Os05g0196500
This is a gene which encodes a histone H3K4 demethylase in rice.
Contents
Annotated Information
Function
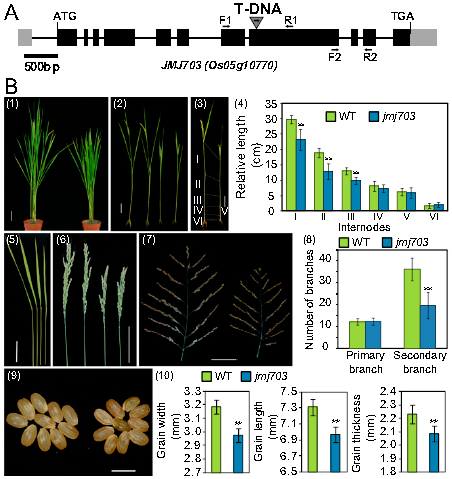
This gene encodes a histone H3K4 demethylase —JMJ703 which involves in both transposon(TE) silencing and stem elongation in rice. jmj703 mutants displayed pleiotropic phenotypes such as erect leaves, significantly decreased secondary panicle branches and abnormal phenotypes of grains, the most obvious of which was dwarfism.[1] JMJ703 is a JmjC domain-containing protein, which can specifically demethylate methylated histone H3 lysine 4 and reverse all three forms of H3K4me(mono-, di-, or trimethylation).[2] It is required for TEs silencing. Impaired JMJ703 activity leads to elevated levels of H3K4me3, misregulation of numerous endogenous genes, and the transpositional reactivation of two families of non-LTR retrotransposons(Karma and LINE1). These changes enhance the movement of transposons that are normally silenced and immobile. So we can conclude from these findings that plants use H3K4me3 demethylase to constitutively remove active chromatin marks and maintain the silent status of a subset of retrotransposons to preserve genome integrity. However, loss of JMJ703 does not affect TEs (such as Tos17) previously found to be silenced by other epigenetic pathways, indicating that different subsets of TEs may be regulated by distinct epigenetic pathways.[1] Besides, loss-of-function mutation of the gene reduces cell division rate of the stem and size of plant stature, showing the importance of this protein in plant growth. The mechanism may be that JMJ703 regulates H3K4me3 on CKX genes and that the mutant phenotype might be due to cytokinin deficiency caused by increased H3K4me3 and increased expression of CKX genes.[2]
Expression
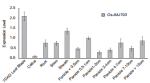
JMJ703 is expressed at relatively high levels in leaves of 7 d-after-germination (DAG) seedlings compared with all of the other tissues tested.[1]
Evolution
Plant JmjC proteins are conserved with yeast and animal homologues, but display several distinct features.[2] JMJ703 is the homolog of the Arabidopsis H3K4 demethylase JMJ14, which is involved in flowering time regulation and gene silencing.[1] The crystal structure of the catalytic domain of the protein shares a general similarity with that of mammalian and yeast proteins that demethylate methylated histone H3 lysine 9 and lysine 36, but displays several distinct structural features that are important for substrate and cofactor binding and enzymatic activity of the protein. It is found that key amino acids involved in the specific structures are conserved within known H3 lysine 4 demethylases, which may be involved in the specificity to histone H3 lysine 4.[2]
Crystal Structure
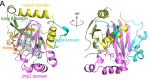
Gel filtration reveals that c-JMJ703 exists as a monomer in solution, suggesting that the monomer may be the biological unit. The c-JMJ703 molecule presents a canonical overall folding of JMJD2 proteins and contains four of the five domains defined in the structure of c-JMJD2A: the JmjN domain (A139-K199), the long b-hairpin (D200-T271), the mixed domain (L272-V377) and the JmjC domain (L378-A498). The JmjC domain, sandwiched by the JmjN and the long b-hairpin with the mixed domain, adopts a jellyroll-like structure with two fourstranded b-sheets as a cupin fold. Although c-JMJ703 shares low primary sequence similarity (less than 25% of the sequence identities) with the reported structural homologues, the core portion, especially the JmjC domain and the catalytic center of c-JMJ703, presents a topology similar to that of JMJD2 proteins with root mean square derivations of 1.75 A ° and 1.81 A ° relative to c-JMJD2A and c-Rph1, respectively. Meanwhile, c-JMJ703 also displays several significant structural differences.[2]
Knowledge Extension
Numerous epigenetic marks, including acetylation, methylation, phosphorylation, and ubiquitination, are introduced at particular sites on histone tails. Posttranslational modifications of histone tails are critical epigenetic marks that regulate diverse cellular processes. Histone lysine methylation activates or represses transcription, depending on the site and degree of these modifications. Two classes of histone lysine demethylases remove histone methylation. Lysine demethylase 1 (KDM1, also known as LSD1) is a flavin adenine dinucleotide (FAD)-containing enzyme that removes mono-/di-methylation. The Jumonji C-terminal domain (JmjC) family of histone demethylases uses Fe2+ and α-ketoglutarate as cofactors to remove all methylation states. Like all other histone modifications, histone lysine methylation is dynamic and reversible. Its addition and removal are catalyzed by histone methyltransferases and demethylases, respectively. Histone demethylases contribute to the regulation of the steady-state levels of histone methylation and are thus crucial for many chromatin-based cellular processes.[3]
Labs working on this gene
- National Key Laboratory of Crop Genetic Improvement, Huazhong Agricultural University, Wuhan, China.
- State Key Laboratory of Plant Genomics and National Center for Plant Gene Research, Institute of Genetics and Developmental Biology, Chinese Academy of Sciences, Beijing, China.
References
- ↑ 1.0 1.1 1.2 1.3 1.4 1.5 Cui X, Jin P, Cui X, et al. Control of transposon activity by a histone H3K4 demethylase in rice[J]. Proceedings of the National Academy of Sciences, 2013, 110(5): 1953-1958.
- ↑ 2.0 2.1 2.2 2.3 2.4 2.5 Chen Q, Chen X, Wang Q, et al. Structural basis of a histone H3 lysine 4 demethylase required for stem elongation in rice[J]. PLoS genetics, 2013, 9(1): e1003239.
- ↑ Hou H, Yu H. Structural insights into histone lysine demethylation[J]. Current opinion in structural biology, 2010, 20(6): 739-748.