreasing and the middle chains with 22 to 38 DP increasing (Figure 2B).[1]
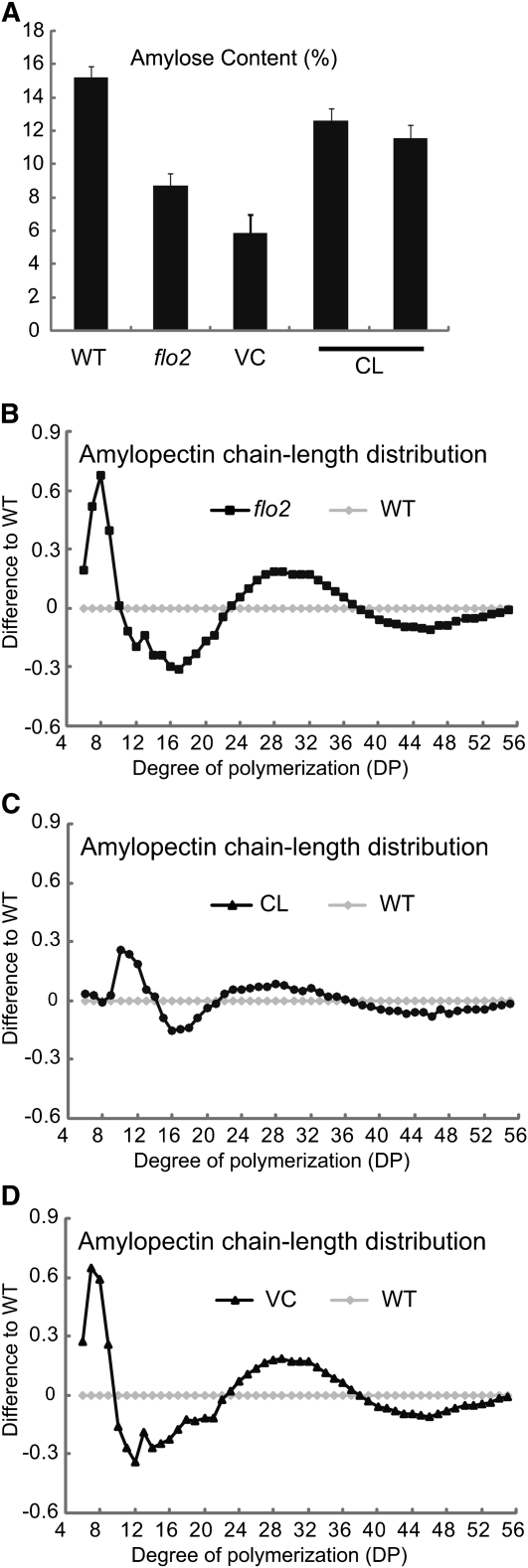
(A) Amylose content of the wild type (Kinmaze), flo2 mutant (EM37), VC, and CL. Representative VC and CL were analyzed. Error bars show sd (n = 6). (B) to (D) Comparison of the chain-length profile of amylopectins. Debranched amylopectin was compared with the wild type. Differences from the wild type in chain length distribution of amylopectins are shown. Amylopectin profile of the flo2 grain (B), CL (C), and VC (D).
Real-time quantitative RT-PCR indicated that expression levels of many genes that participated in starch biosynthesis were significantly decreased, such as genes for BEI, BEIIb, AGPases (AGPL1, AGPL2, AGPL3, AGPL4, AGPS1, AGPS2a, and AGPS2b), soluble SSs (SSI, SSIIa, SSIIc, SSIIIa, and SSIVb), GBSSI, isoamylases (ISA1 and ISA2), and pullulanase (PUL). Expression of genes for SSIIb, SSIIIb, SSIVa, GBSSII, BEIIa, and ISA3 was little decreased (Figure 3). Gene expression for sucrose synthase (Susy1 and Susy2), PPDKB, glucose-6-phosphate isomerases (PGIa and PGIb), and α-amylase (Amy3C, Amy3D, and Amy3E) was also reduced, while expression of the genes for Susy3 and Amy3B was little decreased (Figure 3). The expression of storage protein genes encoding glutelins (GluA1, GluA2, GluA3, and GluB1), globulins (Globulin1, Globulin2, 11S-globulin, and 19-kD globulin), prolamins (10 kD, 13 kD, and 17 kD), major allergenic protein genes (14 to 16 kD; RA16, RA17, RAG5B, RAG2, and RG21), and the alanine aminotransferase gene was also significantly reduced, whereas expression of GluB4 and protein disulfate isomerase was little decreased (Figure 3). In short, the flo2 mutant showed reduced expression of a large portion of the genes participating in storage starch and storage protein biosynthesis in developing rice seeds.[1]
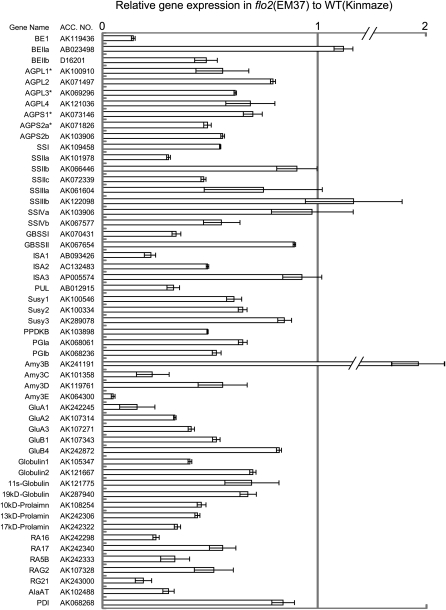
Total RNA extracted from 10 DAF developing seeds was used for real-time RT-PCR analysis. Expression of representative genes involved in storage starch production, storage proteins, and carbon metabolism in the flo2 mutant (EM37) are shown relative to the wild type (Kinmaze), which is set as 1. Each gene name is indicated by a simplified representation with the accession number of the corresponding full-length cDNA. BE1, BEIIa, and BEIIb, branching enzyme I, IIa, and IIb, respectively; AGPL (AGPL1, AGPL2, AGPL3, and AGPL4) and AGPS (AGPS1, AGPS2a, and AGPS2b), ADP glucose pyrophosphorylase large subunit and small subunit, respectively; SSI, SSIIa, SSIIb, SSIIc, SSIIIa, SSIIIb, SSIIIc, SSIVa, and SSIVb, soluble starch synthase I, IIa, IIb, IIc, IIIa, IIIb, IIIc, IVa, and IVb, respectively; GBSSI and GBSSII, granule-bound starch synthase I and II; ISA1, ISA2, and ISA3, corresponding isoamylase isozymes; PUL, pullulanase; Susy1, Susy2, and Susy3, sucrose synthase 1, 2, and 3, respectively; PPDKB, pyruvate phosphate dikinase B; PGI-a and PGI-b, glucose-6-phosphate isomerase a and b; Amy3B, Amy3C, Amy3D, and Amy3E, α-amylase 3B, 3C, 3D, and 3E, respectively; GluA1, GluA2, GluA3, GluB1, and GluB4, glutelin A1, A2, A3, B1, and B4, respectively; Globulin1, Globulin2, 11S-globulin, and 19-kD globulin, corresponding globulin species; 10-, 13-, and 17-kD prolamin, prolamins with each size; RA16, RA17, RAG5B, RAG2, and RG21, species of rice allergenic proteins; AlaAT, alanine aminotransferase; PDI, protein disulfate isomerase. Asterisks show subunits of the plastidic AGPase. Error bar shows sd (n = 3).
In the flo2 grains, reduced accumulation of glutelin, a representative storage protein, was observed (Figure 4A), suggesting decreased production of storage proteins. Protein gel blot analyses also detected an extreme reduction in the amount of BEI and of major allergenic proteins, as well as of RA16 and homologs, in the grain of the flo2 mutant (Figures 4B and 4C). These results suggest that the flo2 mutation reduced the production of storage substances in total in the rice endosperm.[1]
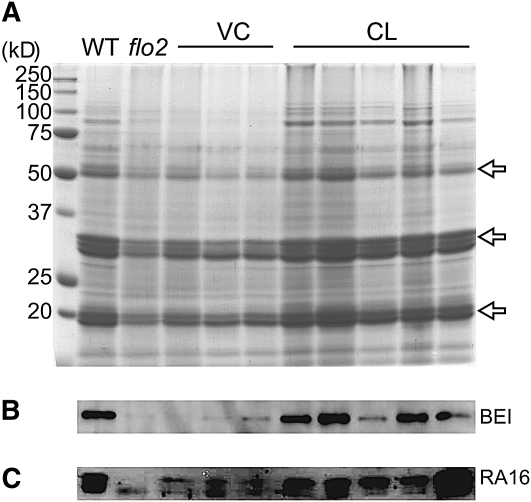
(A) Accumulation of glutelins in developing seeds of the wild type (Kinmaze), flo2 mutant (EM37), VC, and CL. Each lane contains proteins from one-hundredth of the total glutelin fraction, which was extracted from the mixture of 10 rice grains of each line. Proteins are shown by Coomassie blue staining after separation on SDS-PAGE. Arrows indicate 51-kD glutelin precursor, 30- to 36-kD α-subunits of glutelins, and 19- to 22-kD β-subunits of glutelins, respectively. Size markers are shown on the left. (B) and (C) Accumulation of BEI (B) and RA16 (C) in developing seeds 10 DAF of the wild type, flo2 mutant, VC (three independent lines), and CL (five independent lines) is shown by protein gel blot analysis using antiserum raised against BEI and RA16, respectively. The RA16 antiserum detects highly conserved 14- to 16-kD allergenic proteins with slightly different molecular mass. Each lane contains 1 μg of total protein, which was separated by SDS-PAGE.
Expression of the genes involved in storage starch production was greatly reduced in the flo2 mutant (Figure 3). The molecular structure of the storage starch accumulated in flo2 endosperm was also quite different from that of the wild type. Phenotypes similar to those of the flo2 mutant are found in several mutants of the enzymes involved in starch biosynthesis, but many differences are evident in their features. For example, the amylopectin properties of the flo2 grain largely resemble those of the mutant lacking BEI, although the endosperm of this mutant seed exhibits a normal phenotype and contains the same amount of starch as the wild type. The flo2 mutant showed a decrease of BEI protein (Figure 4), suggesting a reduction of BEI activity. Therefore, it is reasonable to state that the reduced BEI activity in the flo2 mutant greatly contributed to the aberrant structure of storage starch. The mutant of BEIIb also produces a white-core endosperm, but its amylopectin structure differs greatly from that of the flo2 mutant where the long side chains contained in its amylopectin are highly enriched in the BEIIb mutant. Amylose is mainly synthesized by the activity of GBSS, suggesting that the decrease of amylose content could be attributed to the reduced expression of GBSSI in the flo2 mutant. It has been shown that AGPase activity strongly affects the yield of grain starch. ISA1 and PUL, both of which have activities as starch debranching enzymes, are essential for the formation of the highly organized clustered structure of amylopectin. Therefore, aberrant features of storage starch in the flo2 endosperm are ascribed to the combined effect of reduced expression of these genes.[1]
The flo2 mutant also showed significantly decreased expression of storage protein genes. In addition, decreased expression was detected for several enzymes involved in metabolism (Figure 3). In particular, Susy is considered a key enzyme for the production of storage starch as well as cellulose synthesis. PPDK has been reported as the gene responsible for the rice flo4 mutant, which has a similar appearance to the flo2 mutant. PGI is important for both glycolysis and gluconeogenesis by catalyzing the reversible isomerization of glucose-6-phosphate to fructose-6-phosphate. Alanine aminotransferase is involved in the efficiency of nitrogen uptake and grain weight increase. These facts suggest that FLO2 widely affects the processes involved in storage substance accumulation in rice endosperm.[1]
There are several reports on the regulatory factors involved in the production of storage substances in endosperm. It has been suggested that there are some regulatory factors that directly or indirectly control the genes involved in the individual enzymes or genes encoding the storage proteins. In the case of direct regulation of gene expression, specific regulatory factors, which interact with the specific nucleotide sequences in the promoter region of the target gene, are proposed.[1]
FLO2 has a TPR motif composed of tandem repeats of 34 amino acid residues. The TPR motif adopts a helix-turn-helix structure and may mediate protein–protein interactions. We detected two candidate proteins, an LEA protein and a bHLH protein, that interact with FLO2 (Table 1, Figures 5 and 6); however, our results indicate that neither protein interacts with the TPR motif region of FLO2, suggesting that additional interactors may remain to be identified. The LEA proteins, constituting a large gene family in rice, and bHLH proteins are involved in many functions, including a multiple stress response through transcriptional regulation, although their individual functions remain unclear. The flo2 mutant showed reduced expression of multiple genes involved in storage starch and storage proteins (Figures 3 and 4). Therefore, it is suggested that FLO2 might interact with the bHLH protein in a transcriptional complex that could regulate storage starch and storage proteins directly or indirectly through influencing the development of endosperm in rice.[1]
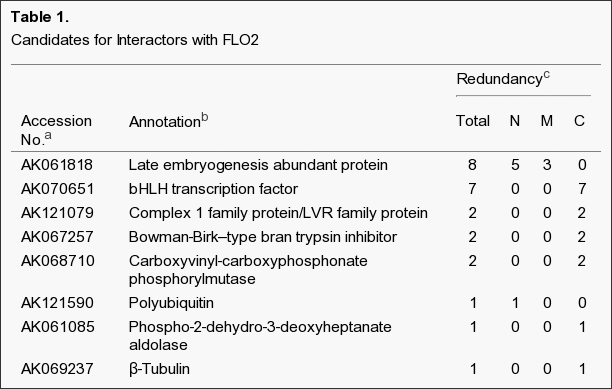
The predicted gene products were listed as the candidate clones by the Y2H screening. a: The corresponding full-length cDNA in the database, in which the isolated cDNA was contained. b: Gene names identified from the database or by similarity. c: Number of clones contained in the pool of candidates isolated from Y2H screening using each part of truncated FLO2 corresponding to FLO2(1–898), FLO2(535–1189), and FLO2(900–1720), which are shown by N, M, and C, respectively. The total number of these clones on each candidate is shown in the column “Total.”
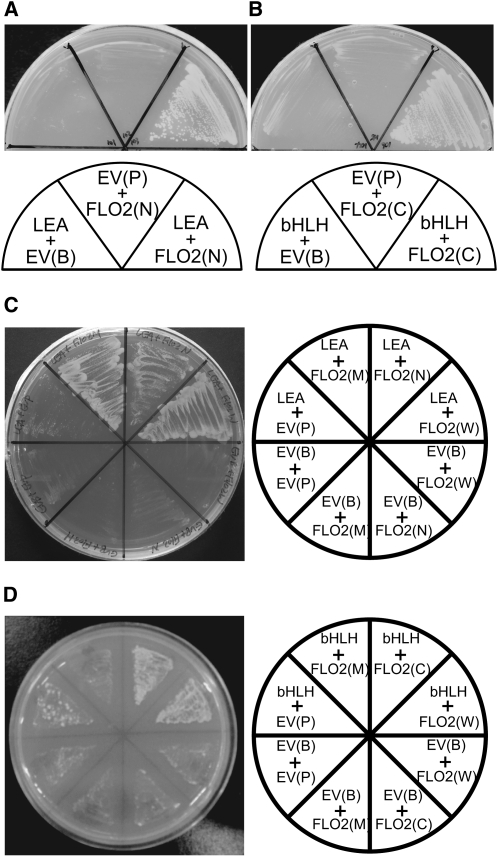
(A) and (B) Y2H analysis of FLO2 with a LEA protein (accession number AK061818[[1]]) (A) and a bHLH protein (accession number AK070651[[2]]) (B). LEA, bHLH, FLO2(N), and FLO2(C) indicate yeast cells containing plasmids for expression of the LEA, bHLH, FLO2(1–898), and FLO2(900–1720), respectively. EV(B) and EV(P) show those containing the empty bait and prey vectors instead of the corresponding genes, respectively. (C) and (D) Reciprocal swapping using LEA and FLO2 (C) and bHLH and FLO2 (D). FLO2(W) and FLO2(M) indicate yeast cells containing plasmids for expression of the full-length FLO2 and FLO2(535–1189), respectively. These cells were grown on agar plates lacking Trp, Leu, and His in the medium.
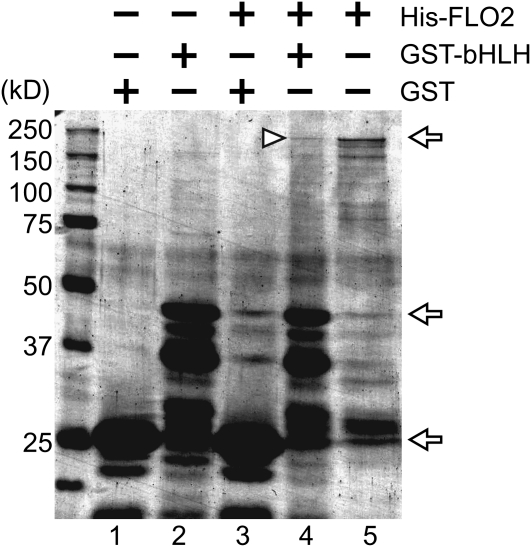
Proteins on SDS-PAGE were detected by Coomassie blue staining. Lanes 1 to 5 indicate GST, GST-fused bHLH, 6xHis tagged FLO2 pulled down with GST, 6xHis tagged FLO2 pulled down with GST-fused bHLH, and 6xHis-tagged FLO2 without GST-fused bHLH, respectively. Arrows indicate the size of 6xHis-tagged FLO2 (215 kD), GST-fused bHLH (48 kD), and GST (25 kD). The triangle indicates the predicted position of the 6xHis-tagged FLO2 bound with GST-fused bHLH. Size markers and their molecular sizes are shown on the left.
In the flo2 mutant, we detected little alteration in the expression of several genes participating in storage starch and storage protein biosynthesis, such as BEIIa, GBSSII, and GluB4 (Figure 3). This result indicates that several genes involved in production of the storage substance may not be regulated by FLO2. Expression profiles of genes involved in starch biosynthesis have been analyzed. Expression patterns of the genes participating in accumulation of storage starch differ depending on the individual genes; GBSSI, SSI, SSIIa, SSIIIa, SSIVb, BEI, BEIIb, AGPL2, and AGPS2b genes are specifically expressed in the endosperm of developing rice seeds at 10 to 20 DAF, whereas GBSSII, SSIIb, SSIIc, SSIVa, BEIIa, AGPL1, and AGPS1 genes are expressed in leaves as well as developing seeds. We observed that expression of all genes in the former group was significantly reduced in the flo2 mutants, whereas little decrease in expression levels was shown in most genes of the latter group (Figure 3). It has been reported that the patterns of gene expression in seeds are tissue and developmental stage specific. Among the genes working in the endosperm, GBSSI, SSIIa, SSIIIa, BEI, BEIIb, AGPS2b, AGPL2, ISA1, and PUL genes, which have transcripts that are low at onset and then rise steeply at the start of starch synthesis in the endosperm, are thought to play essential roles in endosperm starch synthesis at the middle stage of seed development. These genes corresponded to the genes specifically expressed in the endosperm of developing seeds reported by [2]. Our observation revealed that FLO2 expression was highly elevated at 15 to 25 DAF of seed development (Figure 7). Consequently, it is suggested that FLO2 selectively affects the expression of these genes, whose expression patterns were consistent with that of the FLO2 gene.[1]

(A) and (B) Expression of FLO2 in Kinmaze (A) and Nipponbare (B) during seed development at 5, 7, 10, 15, 20, and 25 DAF. Closed box, the wild type (Kinmaze and Nipponbare, respectively) under the normal temperature condition; hatched box, the wild type treated with high-temperature stress after 5 DAF. Amounts of transcripts are shown normalized to Actin I, which is set at 1. Error bars show sd (n = 3). (C) and (D) Expression levels of FLL1 (C) and FLL2 (D) during seed development at 3, 4, 5, 7, 10, and 15 DAF. Closed box, the wild type (Kinmaze) under the normal temperature conditions; hatched box, the wild type treated with high-temperature stress after 5 DAF. Amounts of transcripts are shown normalized to Actin I, which is set at 1. Error bars show sd (n = 3).
It has been reported that high-temperature stress during seed development apparently reduces the accumulation of storage starch in rice grains. In the case of Nipponbare, which is a rice cultivar sensitive to high-temperature stress, high temperature during seed development resulted in significant reduction of FLO2 expression, while FLO2 was expressed most abundantly under normal ripening conditions. The timing of elevation of FLO2 expression seemed to be delayed for several days in response to high-temperature stress (Figure 7B). This result suggests that FLO2 expression was apparently repressed by high-temperature stress in this cultivar. On the other hand, in Kinmaze, which is relatively tolerant to high-temperature stress, it seemed that FLO2 expression was upregulated by high-temperature stress (Figure 7A). These facts support the idea that FLO2 plays an important role in the trait for tolerance to high-temperature stress.[1]
FLO2 was abundantly expressed in immature seeds (Figures 7 and 8 ), and its expression pattern was similar to that of genes involved in the production of storage starch and storage proteins. This temporal expression may account for the appearance of the loss-of-function phenotype observed in the endosperm of the flo2 mutant seeds, such as reduced production of these storage substances. While the flo2 mutant produced smaller grains, the OX lines had larger seeds than those of the wild type (Figures 2 and 9), exhibiting the gain-of-function phenotype of the FLO2 gene. The increase in gene expression for enzymes involved in starch biosynthesis, however, is not directly linked to the level of FLO2 expression. As a matter of fact, the increases in some of the parameters of grain size measured are barely significant (Figure 9). This could be attributed to the fact that expression of the analyzed genes was measured at a single time point and that the impact of FLO2 may not coincide with its expression level at that time. This work suggests that FLO2 has a pivotal role in the accumulation of storage substances in the endosperm and sets the foundation for the understanding of grain size and quality regulation in rice.[1]
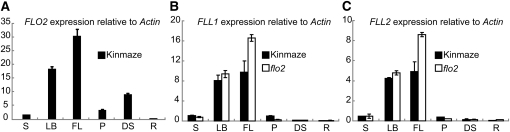
(A) Expression of FLO2 in the stem (S), leaf blade (LB), flag leaf (FL), panicles before heading (P), 10 DAF developing seed (DS), and root (R). Amounts of transcripts are shown as the relative values to those of Actin I. Error bar shows sd (n = 3). (B) and (C) Expression level of FLL1 (B) and FLL2 (C) in the stem (S), leaf blade (LB), flag leaf (FL), panicles before heading (P), 10 DAF developing seed (DS), and root (R). Closed box, wild type (Kinmaze); open box, flo2 mutant (EM37). Amounts of transcripts are shown as the relative values to those of Actin I. Error bar shows sd (n = 3).

(A) Expression level of FLO2 in the developing seed at 20 DAF of the flo2 mutant (EM37), the wild type (Kinmaze), a representative CL, and three overexpression lines (OX) in which the wild-type FLO2 gene including its own promoter was introduced in the flo2 mutant. The number of each OX line is indicated. The amount of the FLO2 transcript was determined by real-time RT-PCR and shown as relative values of the FLO2 transcript to those of rice Actin I. Error bar shows sd (n = 3). (B) Shapes of the grains of the wild type and OX (line 2). The top panel indicates the pictures of grains. The bottom panel shows the grains of the wild type (Kinmaze) and the OX line from a style side. Bars = 0.1 cm. (C) to (F) Averaged grain length (C), grain width (D), grain depth (E), and grain weight (F) of these lines. Error bar shows sd (n = 20). (G) Expression levels of representative genes in the flo2 mutant, wild-type, CL, and OX lines. Total RNA extracted from 10 DAF developing seeds was used for real-time RT-PCR analysis. GluA1, glutelin A1; RA16, a 16-kD rice allergenic protein. Expression levels of the genes are shown normalized to the wild type, which is set at 1. Error bar shows sd (n = 3).
Evolution
In the rice genome FLO2 have two homologous genes, designated FLO2-LIKE1 (FLL1) (Os02g0255700) and FLL2 (Os07g0422000). The predicted proteins encoded by FLL1 and FLL2 showed 59.8 and 52.7% similarity in amino acid sequence, respectively, to FLO2. Orthologous genes to FLO2 and its homologs were also found in other plant genomes, such as in wheat, sorghum (Sorghum bicolor), poplar (Populus spp), grape (Vitis vinifera), castor bean (Ricinus communis), Arabidopsis thaliana, and moss (Physcomitrella patens) (Figure 10), but not in animals or yeast. These homologs in higher plants were classified into three groups, which contained FLO2, FLL1, and FLL2, respectively, whereas homologs of moss diverged from this group. These results suggest that FLO2 constitutes a novel conserved gene family in plants.[1]
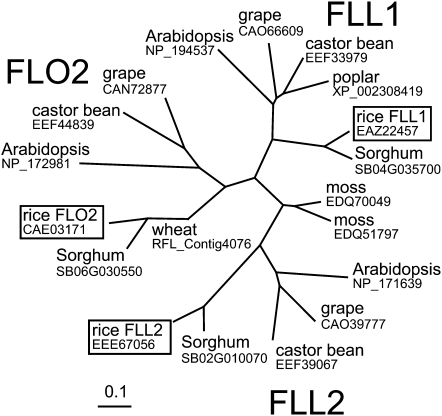
Proteins are shown in the figure as the names of plant species with the corresponding accession numbers registered in protein databases [3] and [4]. Subfamilies including FLO2, FLL1, and FLL2 are indicated. Rice genes are boxed. Scale represents the number of differences between sequences.
The FLO2 gene belongs to a gene family that consists of FLO2, FLL1, and FLL2 in the rice genome. Orthologs of these genes widely exist in plant species, including mosses, but none was found in yeast or animals. The phylogenetic tree indicates that there are three major clades for higher plants, each of which consists of the orthologs of FLO2, FLL1, and FLL2 (Figure 10). The interspecies similarity between members is larger than the similarities among FLO2, FLL1, and FLL2 within a species. These facts suggest that this gene family was acquired in an ancestor plant and diverged within the plant kingdom. The FLO2 homologs in moss (P. patens) form an independent branch in the phylogenetic tree, suggesting that the moss FLO2 family members may have evolved independently.[1]
FLL1 and FLL2 have similar protein structures to FLO2, and presumably have similar functions, but have somewhat different expression patterns. FLO2 was expressed in organs such as leaves and developing seeds (Figure 8A) and is therefore assumed to function in these organs. However, the flo2 mutant showed no apparent phenotype other than in seeds. In both the wild type and the flo2 mutant, a normal level of expression of both FLL1 and FLL2 was observed in leaves, whereas they were poorly expressed in developing seeds. It is presumed that the function of FLO2 is specific to developing seeds.[1]
Labs working on this gene
- Department of Biological Science and Technology, Tokyo University of Science, Noda, Japan
- National Agricultural Research Center, Joetsu 943-0193, Japan
- Institute for Chemical Research, Kyoto University, Uji 611-0011, Japan
- National Institute of Agrobiological Sciences, Tsukuba 305-8602, Japan
- STAFF Institute, Tsukuba 305-0854, Japan
- Institute of Genetic Resources, Faculty of Agriculture, Kyushu University, Fukuoka 812-8581, Japan
References
- ↑ 1.00 1.01 1.02 1.03 1.04 1.05 1.06 1.07 1.08 1.09 1.10 1.11 1.12 1.13 1.14 1.15 1.16 1.17 1.18 1.19 1.20 1.21 1.22 1.23 1.24 1.25 1.26 1.27 She KC; Kusano H; Koizumi K; Yamakawa H; Hakata M; Imamura T; Fukuda M; Naito N; Tsurumaki Y; Yaeshima M; Tsuge T; Matsumoto K; Kudoh M; Itoh E; Kikuchi S; Kishimoto N; Yazaki J; Ando T; Yano M; Aoyama T; Sasaki T; Satoh H; Shimada H.Plant Cell; 22(10): 3280-94, 2010 Oct.
- ↑ Hirose T., Ohdan T., Nakamura Y., Terao T. (2006). Expression profiling of genes related to starch synthesis in rice leaf sheaths during the heading period. Physiol. Plant. 128: 425–435.