Difference between revisions of "Os04g0653000"
(→Annotated Information) |
(→EG2 is a repressor of JA response) |
||
| Line 35: | Line 35: | ||
Please input information here. | Please input information here. | ||
| − | + | yeast two-hybrid (Y2H) assays show OsJAZ1 can interact with OsCOI1b, and the interaction was dependent on JA-Ile’s mimic, coronatine(COR), but not JA or MeJA. No interaction was observed between OsJAZ1 and OsCOI1a or OsCOI1c. | |
| + | |||
| + | [[File:eg-6.jpg]] | ||
| + | |||
| + | Furthermore, the Ala-to-Gly mutation in the Jas domain of OsJAZ1 in eg2-1D blocked the interact between OsJAZ1 and JOsCOI1b, suggesting that the Jas domain is required for the interaction between OsJAZ1 and OsCOI1b. because osjaz1-1D can't interact with OsCOI1b, so it can't be JA-dependent proteasome-mediated degradation. | ||
| + | |||
| + | |||
| + | [[File:eg-7.jpg]] | ||
| + | |||
| + | JA regulates early spikelet development possibly by activating the expression of OsMADS1 through OsMYC2, which is directly regulates OsMADS1 expression.And, OsJAZ1 interacts with OsMYC2 and suppresses the activity of OsMYC2 in triggering the transcription of OsMADS1.Both OsJAZ1 and osjaz1-1D interacted with OsMYC2, and the interaction required the Jas domain. | ||
| + | |||
| + | [[File:eg-8.jpg]] | ||
| + | |||
| + | Inconclude,eg2-1D can't interact with OsCOI1b, so it not degradation, but it also can interact with OsMYC2 to block OsMYC2 regulates the OsMADS1 expression. | ||
| + | |||
| + | [[File:eg-9.jpg]] | ||
==Labs working on this gene== | ==Labs working on this gene== | ||
Revision as of 08:23, 4 June 2014
Please input one-sentence summary here.
Contents
Annotated Information
Function
Please input function information here.
- EG2(extra glume 2)is a JA signalling repressor that interacts with a putative JA receptor, OsCOI1b, to trigger OsJAZ1’s degradation during spikelet development[1].
- Its high sequence similarity with the Arabidopsis JAZ proteins so also called OsJAZ1[1].
- It belong to TIFY family so also called TIFY family[2].
Expression
Please input expression information here.
EG2 expressed ubiquitously in the plant, from roots, stems and leaves, to spikelets and seeds.
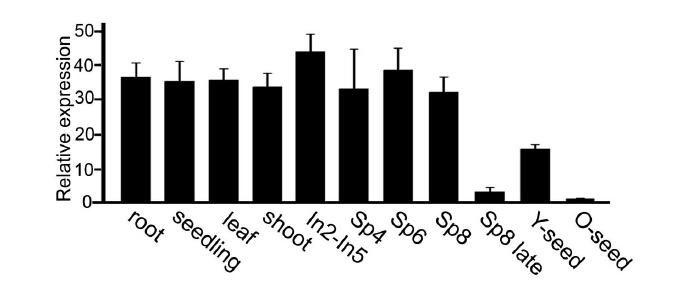
Mutation and Phenotype
Please input information here.
The EG2 gene was mapped to chromosome 4 between CQQ2 and CQQ3, two In-Del markers that define a 2.2 cM.
The mutation of EG2 is eg2-1D, which caused by C-to-G transition, resulting in a substitution of Ala by Gly in the fifth amino acid of the Jas domain. This motif was known to be critical for JAZ’s interaction with the JA receptor COI1.
mutants displayed defects in spikelet development,including extra glume-like structures between the sterile lemma and lemma and altered floral organ numbers and identities.
EG2 is a repressor of JA response
Please input information here.
yeast two-hybrid (Y2H) assays show OsJAZ1 can interact with OsCOI1b, and the interaction was dependent on JA-Ile’s mimic, coronatine(COR), but not JA or MeJA. No interaction was observed between OsJAZ1 and OsCOI1a or OsCOI1c.
Furthermore, the Ala-to-Gly mutation in the Jas domain of OsJAZ1 in eg2-1D blocked the interact between OsJAZ1 and JOsCOI1b, suggesting that the Jas domain is required for the interaction between OsJAZ1 and OsCOI1b. because osjaz1-1D can't interact with OsCOI1b, so it can't be JA-dependent proteasome-mediated degradation.
JA regulates early spikelet development possibly by activating the expression of OsMADS1 through OsMYC2, which is directly regulates OsMADS1 expression.And, OsJAZ1 interacts with OsMYC2 and suppresses the activity of OsMYC2 in triggering the transcription of OsMADS1.Both OsJAZ1 and osjaz1-1D interacted with OsMYC2, and the interaction required the Jas domain.
Inconclude,eg2-1D can't interact with OsCOI1b, so it not degradation, but it also can interact with OsMYC2 to block OsMYC2 regulates the OsMADS1 expression.
Labs working on this gene
Please input related labs here.
References
Please input cited references here.
Structured Information
| Gene Name |
Os04g0653000 |
|---|---|
| Description |
The start codon is not identified. |
| Version |
NM_001060638.1 GI:115461005 GeneID:4337240 |
| Length |
3445 bp |
| Definition |
Oryza sativa Japonica Group Os04g0653000, complete gene. |
| Source |
Oryza sativa Japonica Group ORGANISM Oryza sativa Japonica Group
Eukaryota; Viridiplantae; Streptophyta; Embryophyta; Tracheophyta;
Spermatophyta; Magnoliophyta; Liliopsida; Poales; Poaceae; BEP
clade; Ehrhartoideae; Oryzeae; Oryza.
|
| Chromosome | |
| Location |
Chromosome 4:33705210..33708654 |
| Sequence Coding Region |
33705211..33705317,33707373..33707475,33707567..33707792,33707906..33707963,33708275..33708365 |
| Expression | |
| Genome Context |
<gbrowseImage1> name=NC_008397:33705210..33708654 source=RiceChromosome04 preset=GeneLocation </gbrowseImage1> |
| Gene Structure |
<gbrowseImage2> name=NC_008397:33705210..33708654 source=RiceChromosome04 preset=GeneLocation </gbrowseImage2> |
| Coding Sequence |
<cdnaseq>ggagaggagaggaaggaggaagaggtggtggtggaggagaagagccatcagcagcagcagcagcaaggggaggaggagctcgtcggcctctcgctcgccggcggcaggcccaaagtatttccaatgtcgagccctccacctaatccttcgcaacttacaattttctatggtggatcagtatgtgtttatgactcagtgccgccagagaaggctcaagcaatcatgcttattgccgcagcagctgctgcagcggcgtctgccaccaaaagcaatgctgccattgctgttaagccccctgtgatgcctgcagccaatgctacccaagcagcagtctctcctgtgctcacaagatcgctctcattgcagagcacttctgtagcaactggacaacctcaagttgccgctgaccctagctcaatttgcaagctccaggctgatctccccattgccaggaggcactcccttcagcgcttcctagagaaacgccgtgacaggttggtgagcaaagctccatatcccacaaaatcctccgagggcatggaagcatcagggatggaagtaactgctgagggcaaggcccagtaa</cdnaseq> |
| Protein Sequence |
<aaseq>GEERKEEEVVVEEKSHQQQQQQGEEELVGLSLAGGRPKVFPMSS PPPNPSQLTIFYGGSVCVYDSVPPEKAQAIMLIAAAAAAAASATKSNAAIAVKPPVMP AANATQAAVSPVLTRSLSLQSTSVATGQPQVAADPSSICKLQADLPIARRHSLQRFLE KRRDRLVSKAPYPTKSSEGMEASGMEVTAEGKAQ</aaseq> |
| Gene Sequence |
<dnaseqindica>2..108#2164..2266#2358..2583#2697..2754#3066..3156#gggagaggagaggaaggaggaagaggtggtggtggaggagaagagccatcagcagcagcagcagcaaggggaggaggagctcgtcggcctctcgctcgccggcggcaggtagttttccggctccgccttctcctctcgcaagtcgagtcctattcagctttgggttgatcgattgggtcgcgggagagagagtatttggttggttgttcgggaaatgcatctcgcgcgggattctagccgcttttcagttcatagtatcgccccttctccgagaactcgccgccggggagccgcttcctgctgggagccgagcggcgccgcagccgcctccgaagtggtttaccggaggggatggattttctccggtactggactcctggtgtttgaatttttgggtggaaccggagactctttttttggtcgggtgaaattccttggaccgaccaaaaaaatctcgaatccactcggagtttcctccgttccagatggagctcttgcgttgaaaattttcctaaaaatagagtcaaaatgtttcgacttcgagttaagagtccaccctacaccgagttgcaactttcccttttttacgagttggtgtaatcatggaagagcccttagctttgctgcagtggatcaaattggtgggggttcttgcctaggtcatgtgaagattcttctgattagtggtagtctcatgcttctcctcggcctgcaatccaaatcccctttattcctttcccgctctcttgcgcttggcgtcagctgttcttaaccttgaacaaattagggtgtcattctagttgcacgtgttcttatccagatgttgggagtgcggtgaactgctgatattctactaaaccgatatcaattatatggatgttcaatttggttcacgtcactaggtaaatgatttcccaccttagaccaaacttaacggataggaaagaacttttggcaagtggcagatccaggtagtttttttccttccctttttctgtaattggcagttacttgttctccaaatacttggaactgctgtgaaaagtttccaattgagccttggaatcaggagcaatggggaactcttctaagttacttgcatttcttctttaaagactcagaaataataagcaagaaacacacatagatctggtgaaacttgagttagcgtgcagtttttaacacctccatgtaccaaggggattcaatggtaaaggcctcatccccttgtcttatctttacttcaagactaaatctgtagctggggaccaagtaccatagtttcgcaactcttatataatccactaggctgctgtgaaaacaatagatgcagtgtttgttttgaagtaagtaaggtgaaagtaaaaatacctggtaatggctaatatgaatcgtggctattattaaatgtttcagcattttctgaataggtaaaccacatatatgtaactatgtaagatgtttttaaatctagatgttctattgagcctgtaacctggtcttttagactagattatgctgagactttgaagaaaaaaatttcagtacattccgtccatcttggacaacctttggacagattattaactctgaatcagtcccatcatgctgcacatgttggatatttcagagtggaccataaagttgcaggcctagtttgctatttttagattttagatcttgtgagttttaatcttgatcatgctattagactcagaatgtgtgtaccaaaaacattgataatggtatctatcctgacttgggtataacttgtgttactggcagctgtgggagggctatgttggcgtagcgtactgaccaagagataatcctctcatcaagtagaatatagagagtagaataattatttatgaaggagatgggcctttatctggctgcagacaaggtgcatagtaaagctcattaaacctctaaaaaaggctggaaatacggcctacggtgtaattgcacattatacatcagttagctaggttggcctccaattttatatggcactttacccttggatgcccatcttatgtattgccatgtactcatgtatgcaattagtcaagaacggcccacttatgccaatatctgtgtaactatgcaatctgttattaccttcttgcacatttactaatttcgctttgtcataatcaggcccaaagtatttccaatgtcgagccctccacctaatccttcgcaacttacaattttctatggtggatcagtatgtgtttatgactcagtgccgccagagaaggtcatattcaccccacttatcttcgttatatcctccgactaactgttgaaagtttatgctaattcgttatctgcaacattgtcatcaacaggctcaagcaatcatgcttattgccgcagcagctgctgcagcggcgtctgccaccaaaagcaatgctgccattgctgttaagccccctgtgatgcctgcagccaatgctacccaagcagcagtctctcctgtgctcacaagatcgctctcattgcagagcacttctgtagcaactggacaacctcaagttgccgctgaccctagctcaatttgcaagctccaggctggtaagactacaagatctttttgtgaatctaaaatggggctgttttaatccctcgtcatatgatgtactaagtgattgaagatgctacaaatttattctttatttgctgtgcagatctccccattgccaggaggcactcccttcagcgcttcctagagaaacgccgtgacaggttggcatgaatgaatgtagaaatagcattctatatttgtctgatgcatttccttaggtttagtggtccattagaacatgctaaaagaatgtggatagctggaaactgacttgtaggggataaaacatttgctttagcttgacttggatacgaaccaagtatatgtgcaaggactgttttcagtttcattaccatagcccatgttttcatgatacataacaaagtagccagtagttatagaaacacagctttctgcaaatttgagaaatccttagcttcaactgattaagccaaatgaccatgttaagcaggttggtgagcaaagctccatatcccacaaaatcctccgagggcatggaagcatcagggatggaagtaactgctgagggcaaggcccagtaactgtcaacttcgaatggtggaggctgacgtttggagggggaaggaaggaaaggcgtcgcgttcattgtttagatggtgtcgcaccctagaaccttactaataatcgaacctatgcccttattatcgttgtactgttccttatgtgctagtctattctgtcatgaatttgtttactcgaacagcaaactgctatgatagtagtatatcattgtacatcatgcatatcattgcctgacaatgttatcgtgttcgtctagtaatgatatcatccaggttaattgttactagt</dnaseqindica> |
| External Link(s) |






