2016 Big Data Forum for Life and Health Sciences
2016 Big Data Forum for Life and Health Sciences (December 5-8, 2016)

Invited Speakers

Prof. Amir Ali Abbasi
National Center for Bioinformatics
Quaid-i-Azam University
Pakistan

Prof. Erik Bongcam-Rudloff
Head, SLU-Global Bioinformatics Centre
Swedish University of Agricultural Sciences
Sweden

Dr. Domenica D'Elia
Chair of EMBnet - The Global Bioinformatics Network
National Research Council
Italy

Prof. Yixue Li
Key Laboratory of Systems Biology
Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences
China

Dr. Eugene Yaschenko
Chief of Molecular Software Section
Information Engineering Branch
National Center for Biotechnology Information, USA

Prof. Kai Ye
Xi'an Jiaotong University
China

Prof. Jun Yu
CAS Key Laboratory of Genome Sciences & Information
Beijing Institute of Genomics, Chinese Academy of Sciences
China

Prof. Changqing Zeng
CAS Key Laboratory of Genomic and Precision Medicine
Beijing Institute of Genomics, Chinese Academy of Sciences
China
Agenda
| December 5: Pick-up & Registration | ||
|---|---|---|
| December 6: Invited Talks, Conference Hall at the 1st floor, BIG, CAS | ||
| 09:00 - 09:10 | Welcome and introduction (Zhang Zhang) | |
| 09:10 - 09:50 |
Life Cycle of Big Data - The Sequence Read Archive at NCBI Eugene Yaschenko, Chief of Molecular Software Section, NCBI, USA [Personal Profile] |
|
| Eugene Yaschenko serves as Chief, Molecular Software Section, Information Engineering Branch (IEB) at National Center for Biotechnology Information (NCBI). His experience at NCBI include coordinating public access to sequence, genetics, structural, and bibliographic information, establishing collaborative research projects with NIH institutes and laboratories, and consulting/advising governmental agencies on methods of software and database design. Mr. Yaschenko has worked on several large-scale bioinformatics resources, including GenBank, ClinVar, NIH Manuscript Submission (NIHMS), Database of Genotypes and Phenotypes (dbGaP), and Sequence Read Archive. These efforts have played an important role in the scientific research community. Mr. Yaschenko received his MS degrees in Physics from Lomonosov Moscow State University in 1992 and from The Catholic University of America in 1994. | ||
| 09:50 - 10:30 |
Precision Medicine Research in China: Challenges and Opportunities Changqing Zeng, BIG, CAS, China [Personal Profile] |
|
| Dr. ZENG, director of CAS Key Lab of Genomic & Precision Medicine, obtained her Bachelor and Master degree at Beijing Normal University, and Ph.D. degree at the University of Alabama at Birmingham. Dr. Zeng had her postdoc training at Baylor College of Medicine and worked as a research assistant professor at University of Texas Houston Medical School before 2002. She was the Steering Committee member of the International HapMap Project, the coordinator of the Chinese HapMap Consortium and responsible for the HapMap China Chapter which covers 10% of phase 1 data production. Dr. Zeng was the awardee of National Talents of New Century Project, National Science Fund for Distinguished Young Scholars from the National Natural Science Foundation of China, and the CAS Hundred-Talent Program etc. Currently, Dr. Zeng serves as the CAS (Chinese Academy of Sciences) representative of the Global Alliance for Genomics and Health, a Panel Member of Human Genetic Resources Administration of China, and a Panel Member of the MOST (the Ministry of Science and Technology) for the National Major Research Plan in Precision Medicine. | ||
| 10:30 - 10:50 | Tea & Coffee Break | |
| 10:50 - 11:30 |
Bioinformatics meets Big Data, The Dawn of a New Era
Erik Bongcam-Rudloff, Head of SLU-Global Bioinformatics Centre, Swedish University of Agricultural Sciences, Sweden |
|
| 11:30 - 12:10 |
Systematic discovery of complex insertions and deletions in human cancers [Abstract] Kai Ye, Xi'an Jiaotong University, China |
|
| Complex insertions and deletions (indels) are formed by simultaneously deleting and inserting DNA fragments of different sizes at a common genomic location. Here we present a systematic analysis of somatic complex indels in the coding sequences of samples from over 8,000 cancer cases using Pindel-C. We discovered 285 complex indels in cancer-associated genes (such as PIK3R1, TP53, ARID1A, GATA3 and KMT2D) in approximately 3.5% of cases analyzed; nearly all instances of complex indels were overlooked (81.1%) or misannotated (17.6%) in previous reports of 2,199 samples. In-frame complex indels are enriched in PIK3R1 and EGFR, whereas frameshifts are prevalent in VHL, GATA3, TP53, ARID1A, PTEN and ATRX. Furthermore, complex indels display strong tissue specificity (such as VHL in kidney cancer samples and GATA3 in breast cancer samples). Finally, structural analyses support findings of previously missed, but potentially druggable, mutations in the EGFR, MET and KIT oncogenes. This study indicates the critical importance of improving complex indel discovery and interpretation in medical research. | ||
| 12:10 - 14:00 | Lunch and Group Photo | |
| 14:00 - 14:40 |
The Strategic Planning for National Biomedical Big Data Infrastructure Yixue Li, Shanghai Institutes for Biological Sciences, CAS, China [Personal Profile] |
|
| Yi-Xue Li is the director in Shanghai Center for Bioinformation Technology, vice director and a full research professor of Key Laboratory of Systems Biology at Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences. Dr. Li received his BSc. and Msc. degrees in theoretical physics from Xinjiang University, China, in 1982 and 1987, respectively, and his Ph.D. degree in theoretical physics from the University of Heidelberg, Germany, in 1996. After Dr. Li got his Ph.D. degree, he worked as a bioinformatics research staff in European Molecular Biology Laboratory (EMBL) from 1997-2000, and came back to Shanghai, China in the middle of 2000. He has served as a reviewer/panelist for many national research foundations/agencies such as the Chinese National Science Foundation, the National High-Tech Program(863) and National Key Basic Research Program(973). He has organized several international conferences and workshops and has also served as a program committee member for several major national and international conferences like GIW, HUPO and National Bioinformatics Conference etc. | ||
| 14:40 - 15:20 |
Non-coding RNAs: from “Junk” matter to “Key” regulators [Abstract] Domenica D'Elia, Chair of EMBnet - The Global Bioinformatics Network, National Research Council, Italy [Personal Profile] |
|
| Domenica D’Elia works at the Institute for Biomedical Technologies of the Italian National Council (CNR) in Bari, Italy. She is Chair of EMBnet - The Global Bioinformatics Network, and member of the Bioinformatics Italian Society (BITS). She is also Deputy of the Training Coordinator of ELIXIR-ITA, member of GOBLET, the “Global Organisation for Bioinformatics Learning, Education & Training”, and scientific peer advisory and project reviewer for the Knowledge Transfer Programme for Bioinformatics, an initiative of the Centre for Proteomic & Genomic Research in South Africa. She holds a PhD in Biochemistry and Molecular Biology from the University of Bari; after few years working at the University on the biogenesis of mitochondria she moved to the CNR, where she has been working as a Project and Team Coordinator for the development of specialised biological databases. In this role, she has leaded and contributed to the development of diverse databases, bioinformatics integrated platforms and to the development of data mining tools. Presently her main research interest is the study of mechanisms of action and functions of miRNAs that may open new perspectives in the use of miRNAs as therapeutic targets and non-invasive biomarkers of diseases and of therapy response. Another research area she is investigating is the protective role of plant miRNAs in human cancer and age-related diseases. She also collaborates in a project, based at the Institute for Biomedical Technologies in Bari, for the development of a bioinformatics suite for the classification and analysis of non-coding RNAs. This bioinformatics platform has been designed to include also tools for gene expression profiling in total RNA-seq and small RNA-seq experiments, and for the discovery of ncRNAs functional interactions. Her activities are in support of data curation and functional analysis. She focuses on the discovery of ncRNAs functional role in the regulation of gene expression in the biomedical domains, collaborating with the Computer Science Departments of the University of Bari and Verona (IT) and with data producers at the Italian “Research Council for the agricultural research and the analysis of agricultural economy” (CREA). She is also involved as Management Committee member and Chair of WG4 in the COST Action CHARME ”Harmonising standardisation strategies to increase efficiency and competitiveness of European life-science research”. | ||
| The discovery of the pivotal roles of non-coding RNAs (ncRNAs) on gene expression and genome maintenance represents one of the most significant revolution of the last decades in life science research. Some ncRNA classes, such as ribosomal RNAs and transfer RNAs, have been known for a very long time, others, such as micro RNAs (miRNAs), Piwi interacting RNAs (piRNAs), and long non-coding RNAs (lnRNAs) were discovered in more recent years. However, the discovery of the great diversity and magnitude of this family of regulators is a recent achievement. Indeed, it is only in the last decade that, thanks to the advent of next generation sequencing technologies, large-scale sequencing studies have allowed scientists to systematically analyse ncRNAs for their real size and functional activities. These studies have identified a surprisingly large number of new and diverse ncRNA genes that are emerging to be central to many aspects of plant and animal gene regulation. Domenica D’Elia will expose some of the most relevant and recent results in this fascinating research domain, highlighting her research and recent achievements. | ||
| 15:20 - 15:40 | Tea & Coffee Break | |
| 15:40 - 16:20 |
Ancient duplication history of vertebrate (Human) genome [Abstract] Amir Ali Abbasi, National Center for Bioinformatics, Pakistan [Personal Profile] |
|
| Dr. Amir Ali Abbasi obtained his doctorate in human biology in 2008 from Philipps University, Marburg, Germany. He is long fascinated in several areas of research landscapes within the scope of Evolutionary and Functional Genomics, Population and Medical Genetics, Evolutionary Developmental Biology and Genome Bioinformatics. He is very keen to integrated computational methodological work which is tight in with experimental research and external collaborations, where his group studies the variability of molecular traits in different systems, including Zebrafish and transgenic mice models. Another enthralling area of his lab lies in deciphering evolution of vertebrate gene families and genomes in order to pin down the major morphological transitions that vertebrates accomplished during their deep history (>450 mya). With the recent availability of vast amount of population-wide genomic data, his current interests lie in the field of evolutionary medicine with the application of modern evolutionary theory to understanding health and disease. Currently his group is trying to answer how evolution has shaped the human physiology under different environmental pressures and life styles over-time that may leave us susceptible to certain diseases. Such evolutionary approaches have important implications in advancing the knowledge of medical science regarding systematic molecular diagnostics and treatment of certain diseases like, antibiotic resistance, cancer, autoimmune disease and human anatomy. | ||
| Among the bilaterians, vertebrates have unique anatomical features and possess the greatest number of cell/tissue types. It was speculated that the invention of genes with new functions underlies increasing developmental, morphological, and metabolic complexity during vertebrate early history. Particularly, in the early years of genomic research prior to the availability of large-scale animal genome sequence data, based on rather inaccurate indicators such as genome size and gene number, it was suggested that whole-genome duplications (WGDs) generated a large amount of raw material to prompt evolution of novelty and complexity in a short time during early vertebrate history. This notion popularly theorized “2R hypothesis” (two rounds of WGDs) has been widely debated. This talk would compare the evidence in favor of and against the 2R hypothesis. | ||
| 16:20 - 17:00 |
Convergent thinking: theoretical frameworks of genomic data Jun Yu, BIG, CAS, China [Personal Profile] |
|
| Dr. YU obtained his B.S. degree in biochemistry from Jilin University in 1983 and Ph.D. degree in biomedical sciences from New York University Medical School in 1990. He had served as a Research Assistant Professor at NYU since 1990 until he joined University of Washington Genome Center in 1993. Dr. YU is one of the founders of Beijing Institute of Genomics and has been the deputy director until 2012. Dr. YU participated in and presided many important projects including The International Human Genome Project (Chinese part), Super Hybrid Rice Genome Project, The Silkworm Genome Project, The Date Palm Genome Project etc. | ||
| 17:00 - 18:00 | Free time and transfer | |
| 18:00 - 20:00 | Welcome reception | |
| December 7: Forum Discussion on Precision Medicine & Big data, Conference room at the 2nd floor, BIG, CAS | ||
| 09:00 - 09:30 | Warmup with Coffee & Tea | |
| 09:30 - 09:40 | Introduction (Zhang Zhang) | |
| 09:40 - 10:00 | Overview of China Precision Medicine Projects (Jun Yu) | |
| 10:00 - 12:00 |
Brainstorming session 1: Precision Medicine and Health Data (break at 10:40)
|
|
| 12:00 - 14:00 | Lunch | |
| 14:00 - 14:20 | Overview of BIG Data Center (Zhang Zhang) | |
| 14:20 - 16:50 |
Brainstorming session 2: BIG Data Center Database Resources (break at 15:20)
|
|
| 16:50 - 17:00 | Final remarks | |
| 17:00 - 18:00 | Free time and Transfer | |
| 18:00 - 20:00 | Dinner | |
| December 8: Forum Discussion on BIG Data Center, Conference Room at YaAo (Best Western) Hotel | ||
| 09:00 - 10:20 | Suggestions on Future Directions | |
| 10:20 - 10:40 | Tea & Coffee Break | |
| 10:40 - 12:00 | International Connections and Collaborations | |
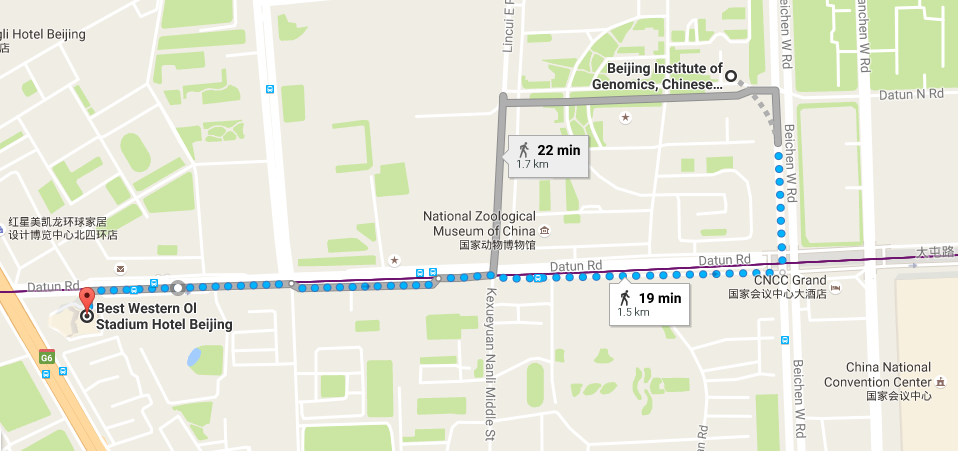
Location
-
Beijing Institute of Genomics, Chinese Academy of Sciences
No.1 Beichen West Road, Chaoyang District
Beijing 100101, China -

Hotel
-
Best Western Ol Stadium Hotel (北京亚奥国际酒店)
No.1 Datun Road Jia, Chaoyang District
Beijing 100101, China -