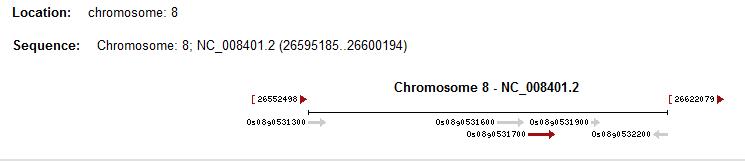
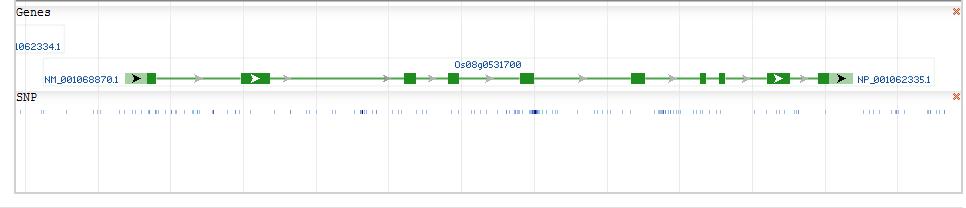
Os08g0531700
Please input one-sentence summary here.
Contents
Annotated Information
Function
The basic function of molecular behavior, such as catalytic or combination. It describes the activity of the gene product at the molecular level or function, a gene product usually has one or more molecular function. The function of binding to a specific DNA sequence in order to modulate transcription. The transcription factor may or may not also interact selectively with a protein or macromolecular complex.It has so many molecular functions,such as: antioxidant activity,enzyme activity,the ability of combination,nutrient storage activity,transcriptional regulation activity, translation regulation factor activity,molecular conduction activity and protein transport activity.
Expression
For expression analysis, the rice panicle and seed develop-ment stages were divided into 6 and 5 broad categories, respectively, based on landmark developmental events as described by Itoh and coworkers; information avail-able at oryzabase and our preliminary histochemical analysis.Seedlings subjected to three stress conditions, viz. desiccation, cold and salt stress, were also included in this analysis.
MADS-box transcription factors, besides being involved in floral organ specification, have also been implicated in several aspects of plant growth and development. In recent years, there have been reports on genomic localization, protein motif structure, phylogenetic relationships, gene structure and expression of the entire MADS-box family in the model plant system, Arabidopsis. Though there have been some studies in rice as well, an analysis of the complete MADS-box family along with a comprehensive expression profiling was still awaited after the completion of rice genome sequencing. Furthermore, owing to the role of MADS-box family in flower development, an analysis involving structure, expression and functional aspects of MADS-box genes in rice and Arabidopsis was required to understand the role of this gene family in reproductive development.

Evolution
In the process of evolution of monocotyledonous SEP - like genes,some new functionalization or subfunctionalization. To examine the evolutionary relationships of MADS-box genes in rice (including the 10 new genes identified in this study) and Arabidopsis, a tree was constructed using only the conserved MADS-box domain. Five groups, as described by Parenicova and coworkers were identified containing representative genes of both rice and Arabidop-sis. All the Arabidopsis proteins were found to lie in groups similar to those identified previously , except AGL47 and AGL82, which instead of forming a basal branch of the Mβ, grouped with Mγ proteins in our analysis as shown in supplementary figure S1. OsMADS64 grouped separately with AGL33 of Arabidop-sis, which does not cluster with any of the MADS groups described above.
Labs working on this gene
MADS-box transcription factors, besides being involved in floral organ specification, have also been implicated in several aspects of plant growth and development. In recent years, there have been reports on genomic localization, protein motif structure, phylogenetic relationships, gene structure and expression of the entire MADS-box family in the model plant system, Arabidopsis. Though there have been some studies in rice as well, an analysis of the complete MADS-box family along with a comprehensive expression profiling was still awaited after the completion of rice genome sequencing. Furthermore, owing to the role of MADS-box family in flower development, an analysis involving structure, expression and functional aspects of MADS-box genes in rice and Arabidopsis was required to understand the role of this gene family in reproductive development. Through the mutation research of flowering plants that SEPALLATA - like by MADS - box genes on the differentiation of the sepals, petals, stamens and carpels in floral decisions are required, as well as to the decision to determine E class flower organs.SEP - like genes encode transcription factor by MADS domain structure, and system analysis showed that such genes of angiosperms constitute a special branch of gene functions in dicotyledonous plants and show different degrees of redundancy and the functionalization. When OsMADS7 and OsMADS8 expression are cut at the same time, the transgenic plants show serious phenotypic defects,flowering delay, homologous into stamens and carpels lodicule, palea/lemma of organs, and lost to take decisions.In the analysis of different SEP - like rice gene expression and protein interaction characteristics of space,scientists found that these genes are very similar, but also show some specific differences.
References
1.The Rice Annotation Project Database (RAP-DB): 2008 update. Rice Annotation Project, et al. Nucleic Acids Res, 2008 Jan. 2.Curated genome annotation of Oryza sativa ssp. japonica and comparative genome analysis with Arabidopsis thaliana. Rice Annotation Project, et al. Genome Res, 2007 Feb. 3.The Rice Annotation Project Database (RAP-DB): hub for Oryza sativa ssp. japonica genome information. Ohyanagi H, et al. Nucleic Acids Res, 2006 Jan 1. 4.The map-based sequence of the rice genome. International Rice Genome Sequencing Project. Nature, 2005 Aug 11. 5.Rongfeng Cui;Jiakun Han;Suzhen Zhao;Kunmei Su;Feng Wu;Xiaoqiu Du;Qijiang Xu;Kang Chong;Günter Theißen;Zheng Meng Functional conservation and diversification of class E floral homeotic genes in rice (Oryza sativa) The Plant Journal, 2010, 61(5): 767-781 6.Rita Arora;Pinky Agarwal;Swatismita Ray;Ashok K Singh;Vijay P Singh;Akhilesh K Tyagi;Sanjay Kapoor MADS-box gene family in rice: genome-wide identification, organization and expression profiling during reproductive development and stress BMC Genomics, 2007, 8: 242