Os08g0524100
This gene belongs to Histone Deacetylase Gene family, which symbol is OsSRT1.
Contents
Annotated Information
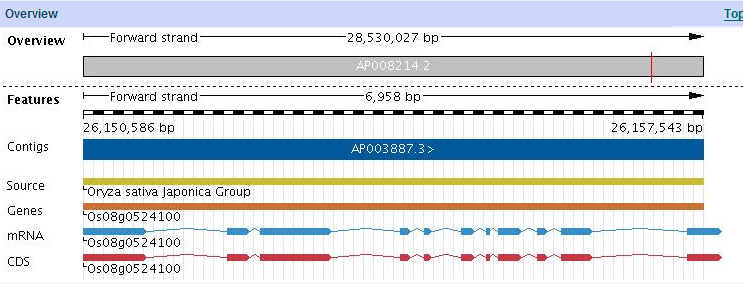
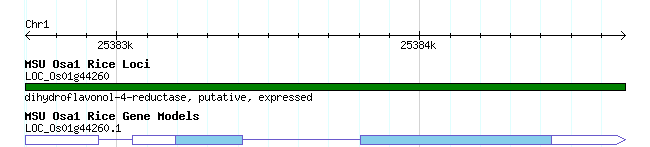
gene overview
Function
OsSRT1 is a widely expressed nuclear protein with higher levels in rapidly dividing tissues. OsSRT1 RNA interference induced an increase of histone H3K9 (lysine-9 of H3) acetylation and a decrease of H3K9 dimethylation, leading to H2O2 production, DNA fragmentation, cell death, and lesions mimicking plant hypersensitive responses during incompatible interactions with pathogens, whereas overexpression of OsSRT1 enhanced tolerance to oxidative stress. Transcript microarray analysis revealed that the transcription of many transposons and retrotransposons in addition to genes related to hypersensitive response and/or programmed cell death was activated. Chromatin immunoprecipitation assays showed that OsSRT1 down-regulation induced histone H3K9 acetylation on the transposable elements and some of the hypersensitive response-related genes, suggesting that these genes may be among the primary targets of deacetylation regulated by OsSRT1.The rice SIR2-like gene is required for safeguard against genome instability and cell damage to ensure plant cell growth, but likely implicates different molecular mechanisms than yeast and animal homologs. Overexpression of OsSRT1 Enhanced Tolerance to Oxidative Stress
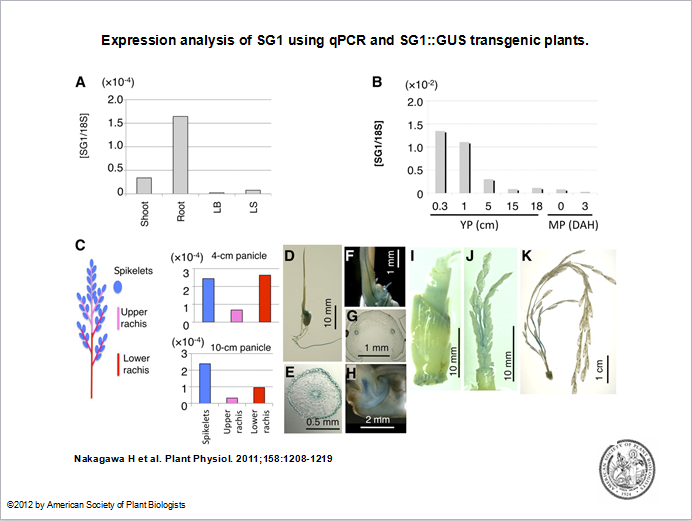
Expression
Please input expression information here. Baseline expression in tissue(s) was found for OS08G0524100.And no differential experiments were found for OS08G0524100.Related picture is shown as followed.
Evolution
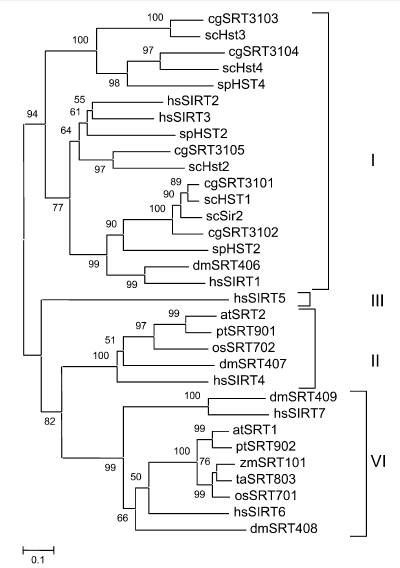
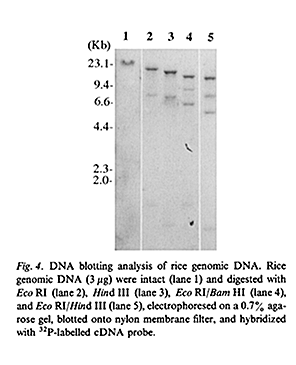
Rice Genome Contains Two SIR2-Related Genes(figure 1A,2A,2B)
Down-Regulation of OsSRT1 by RNAi Induced Programmed Cell Death in Rice.the OsSRT1 RNAi had little effect on overall histone H3 acetylation. However, the acetylation of H3K9 was induced, whereas the dimethylation of H3K9 was reduced, in agreement with the antagonistic relationship between H3K9 acetylation and dimethylation(3A,3B,3C)
Overexpression of OsSRT1 Enhanced Tolerance to Oxidative Stress. Transcriptomic Analysis Revealed Activation of ManyTransposon and PCD-Related Genes. OsSRT1 RNAi Induced Histone H3K9 Acetylation on Transposable Elements.
OsSRT1 RNAi Induced Histone H3K9 Acetylation on Transposable Elements
The latest report(in two years) Rice epigenomics and epigenetics: challenges and opportunities Rice is one of the most important food crops in the world and has been established as a model for plant genome study. Genomic sequences of two of the three subspecies of rice are available [1 and 2]. With its accurate genomic sequences, abundant genetic resources and availability of highly efficient reverse genetic tools, rice has become a model for cereal epigenomics and epigenetics as well. Epigenomics refers to genome-wide chromatin modification profiles mainly including DNA methylation and histone modifications, which control chromatin accessibility to DNA replication and repair and transcription machineries, thereby modulating the genome activities. Epigenetics studies mechanisms that are involved in variation and inheritance of epigenomic modifications. A wealth of trait variations among different rice species, subspecies and cultivars related to epigenomic variations, which may have been accumulated during the long history of rice evolution, domestication and selection, provides unique opportunities for crop epigenetic study. We summarize and discuss features of rice epigenomes and epigenetic variations and its possible implication in heterosis and genetic improvement.
Labs working on this gene
Benhamed M, Bertrand C, Servet C, Zhou DX Blander G, Guarente L Brodersen P, PetersenM, Pike HM, Olszak B, Skov S, Odum N Carrozza MJ, Utley RT, Workman JL, Coˆ te´ J Chu Z, Yuan M, Yao J, Ge X, Yuan B, Xu C, Li X, Fu B, Li Z, Bennetzen JL, Czernic P, Huang HC, Marco Y DaiM, Hu Y, Zhao Y, Liu H, Zhou D-X Donahue JL, Okpodu CM, Cramer CL, Grabau EA, Alscher RG
References
[1] Down-Regulation of a SILENT INFORMATION REGULATOR2-Related Histone Deacetylase Gene, OsSRT1,Induces DNA Fragmentation and Cell Death in RiceRice1.
[2] Rice histone deacetylase genes display specific expression patterns and developmental functions
[3] Involvement of Histone Modifications in Plant Abiotic Stress Responses
[4] Arabidopsis Putative Deacetylase AtSRT2 Regulates Basal Defense by Suppressing PAD4, EDS5 and SID2 Expression
[5] HISTONE DEACETYLASE6 Interacts with FLOWERING LOCUS D and Regulates Flowering in Arabidopsis
[6] Rice epigenomics and epigenetics: challenges and opportunities
[7] Regulatory Function of Histone Modifications in Controlling Rice Gene Expression and Plant Growth
[8] A comparative transcriptomic analysis of the extremely boron tolerant plant Puccinellia distans with the moderately boron tolerant Gypsophila arrostil
Structured Information
- ↑ Down-Regulation of a SILENT INFORMATION REGULATOR2-Related Histone Deacetylase Gene, OsSRT1,Induces DNA Fragmentation and Cell Death in RiceRice1.
- ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedref2 - ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedref3 - ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedref4 - ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedref5 - ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedref6 - ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedref7 - ↑ Cite error: Invalid
<ref>tag; no text was provided for refs namedref8