Os04g0653000
Please input one-sentence summary here.
Contents
Annotated Information
Function
Please input function information here.
- EG2(extra glume 2)is a JA signalling repressor that interacts with a putative JA receptor, OsCOI1b, to trigger OsJAZ1’s degradation during spikelet development.
- Its high sequence similarity with the Arabidopsis JAZ proteins so also called OsJAZ1.
- It belong to TIFY family so also called TIFY family.
Expression
Please input expression information here.
EG2 expressed ubiquitously in the plant, from roots, stems and leaves, to spikelets and seeds.
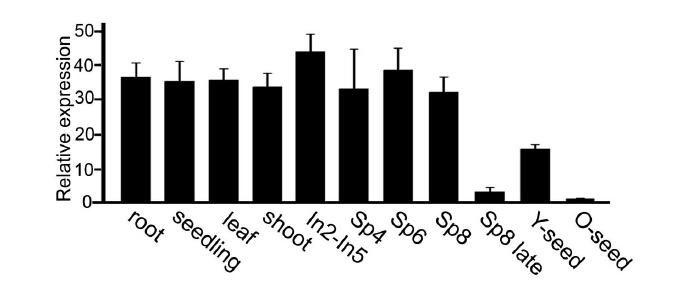
Mutation and Phenotype
Please input information here.
The EG2 gene was mapped to chromosome 4 between CQQ2 and CQQ3, two In-Del markers that define a 2.2 cM.
The mutation of EG2 is eg2-1D, which caused by C-to-G transition, resulting in a substitution of Ala by Gly in the fifth amino acid of the Jas domain. This motif was known to be critical for JAZ’s interaction with the JA receptor COI1.
mutants displayed defects in spikelet development,including extra glume-like structures between the sterile lemma and lemma and altered floral organ numbers and identities.
EG2 is a repressor of JA response
Please input information here.
yeast two-hybrid (Y2H) assays show OsJAZ1 can interact with OsCOI1b, and the interaction was dependent on JA-Ile’s mimic, coronatine(COR), but not JA or MeJA. No interaction was observed between OsJAZ1 and OsCOI1a or OsCOI1c.
Furthermore, the Ala-to-Gly mutation in the Jas domain of OsJAZ1 in eg2-1D blocked the interact between OsJAZ1 and JOsCOI1b, suggesting that the Jas domain is required for the interaction between OsJAZ1 and OsCOI1b. because osjaz1-1D can't interact with OsCOI1b, so it can't be JA-dependent proteasome-mediated degradation.
JA regulates early spikelet development possibly by activating the expression of OsMADS1 through OsMYC2, which is directly regulates OsMADS1 expression.And, OsJAZ1 interacts with OsMYC2 and suppresses the activity of OsMYC2 in triggering the transcription of OsMADS1.Both OsJAZ1 and osjaz1-1D interacted with OsMYC2, and the interaction required the Jas domain.
Inconclude,eg2-1D can't interact with OsCOI1b, so it not degradation, but it also can interact with OsMYC2 to block OsMYC2 regulates the OsMADS1 expression.
Extend knowledge
Please input information here.
The TIFY family is a novel plant-specific gene family involved in the regulation of diverse plant-specific biologic processes. This report identifies 20 TIFY genes in rice, On the basis of their protein structures, members of the TIFY family can be divided into two groups. Almost all the OsTIFY genes were responsive to one or more abiotic stresses including drought, salinity, and low temperature.Suggesting that the OsTIFY family may have important roles in response to abiotic stresses.
To analyze the evolutionary relationships of the OsTIFY genes, a phylogenetic tree was generated using the sequence alignments of 38 full-length TIFY proteins, including 20 from rice and 18 from Arabidopsis thaliana. The result show in the picture.
Labs working on this gene
Please input related labs here.
- State Key Laboratory of Hybrid Rice, School of Life Sciences and Biotechnology, Shanghai Jiao Tong University
- Jiangsu Key Laboratory for Eco-Agricultural Biotechnology around Hongze Lake, Huaiyin Normal University
- Department of Energy Plant Research Laboratory, Michigan State University
- National Key Laboratory of Crop Genetic Improvement,National Center of Plant Gene Research,Huazhong Agricultural University
References
Please input cited references here.
- Qiang Cai, Zheng Yuan, Mingjiao Chen, et al. Jasmonic acid regulates spikelet development in rice. NATURE COMMUNICATIONS, 2014
- Haiyan Ye, Hao Du, Ning Tang, et al. Identification and expression profiling analysis of TIFY family genes involved in stress and phytohormone responses in rice. Plant Mol Biol, 2009







