Os05g0111300
Please input one-sentence summary here.
Contents
Annotated Information
Function
Os05g0111300 gene was named OsMT2b,have been reported in several articles. OsMT2b transcript levels were regulated by cytokinin, and abnormal expression of OsMT2b affected the endogenous cytokinin levels in organs and tissues by the analysis of OsMT2b-RNAi and OsMT2b-overexpressing transgenic plants of rice. Considering the interrelation of OsMT2b and cytokinin as well as the phenotypes of roots and germinating embryos in transgenic plants,it indicates that OsMT2b plays important roles in initiation of lateral root and seed embryo germination.
Expression
OsMT2b in the MT Gene Family of Higher Plants
To study the molecular events of rice early embryo development, suppression subtractive hybridization was performed by using complementary DNA (cDNA) of rice embryos (indica ‘Jiayu 948’) at 5 to 7 d after pollination (DAP) as tester and cDNA of the embryos at 15 to 17 DAP as driver (J. Yuan, Y. Ren, and J.zhao, unpublished data). One clone of the subtracted cDNA library showed high similarities to the cDNA of OsMT2b (U77294), and the full cDNAclone was denominated to OsMT2bL (O. sativa METALLOTHIONEIN2bLIKE) and submitted to GenBank with accession number EF584509. The only difference of amino acid sequence between OsMT2bL and OsMT2b is Ser in OsMT2b and Gly in OsMT2bL at the position of the 21st amino acid. We chose OsMT2b for further study. Multiple sequence alignment was conducted among OsMT2b, OsMT2bL, and other proteins, including AtMT2b (NP195858) from Arabidopsis (Arabidopsis thaliana), AbMT2b (CAC40742) from Atropa belladonna, HvMTL (BAA23628) from Hordeum vulgare and ZeaMTL (CAA57676) from maize (Zea mays; Fig. 1A). Type-2 MTs contain two Cys-rich domains separated by a space of approximately 40 amino acid residues.The sequences of the N-terminal domain of this type of MT are highly conserved (MSCCGGNCGCGS), and the C-terminal domain contains three Cys-Xaa-Cys motifs (Fig. 1A). The analysis shows that homologous genes of OsMT2b exist ubiquitously in monocotyledon and dicotyledon. It suggests that this gene family may play important roles in higher plants.There are seven members of this protein family in Arabidopsis and 15 members in rice. By phylogenetic analysis, the protein members are divided into several small subgroups (Fig. 1B). Some subgroups contain both rice and Arabidopsis representatives. For instance, OsMT3a, OsMT3b, and OsMT3c are clustered with AtMT3. Similarly, OsMT4a is clustered with AtMT4a and AtMT4b. The 5#-flanking region of OsMT2b, including a region of about 961 bp upstream of the translation initiation codon ATG, is derived from the National Center for Biotechnology Information database and analyzed as a promoter (Fig. 1C). There are five ARABIDOPSIS RESPONSE REGULATOR1 (ARR1)-binding elements(ARR1AT) with core sequence 5#-AGATT-3# in the promoter region. One ARR1AT element was found in the promoter of the O. sativa NONSYMBIOTIC HAEMOGLOBIN2 (OsNSHB2) gene, and the mutation of this element abolished promoter activation in response to cytokinin (Ross et al., 2004). The presence of these elements suggests that the OsMT2b gene is probably regulated by cytokinin. The predicted transcription initiation site of OsMT2b is located at 88 bp upstream of ATG, and is designated as11 (Fig. 1C). The comparison of the full-length cDNA sequenceswith the corresponding genomic DNA sequences showed that the coding sequence (CDS) of the OsMT2b gene has three exons that are disrupted by two introns (Fig. 1C).
Temporal and Spatial Expression Patterns of OsMT2b
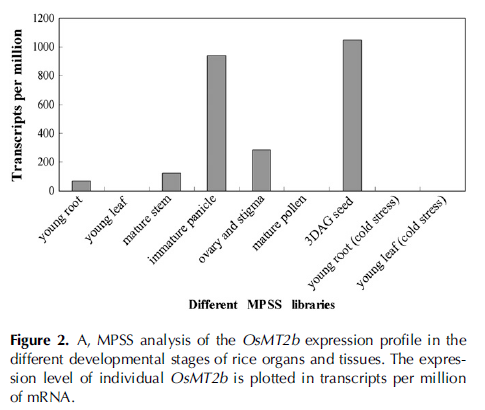
The expression levels of OsMT2b were analyzed using the Massively Parallel Signature Sequencing (MPSS) database that was constructed by Nakano et al., 2006. A practical tag, GATCTCATCATGTACTC, was available for OsMT2b after the BLAST in the MPSS database. The expression levels of OsMT2b were measured in transcripts per million of mRNA in the selected rice organs and tissues (Fig. 2A).
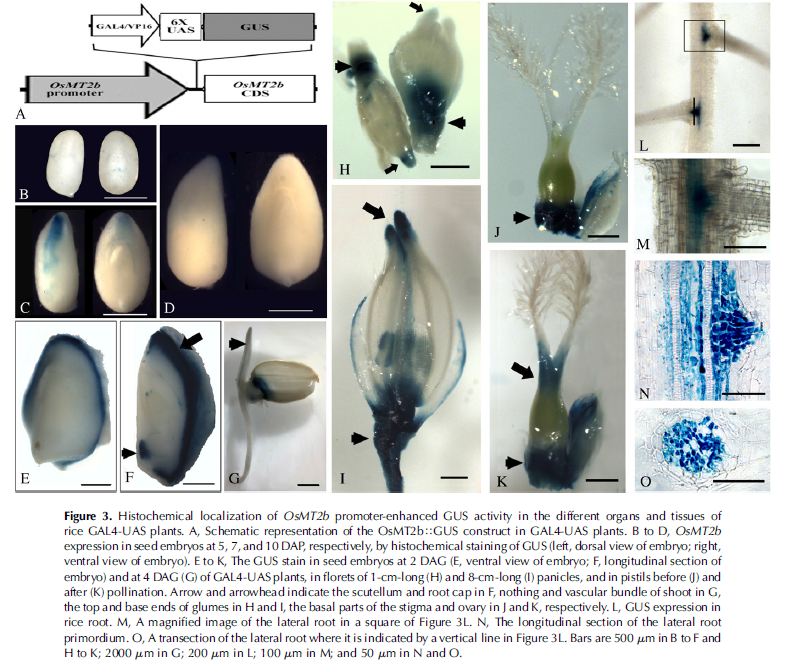
The results indicate that the expression level of OsMT2b is higher in immature panicles and seeds at 3 d after germination (DAG) than in ovaries and stigmas, young roots, and mature stems, but, hardly detected in leaves and mature pollens; its expression level reduces in the roots of 2-week-old seedlings treated at 4�C cold, but still does not change in the leaves of the same treating condition.To further study the spatial expression pattern of OsMT2b,we performed GUS staining in a T-DNAinsert mutant 03Z11AN35 that has a GAL4-UAS enhancer trap system located between the promoter and the CDS of OsMT2b (Fig. 3A). In this system, the GAL4-UAS is made up of GAL4/VP16, a fusion gene of yeast transcriptional activator GAL4 DNA-binding domain with the herpes simplex virus VP16 activation domain, and the upstream activator sequence with six repeats of UAS (6 3 UAS). The following part of GAL4-UAS is GUS, a b-glucuronidase gene. The enhancer trap system has a higher probability of detecting the expression of gene whose promoter locates near the trap system, and hence is possibly more effective for identifying gene functions in reverse genetic studies (Greco et al., 2001; Wu et al., 2003; Zhang et al., 2006). The histochemical staining of GUS in different developmental stages of rice embryos showed that the activity of GUS enhanced by the promoter of OsMT2b was strong in the top of the scutellum at 7 DAP (Fig. 3C), but almost not detected in the embryos at 5 and 10 DAP (Fig. 3, B and D). In the germinating embryos, GUS staining signal was intense in the scutellum (arrow) and the cap of the radicle at 2 DAG (arrowhead; Fig. 3, E and F), which accorded with the result of MPSS that the OsMT2b transcript level was high in seeds at 3 DAG. In the embryos at 4 DAG, with shoots and roots emerging out, the GUS signal still concentrated in rice scutellum, and also appeared in the vascular bundle of shoot (arrowhead; Fig. 3G), which suggests that OsMT2b may play a role in the development of rice embryo scutellum. The GUS signal in immature panicles verified the second expression peak of OsMT2b in MPSS analysis.In florets of a 1-cm-long panicle, the GUS staining located in the pedicel and the basement of glumes (Fig.3H). In florets of an 8-cm-long panicle, besides the positions described above, GUS expressions were also detected in the anthers and pistils (Fig. 3I). The signal was intense in the basal parts of ovaries before pollination (Fig. 3J), and in the basal parts of stigmas and ovaries after pollination (Fig. 3K).To locate OsMT2b precisely at rice roots as described in MPSS results, GUS staining was performed. The strong signals were detected predominantly in the basal parts of lateral roots and nearby vascular cylinders of roots (Fig. 3, L and M). In the part of the root with lateral root primordium, the GUS staining mainly focused in root primordium, the passage cell of endodermis, and cells of pericycle and phloem, but not in the cells of epidermis, cortex, endodermis, and vessels
Cytokinin Regulates the Transcript Levels of OsMT2b
To investigate whether metal ions, hormones, and stress-related factors are involved in the regulation of OsMT2b, we harvested rice seedlings treated with various factors and detected the transcriptional level of OsMT2b by real-time quantitative reverse transcription (RT)-PCR. The expression levels of OsMT2b were markedly increased in the roots treated with iron (Fe), zinc (Zn), and indole-3-acetic acid (IAA), but observably decreased in the treatments of copper (Cu), 6-bezyladenine (6-BA), kinetin (KT), and NaCl (Fig. 4A). In shoots, its expression was elevated in the treatments of manganese (Mn), but remarkably decreased in the treatment of 6-BA and KT (Fig. 4B). Besides that, 4�C cold treatment also obviously reduced the transcript level of OsMT2b in roots and shoots, which is consistent with the results of MPSS. In view of the intensive response of OsMT2b transcripts to the treatment of cytokinin, we chose zeatin,a kind of monocotyledonous endogenous cytokinin, to treat rice seedlings and assayed its effect on rice roots by real-time quantitative RT-PCR. The result showed that the OsMT2b expression level started to distinctly decrease by the treatment of zeatin from 1 to 20 mM and reached the lowest value in 10 mM (Fig. 4C). To further confirm the down-regulation of OsMT2b transcripts by zeatin, we performed GUS activity assay in the roots of GAL4-UAS rice seedlings treated with zeatin. Quantitative analysis showed the alteration of GUS translation levels enhanced by the promoter of OsMT2b was dependent on the change of zeatin concentration. GUS activities decreased in the treatments of 1 to 20 mM zeatin, and yet reached the lowest value in 10 mM zeatin (Fig. 4D), which was accorded with the alteration of OsMT2b transcript levels. Based on the analysis of real-time quantitative RT-PCR in wild-type rice and the assay of GUS activity in GAL4-UAS rice, the results indicate that the expression of OsMT2b is down-regulated by cytokinin.
Evolution
Please input evolution information here.
You can also add sub-section(s) at will.
Labs working on this gene
College of Life Sciences, Wuhan University, Wuhan 430072, China
References
Jing Yuan, Dan Chen, Yujun Ren,et a. Characteristic and Expression Analysis of a Metallothionein Gene, OsMT2b, Down-Regulated by Cytokinin Suggests Functions in Root Development and Seed Embryo Germination of Rice.[J]Plant Physiology,2008.146,1637–1650.