Os11g0703900
Gene Os11g0703900, namely Hsp70,meaning rice heat shock protein gene,is a multigenic family, andregulates the expression of stress response genes, such as the regulation of the heat shock protein transcription factor (heat-shock factor, HSF).
Contents
Annotated Information
Function
- Heat shock proteins (heat shock protein, Hsp) are a class of resistance protein molecules playing an important role in the process of environmental stress[1][2][3].
- Hsp70 genes encode for a group of highly conserved chaperone proteins across the living systems encompassing bacteria, plants, and animals[4].
- Hsp70 can not only prevent proteins aggregation and help inactivated proteins refolding, but also assist the functions of transport and participate in the intracellular signal transduction, such as protein kinase C (protein kinase C, PKC) and protein phosphorylation activity (protein phosphatase)[5].
- It is noted that Hsp70 may function as a negative feedback regulator of HSF activity[6][7].In this process, Hsp70 binds to denatured proteins and releases HSF leading to its activation during heat stress[6].
- Heat stress results in misfolding and aggregation of cellular proteins. Heat shock proteins (Hsp) enable the cells to maintain proper folding of proteins, both in unstressed as well as stressed conditions. Hsp70 genes encode for a group of highly conserved chaperone proteins across the living systems encompassing bacteria, plants, and animals. In the cellular chaperone network, Hsp70 family proteins interconnect other chaperones and play a dominant role in various cell processes.
- In animal cells, Hsp70s play an anti-apoptotic role via two pathways. In one, Hsp70 binds directly to Apaf-1 and inhibits the formation of apoptosomes. In the other, Hsp70 interacts with members of the Bcl-2 family, and inhibits the release of cytochrome c, endonucleases, and apoptosis-inducing factor from mitochondria to the cytosol[8].
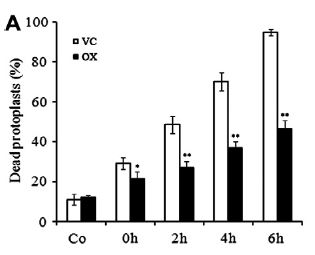
- In animal cells, the loss of mtHsp70 protein content in the marginal zone may lead to decreased functional mitochondria, whereas mtHsp70 over-expression preserves mitochondrial function, lessens oxidative stress and subsequent apoptosis[9].
- overexpression of mtHsp70 effectively reduced the percentage of dead cells (39.8% after 6 h recovery) compared with VC (63.2% after 6 h recovery).
Expression
- Rice Hsp70 superfamily genes are represented by 24 Hsp70 family and 8 Hsp110 family members[4].
- In rice, ER Hsp70 (BiP) is found to associate with nascent polypeptides that emerge into the ER lumen during prolamin translocation and facilitate peptide folding and assembly[10]. Over-expression of BiP relieves "ER stress"[8].
- Rice mitochondrial Hsp70 has been proposed as suppressor of heat and H2O2-induced programmed cell death[11].

- Hsp70 regulates the expression of stress response genes, such as the regulation of the heat shock protein transcription factor (heat-shock factor, HSF)[12][13].
- Overexpression of Hsp70 will decrease the concentration of Ca2+ intracellular[14], which usually works as a second messenger to stimulate the activity of protein kinases in downstream when the organism faces an environmental stress[15].
- Hsp70 are ATP-dependent chaperones having a conserved ~44-kD N-terminal ATPase domain (also called nucleotide binding domain; NBD), a ~18-kD substrate binding domain (SBD) and a ~10-kD variable C-terminal “lid.”.
- Hsp70 proteins require two cochaperones in the form of J-domain proteins and nucleotide exchange factors (NEFs) for their functions. Binding of J-domain proteins stimulates ATP hydrolysis that facilitates trapping of exposed hydrophobic region of substrates and closed conformation of SBD. NEFs participate in ATP–ADP exchange by catalyzing the release of ADP from Hsp70, leading to conversion of open conformation Hsp70 and substrate release[4].
- The stress induced expression profile of Hsp70 superfamily genes showed diverse response.
- During heat stress the expression of hsp70 correlates well with increases in internal ion concentrations, and can also be induced by excess salt or ethanol at normal growth temperatures.
Evolution
- Hsp70 is a multigenic family. Various members constituting this family are present in different cellular compartments. In Saccharomyces cerevisiae, 14 Hsp70 genes representing nine cytosolic, three mitochondrial (Ssc, Ecm, and Ssq), and two ER (Kar2 and Lhs1) isoforms have been identified[16].In E. coli, there are three Hsp70 genes. Arabidopsis has 18 Hsp70 superfamily members, of which 14 belong to Hsp70 family and 4 belong to Hsp110/SSE family[17].The C-terminus of organellar Hsp70 proteins is highly conserved and unique for each organelle. The motif for the cytosolic group is EEVD, for the mitochondrion is PEAEYEEAKK and for the plastid is PEGDVIDADFTDSK [18].The C terminus EEVD motif of cytosolic Hsp70 proteins interacts with co-chaperones that contain several degenerate 34 amino acid repeats, called tetratricopeptide repeats.
- While Hsp70 genes clustered on clades A, B, and C, clade D contained Hsp110 family genes. Hsp110 family is a subfamily of Hsp70 superfamily, structurally very similar to Hsp70 having N-terminal ATPase domain and C-terminal peptide binding domain and functions as NEFs for Hsp70 family proteins[5].
- During heat stress the expression of hsp70 correlates well with increases in internal ion concentrations, and can also be induced by excess salt or ethanol at normal growth temperatures. These results appear to indicate that whereas hsp70 is induced by all stresses that lead to protein denaturation — including heat stress[19].
Labs working on this gene
- 1.Sarkar N K, Kundnani P, Grover A.Department of Plant Molecular Biology, University of Delhi South Campus, N Delhi 110021, India
- 2.Wei ZhangCollege of Life Science, Nanjing Agricultural University, Nanjing, Jiangsu 210095, China
- 3.Laboratorium voor Genetica, Universiteit Gent, K.L. Ledeganckstraat 35, B-9000 Gent, Belgium
- 4.An Department of Plant Molecular Systems Biotechnology and Crop Biotech Institute, Kyung Hee University, Yongin 446-701, South Korea
- 5.Institute of Microbiology, Chinese Academy of Sciences, Beijing 100101, China
References
<references> [1]
<references>
Structured Information
- Hsp70 are ATP-dependent chaperones having a conserved ~44-kD N-terminal ATPase domain (also called nucleotide binding domain; NBD), a ~18-kD substrate binding
domain (SBD) and a ~10-kD variable C-terminal “lid.” The flexible C-terminal lid assists in holding the substrates at SBD. ATPase domain performs regulatory role and SBD associates to hydrophobic regions exposed in non-native substrates.
| Gene Name |
Os11g0703900 |
|---|---|
| Description |
Heat shock protein 70 |
| Version |
NM_001075072.1 GI:115486792 GeneID:4351208 |
| Length |
4293 bp |
| Definition |
Oryza sativa Japonica Group Os11g0703900, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 11:30612818..30617110 |
| Sequence Coding Region |
30612889..30613102,30615017..30616752 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008404:30612818..30617110 source=RiceChromosome11 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008404:30612818..30617110 source=RiceChromosome11 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>atggcgggcaagggcgagggtccggccatcggcatcgaccttggcacgacctactcgtgcgtgggcgtttggcagcacgaccgcgtggagatcatcgccaacgaccagggcaaccgcaccaccccctcctacgtcggcttcaccgactccgagaggctcatcggagatgctgccaagaaccaggtcgccatgaaccccatcaacaccgtctttgatgccaagcgtctcattggcaggaggtttagcgatgcttctgttcagagtgacattaagctctggcccttcaaggtgattgctggacctggtgacaagcctatgattgttgtccagtacaagggtgaggagaagcagtttgctgcagaagagatctcctccatggtcctcatcaagatgcgtgagattgctgaggcctaccttggcaccaccatcaagaatgccgttgtcactgttcctgcctacttcaatgactcccagaggcaggccaccaaggatgctggagtgattgctggtctcaatgtcatgcgtatcatcaacgagccgactgctgcagctattgcctatggtcttgataagaaggccaccagcgttggtgagaagaatgtcctcatctttgaccttggaggtggtacctttgatgtctccctccttaccattgaggagggtatctttgaggtcaaggccacagctggtgacacccatcttggtggtgaagattttgacaatcgtatggtcaaccactttgtgcaagaattcaagaggaagaacaagaaggatatcactggcaaccccagggctctcaggaggttgaggacagcttgtgagagggcaaagaggaccctgtcctccactgcccagaccaccattgagatcgattccctgtatgagggcatcgacttctactcaaccatcacccgtgccaggtttgaggagctcaacatggatctcttcaggaagtgtatggagcctgtggagaagtgcctcagggatgctaagatggacaagagctctgttcatgatgttgtccttgttggtggctccactaggatcccgagggtgcagcagctcctgcaggatttcttcaacggcaaggagctttgcaagaacatcaacccagatgaggctgttgcttatggtgctgctgtccaggctgccatcttgagtggtgagggcaacgagaaggtccaggacctcctcctgttggatgttacccctctctctctcggtttggagactgctggtggtgtcatgaccgttttgatcccaaggaacaccaccattcccaccaagaaggagcaggtcttctccacctactccgacaaccagcctggtgtgctcatccaggtttatgagggtgagaggaccaggacacgtgacaacaacctgctggggaagtttgagctctctggaatccctcctgctcccaggggtgttccacagatcactgtttgcttcgacattgatgccaatggtatcctgaacgtgtctgctgaggacaagaccaccgggcagaagaacaagatcaccattaccaacgacaagggcaggcttagcaaggaggagattgagaagatggtccaggaggccgagaagtacaagtcagaggatgaggagcacaagaagaaggtggagtccaagaacgcgctggagaactacgcctacaacatgcgcaacaccatcaaggatgagaagatcgcctcgaagctcccggcagcggacaagaagaagatcgaggatgccatcgaccaggccatccagtggctggacggcaaccagctcgctgaggctgatgagttcgatgacaagatgaaggagctggagggcatctgcaaccccatcatcgccaagatgtaccagggcgctggcgcggacatggccggcggcatggacgaggacgatgctcccccggctggcggcagcggtgctggccccaagatcgaggaggtcgactaa</cdnaseq> |
| Protein Sequence |
<aaseq>MAGKGEGPAIGIDLGTTYSCVGVWQHDRVEIIANDQGNRTTPSY VGFTDSERLIGDAAKNQVAMNPINTVFDAKRLIGRRFSDASVQSDIKLWPFKVIAGPG DKPMIVVQYKGEEKQFAAEEISSMVLIKMREIAEAYLGTTIKNAVVTVPAYFNDSQRQ ATKDAGVIAGLNVMRIINEPTAAAIAYGLDKKATSVGEKNVLIFDLGGGTFDVSLLTI EEGIFEVKATAGDTHLGGEDFDNRMVNHFVQEFKRKNKKDITGNPRALRRLRTACERA KRTLSSTAQTTIEIDSLYEGIDFYSTITRARFEELNMDLFRKCMEPVEKCLRDAKMDK SSVHDVVLVGGSTRIPRVQQLLQDFFNGKELCKNINPDEAVAYGAAVQAAILSGEGNE KVQDLLLLDVTPLSLGLETAGGVMTVLIPRNTTIPTKKEQVFSTYSDNQPGVLIQVYE GERTRTRDNNLLGKFELSGIPPAPRGVPQITVCFDIDANGILNVSAEDKTTGQKNKIT ITNDKGRLSKEEIEKMVQEAEKYKSEDEEHKKKVESKNALENYAYNMRNTIKDEKIAS KLPAADKKKIEDAIDQAIQWLDGNQLAEADEFDDKMKELEGICNPIIAKMYQGAGADM AGGMDEDDAPPAGGSGAGPKIEEVD</aaseq> |
| Gene Sequence |
<dnaseqindica>72..285#2200..3935#ctctcccactcttcctcctccccaaaccctagccgccgccgccggcgtactcgagagaaacatcagcatccatggcgggcaagggcgagggtccggccatcggcatcgaccttggcacgacctactcgtgcgtgggcgtttggcagcacgaccgcgtggagatcatcgccaacgaccagggcaaccgcaccaccccctcctacgtcggcttcaccgactccgagaggctcatcggagatgctgccaagaaccaggtcgccatgaaccccatcaacaccgtctttggtaactacttattccctgctcctcagtcctctctctttctctctctccccccttagatctcatctctaaggtcttcgtcgtcgtcgtatgctagatctactcttgcccagctcgttcagatctgttagatttgtgtgggatgctctaattaattactaccaacttagtagatctggaactcttttgacttgcagatcttgctccactcctgttgcattttgaatctgtgttgatgtgatttaagttgttggcagtgctttttgcatactatttgaatttgtggatggttagcagtcagtacactaggatttattactggatctggacagatatgctcgtttcaatcggctcgatttgctatgttgatagctaacatgttgtgcttgtcctactgccctggcgcaaatttgatccatgcattcttttcgttaaatgatttgtcaagtagtaatgcttgtttgaaaatgatattattattattattattattttaaataaatatagcctgaaacattcttagtttcgaggagaagtgatgatgtcttatccataccacatgtctgcatctcatatgatgattaatctacatcatgcactaccatctgatctcctctagctcatacagcttttaacaagttaggcgatagatgtttgctacaaacagtcaactggatttgctttcttgaaagaaaaaaaatgaaccgtgtagtttgcaacctgtgatgtaaattaacccatttactgaactacattgattatttcagtaaaatcaccatctttcggctgaggttgcatgcttagttattatatttttaccaatgatgattttgttatgtagccagatactgtactttcttatgccatttggtatctgactcattgtctacttttgtgactatgtgtctcgaaatatggcatataataagggggaacggatcttgcgacattgatggctcctgtatgagcctccaattactatatagccgtttacatgttcttatcatgttacttttccaaattagtgctgcacatcttgtctgtttgcaagtttgggttcattcatattgctgtgggcattttaatacatcttaactataatgtaaatttcttgtacatcgctaaaattttctgccttgattgagtagactcttagatgacatctatcttaccttatgcacttgattcgtatttctgttacaacattttgattacttgatgtagttcacacaagcatgacacatctctttgaaaaaaagatttcattttttaaagcaagcctaatgctttagggtgggagaagtgatgatacttaatccatacaacatttctgcatctcagatgatgatcaatctacatcatgcactaccatctgatctccccatcatacaattttttattttgttttgttggattatataactttaacatttgataaagtgccaacagatttttatacacaatttcttatagatattaccatgacagcttgcagcctgtgatgcaaaagtacccattaaactgaattacactgattattccaaacaaatcaccatctttcggctgaggttgcttgcttgtttaaaattatgaatgcttcttgcacaaccatttaggactttcggtagtttgtaatgccatttggtatctgacacatggtctgcgcttgtgactatttgtcttgaaatatggcataaaacaagggggaacggatcttgcgacattggtggttcctctatgatcctccaattactatatagctgtttacaatttgttaagttaggagcataccttttgctttgtcatcaaagtcttggttcctttgtatttctagttcagattcatttttcttcagcttttgaaacataaactcagtatatttttactgtaattgctgacattgagtcttctctgcagatgccaagcgtctcattggcaggaggtttagcgatgcttctgttcagagtgacattaagctctggcccttcaaggtgattgctggacctggtgacaagcctatgattgttgtccagtacaagggtgaggagaagcagtttgctgcagaagagatctcctccatggtcctcatcaagatgcgtgagattgctgaggcctaccttggcaccaccatcaagaatgccgttgtcactgttcctgcctacttcaatgactcccagaggcaggccaccaaggatgctggagtgattgctggtctcaatgtcatgcgtatcatcaacgagccgactgctgcagctattgcctatggtcttgataagaaggccaccagcgttggtgagaagaatgtcctcatctttgaccttggaggtggtacctttgatgtctccctccttaccattgaggagggtatctttgaggtcaaggccacagctggtgacacccatcttggtggtgaagattttgacaatcgtatggtcaaccactttgtgcaagaattcaagaggaagaacaagaaggatatcactggcaaccccagggctctcaggaggttgaggacagcttgtgagagggcaaagaggaccctgtcctccactgcccagaccaccattgagatcgattccctgtatgagggcatcgacttctactcaaccatcacccgtgccaggtttgaggagctcaacatggatctcttcaggaagtgtatggagcctgtggagaagtgcctcagggatgctaagatggacaagagctctgttcatgatgttgtccttgttggtggctccactaggatcccgagggtgcagcagctcctgcaggatttcttcaacggcaaggagctttgcaagaacatcaacccagatgaggctgttgcttatggtgctgctgtccaggctgccatcttgagtggtgagggcaacgagaaggtccaggacctcctcctgttggatgttacccctctctctctcggtttggagactgctggtggtgtcatgaccgttttgatcccaaggaacaccaccattcccaccaagaaggagcaggtcttctccacctactccgacaaccagcctggtgtgctcatccaggtttatgagggtgagaggaccaggacacgtgacaacaacctgctggggaagtttgagctctctggaatccctcctgctcccaggggtgttccacagatcactgtttgcttcgacattgatgccaatggtatcctgaacgtgtctgctgaggacaagaccaccgggcagaagaacaagatcaccattaccaacgacaagggcaggcttagcaaggaggagattgagaagatggtccaggaggccgagaagtacaagtcagaggatgaggagcacaagaagaaggtggagtccaagaacgcgctggagaactacgcctacaacatgcgcaacaccatcaaggatgagaagatcgcctcgaagctcccggcagcggacaagaagaagatcgaggatgccatcgaccaggccatccagtggctggacggcaaccagctcgctgaggctgatgagttcgatgacaagatgaaggagctggagggcatctgcaaccccatcatcgccaagatgtaccagggcgctggcgcggacatggccggcggcatggacgaggacgatgctcccccggctggcggcagcggtgctggccccaagatcgaggaggtcgactaagcgcgaaatttgctagtgatccgcgcttcaagttatcttattgcattattagtgtctcttttatgttgtgttaaaaaactttgctgctatgacttatgatgatacccttggtctgtttgggcagtccttttgaattgaagaactgcgttgggagctctatgagaactatgaattgcgtgagaaatttaattattgtggtgcttgcagcgcctattatttggtctctgtcccacgtttattatttgcatgtgcgcattgtttgctagcaattgctgttgtgcttctcagccaacatgtaaactgtttggattgtttctgatttttggatgcattttttaaagcggatagtgaaggacgg</dnaseqindica> |
| External Link(s) |
- ↑ 1.0 1.1 Ahuja I, RCH. De Vos, Bones AM, Hall RD: Plant molecular stress responses face climate change. Trends in Plant Science 2010, Vol. 15, No. 12.
- ↑ 2.0 2.1 Wang W, Vinocur B, Shoseyov O, Altman A: Role of plant heat-shock proteins and molecular chaperones in the abiotic stress response. Trends in Plant Science 2004, Vol. 9, No. 5.
- ↑ 3.0 3.1 Timperio AM, Egidi MG, Zolla L: Proteomics applied on plant abiotic stresses: Role of heat shock proteins (HSP). Journal of Proteomics 2008, 71: 391-411.
- ↑ 4.0 4.1 4.2 4.3 Sarkar N K, Kundnani P, Grover A. Functional analysis of Hsp70 superfamily proteins of rice (Oryza sativa)[J]. Cell Stress and Chaperones, 2013, 18(4): 427-437.
- ↑ 5.0 5.1 5.2 Ding XZ, Tsokos GC, Kiang JG: Overexpression of HSP-70 inhibits the phosphorylation of HSF1 by activating protein phosphatase and inhibiting protein kinase C activity. The FASEB Journal 1998, 12:451–459.
- ↑ 6.0 6.1 6.2 Shi Y, Mosser D D, Morimoto R I. Molecularchaperones as HSF1-specific transcriptional repressors[J]. Genes & development, 1998, 12(5): 654-666.
- ↑ 7.0 7.1 Kim B H, Schöffl F. Interaction between Arabidopsis heat shock transcription factor 1 and 70 kDa heat shock proteins[J]. Journal of experimental botany, 2002, 53(367): 371-375.
- ↑ 8.0 8.1 8.2 Snowden C J, Leborgne‐Castel N, Wootton L J, et al. In vivo analysis of the lumenal binding protein (BiP) reveals multiple functions of its ATPase domain[J]. The Plant Journal, 2007, 52(6): 987-1000.
- ↑ 9.0 9.1 Kuzmin, E.V., Karpova, O.V., Elthon, T.E. and Newton, K.J. Mitochondrial respiratory deficiencies signal up-regulation of genes for heat shock proteins. J. Biol. Chem. 2004. 279, 20672–20677.
- ↑ 10.0 10.1 Muench D G, Wu Y, Zhang Y, et al. Molecular cloning, expression and subcellular localization of a BiP homolog from rice endosperm tissue[J]. Plant and cell physiology, 1997, 38(4): 404-412.
- ↑ 11.0 11.1 11.2 11.3 Qi Y, Wang H, Zou Y, et al. Over-expression of mitochondrial heat shock protein 70 suppresses programmed cell death in rice[J]. FEBS letters, 2011, 585(1): 231-239.
- ↑ 12.0 12.1 Sarkar N K, Kundnani P, Grover A. Functional analysis of Hsp70 superfamily proteins of rice (Oryza sativa)[J]. Cell Stress and Chaperones, 2013, 18(4): 427-437.
- ↑ 13.0 13.1 Kim BH, Schöffl F. Interaction between Arabidopsis heat shock transcription factor 1 and 70 kDa heat shock proteins. Journal of Experimental Botany 2002, 53(367):371-375.
- ↑ 14.0 14.1 Kiang JG, Ding XZ, McClain DE. Overexpression of HSP-70 attenuates increases in [Ca2+]I and protects human epidermoid A-431 cells after chemical hypoxia. Toxicology and Applied Pharmacology 1998, 149(2):185-194.
- ↑ 15.0 15.1 Knight H: Calcium signaling during abiotic stress in plants. International Review of Cytology 2000, 195:269-324.
- ↑ 16.0 16.1 Walsh P, Bursać D, Law Y C, et al. The J‐protein family: modulating protein assembly, disassembly and translocation[J]. EMBO reports, 2004, 5(6): 567-571.
- ↑ 17.0 17.1 Lin B L, Wang J S, Liu H C, et al. Genomic analysis of the Hsp70 superfamily in Arabidopsis thaliana[J]. Cell stress & chaperones, 2001, 6(3): 201.
- ↑ 18.0 18.1 Guy C L, Li Q B. The organization and evolution of the spinach stress 70 molecular chaperone gene family[J]. The Plant Cell Online, 1998, 10(4): 539-556.
- ↑ 19.0 19.1 Frank Van Breusegem, Rudy Dekeyser, Ana Beatriz Garcia,et al.Heat-inducible rice hsp82 and hsp70 are not always co-regulated[J].Planta, 1994, 193(1): 57-66.