Documentation
1. Database Overview
Single-cell multimodal sequencing technology represents a significant advancement over traditional single-modality approaches such as scRNA-seq and scATAC-seq. Unlike these methods that focus on profiling either the transcriptome or the epigenome, multimodal sequencing enables the simultaneous analysis of multiple types of molecular information from the same cell, including but not limited to gene expression, chromatin accessibility, and DNA methylation status. This comprehensive view allows for an unprecedented understanding of cellular heterogeneity and regulatory mechanisms at the single-cell level.
The uniqueness of single-cell multimodal data lies in its ability to reveal complex relationships between different layers of biological regulation within individual cells, providing insights into how various molecular features interact to determine cell states and functions. However, this richness also introduces analytical challenges. Integrating and interpreting multimodal data require sophisticated computational tools capable of handling high-dimensional datasets and uncovering meaningful patterns across diverse data types. Furthermore, ensuring the accuracy and reproducibility of such analyses remains a critical issue, demanding rigorous quality control measures and validation strategies.
Given the unique characteristics and analytical challenges of single-cell multimodal data, scMultiModalMap has established a comprehensive analysis pipeline, with a particular emphasis on high-quality data preprocessing and deep learning-based integration methods. The platform systematically collects, processes, integrates, and visualizes a total of 174 single-cell multimodal datasets. These datasets span multiple modalities, such as gene expression, surface protein abundance, and chromatin accessibility, enabling a more holistic understanding of cellular states and functions.
Our goal is to provide researchers with a powerful and user-friendly platform for studying cellular heterogeneity at an unprecedented level of detail. By leveraging advanced computational tools and deep learning techniques, scMultiModalMap not only ensures robust data quality control but also facilitates the discovery of cross-modal biological insights that would be difficult to obtain using single-modality approaches alone. This integrated resource serves as a valuable reference for exploring gene regulatory mechanisms, cell development trajectories, and disease-related cellular states in a wide range of biological contexts.
2. Data Processing Pipeline
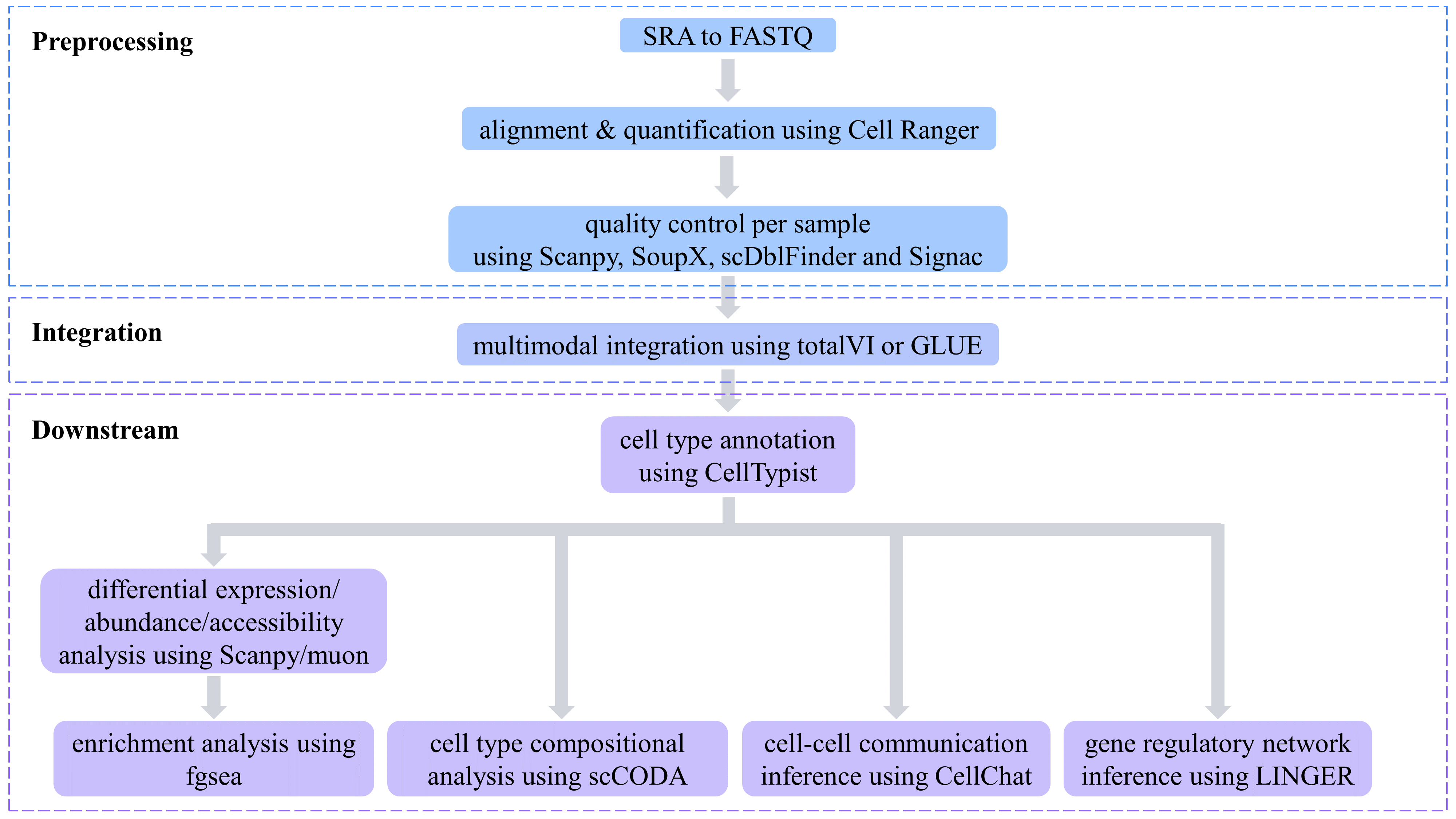
Raw sequencing data (in SRA format) from datasets obtained via GEO and ENCODE were first converted to FASTQ format using the fasterq-dump command from SRA Toolkit 3.0.0. Subsequently, Cell Ranger 8.0.0 and Cell Ranger ARC 2.0.2 were employed to align sequencing reads and quantify gene expression levels, surface protein levels, and chromatin accessibility peaks. The resulting data were then compiled into HDF5 files containing feature count matrices and other essential metadata. For datasets sourced directly from 10x Genomics, pre-processed HDF5 files along with corresponding metadata were downloaded and used for downstream analyses.
During the quality control process, each sample and each modality within the datasets was processed independently to minimize variations arising from sample heterogeneity, sequencing biases, and differences across modalities.
For the gene expression modality in both CITE-seq and 10x Multiome datasets, Scanpy 1.10.2 was used to select high-quality cells based on the following criteria: (1) the log-transformed total gene counts were within the range of the median counts ± 5 median absolute deviations (MAD) across all cells; (2) the log-transformed number of detected genes per cell was within the same range (median ± 5 MAD); and (3) the proportion of mitochondrial gene counts was less than 10% for human samples or 5% for mouse samples. Subsequently, ambient RNA contamination was corrected using SoupX 1.6.2, and doublets were filtered out using scDblFinder 1.16.0.
For the surface protein abundance modality in CITE-seq datasets, Scanpy 1.10.2 was used to select high-quality cells based on the following criteria: (1) the log-transformed total protein counts were within the range of the median ± 5 MAD across all cells; (2) the log-transformed number of detected proteins was within the same range (median ± 5 MAD); and (3) no more than 20% of the total protein counts originated from antibody isotype controls. Additionally, the percentages of counts from B cell surface protein markers (CD19 and IgM) and T cell surface protein marker (CD3) were calculated. If both B cell marker percentages and T cell marker percentages were greater than 10%, and the difference between them was less than 5%, the corresponding cells were identified as doublets and removed.
For the chromatin accessibility modality in 10x Multiome datasets, Signac 1.14.0 was used to select high-quality cells based on the following criteria: (1) the log-transformed total peak counts were within the range of the median ± 5 MAD across all cells; (2) the nucleosome signal scores were less than 4; (3) the transcription start site (TSS) enrichment scores were greater than 1; (4) the percentage of fragments in peaks was greater than 40%; and (5) the ratio of reads in genomic blacklist regions was less than 0.01 for human samples or 0.05 for mouse samples.
Different modalities within the same dataset were aligned based on cell barcodes using muon 0.1.7. To integrate samples and modalities, deep learning models were applied: TotalVI 1.2.1 for CITE-seq datasets and GLUE 0.3.2 for 10x Multiome datasets. Subsequently, Scanpy 1.10.2 was used to cluster cells into groups based on the latent representations generated by these models.
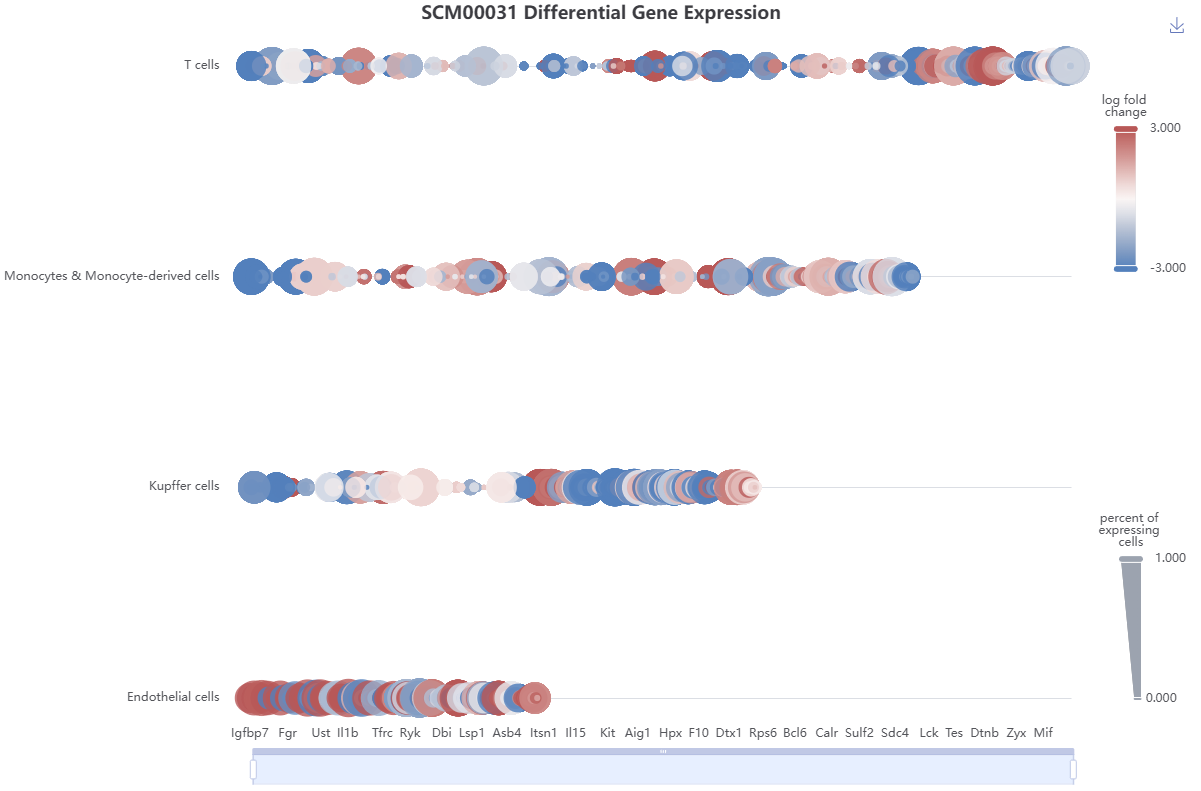
Cell type annotations were performed using CellTypist 1.6.3, employing both the official CellTypist models and customized models to improve annotation accuracy. Differential gene expression analysis and differential protein abundance analysis were carried out using Scanpy 1.10.2 to identify cell type-specific gene markers and surface protein markers. Similarly, differential chromatin accessibility analysis was performed using muon 0.1.7. The Wilcoxon rank-sum test was selected as the statistical method for all differential analyses. Results were filtered within each cell type based on the following criteria: (1) the adjusted p-values were less than 0.05; (2) the absolute log fold change were greater than or equaled 0.1; and (3) the percentage of expressing cells within this cell type were greater than or equaled with 0.1. Subsequently, the results were sorted by ascending adjusted p-values and descending absolute z-scores. Finally, the top 200 positively and 200 negatively differential genes were retained. Likewise, the top 2000 positively and 2000 negatively differential peaks were retained.
Following differential gene expression analysis, gene set enrichment analysis was performed using fgsea 1.28.0. Three gene set collections were selected from the Molecular Signatures Database (MSigDB): C5 Gene Ontology (GO) Biological Process (BP) gene sets, C5 GO Molecular Function (MF) gene sets, and C2 Canonical Pathways (CP) Reactome gene sets. Cell type compositional analysis was conducted using scCODA (implemented in pertpy 0.9.4), to identify cell type-specific changes in abundance influenced by experimental or biological conditions. Cell-cell communication was inferred using CellChat 2.1.2, based on the expression levels of ligand-receptor pairs. Finally, gene regulatory networks were inferred using LINGER 1.101, which integrates information from both the gene expression and chromatin accessibility modalities. This multimodal approach enables improved performance compared to conventional methods that rely solely on gene expression data.
3. Database Usage
3.1. Homepage Quick Search
The homepage features a quick search function that enables users to effortlessly explore our curated datasets by filtering across six key categories: multimodality type, sequencing protocol, species, CITE-seq antibody panel, sample description, and sample source. This intuitive search interface allows researchers to rapidly locate and access the most relevant datasets to support their studies.

3.2. Datasets
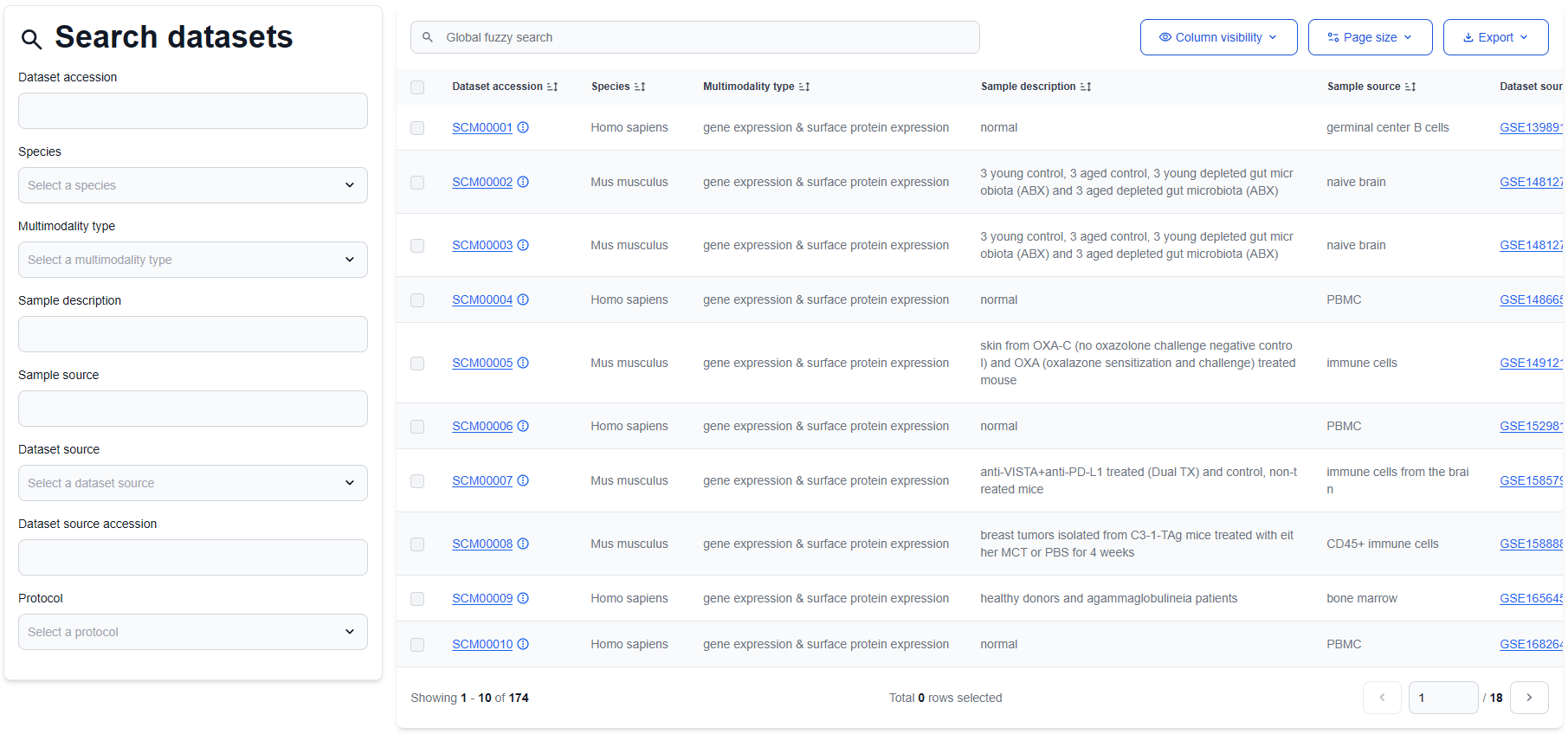
On the Datasets page, users can browse the collection of curated datasets and perform targeted searches to locate those most relevant to their research. Additionally, an integrated global fuzzy search feature enables users to quickly filter and identify datasets using broad or partial search terms, enhancing the efficiency of data discovery.
3.3. Dataset Detail
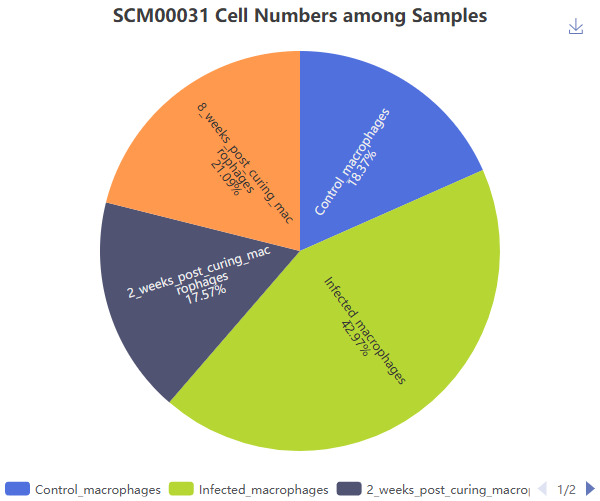
Each dataset is accompanied by comprehensive metadata and analytical summaries, including dataset meta information, sample quality control metrics, cell count distributions across samples and biological conditions, cell type composition, UMAP visualizations (both integrated and single-modal), differential gene expression, differential protein abundance, differential chromatin accessibility, and results from enrichment analysis.







3.4. UMAP Visualization
The UMAP visualization consists of two interactive panels. The first UMAP panel enables users to explore cell populations across different samples, conditions, clusters, and cell types within the selected dataset. The second UMAP panel allows for interactive examination of modality-specific features, such as gene, protein, or chromatin accessibility peaks at single-cell resolution. By combining both panels, users can investigate the expression or presence of specific features across distinct cell types.
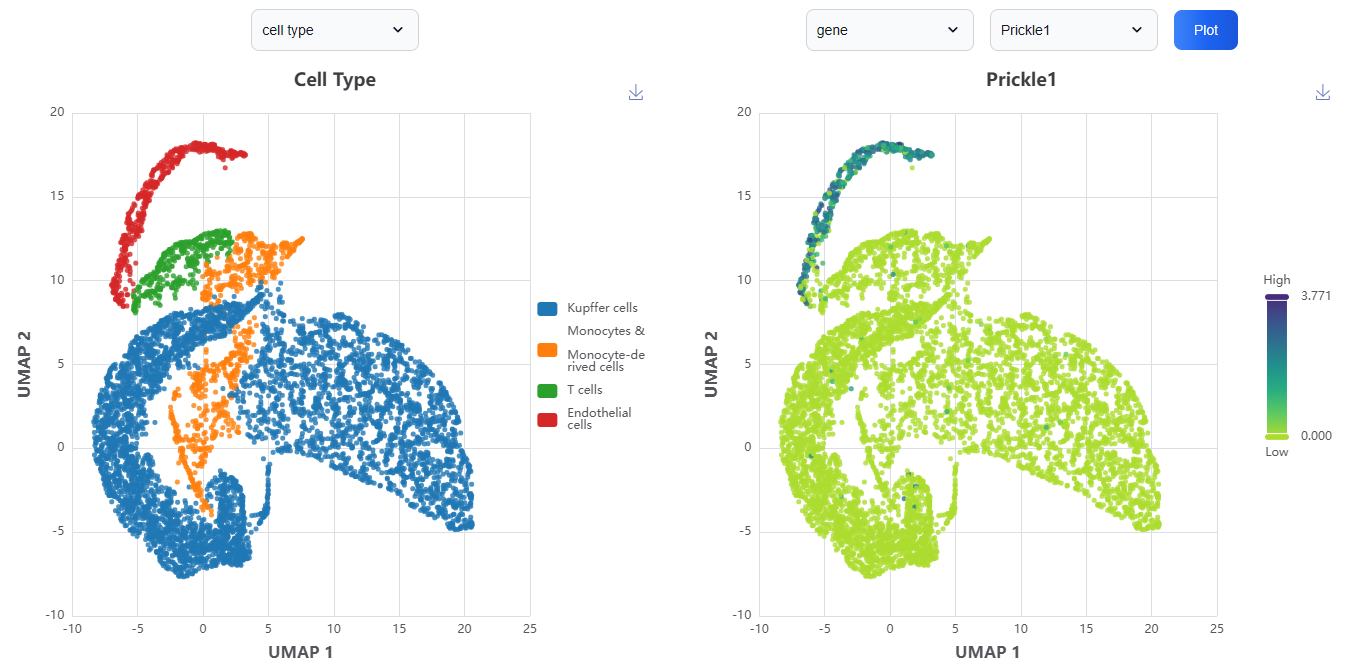
3.5. Feature Level Comparison
The feature level comparison enables users to directly explore and compare the expression or enrichment levels of modality-specific features, such as genes, proteins, or chromatin accessibility peaks across different cell types. This interactive visualization facilitates the identification of cell type-specific markers, functional differences, and regulatory patterns within datasets.

3.6. Cross-Modal Feature Correlation Analysis
Use the cross-modal feature correlation analysis to explore pairwise relationships between features, such as genes, proteins, or chromatin accessibility peaks across different modalities. This analytical tool helps uncover statistically significant correlations, potentially revealing functional associations, co-regulation mechanisms, or candidate biomarkers for further experimental validation.
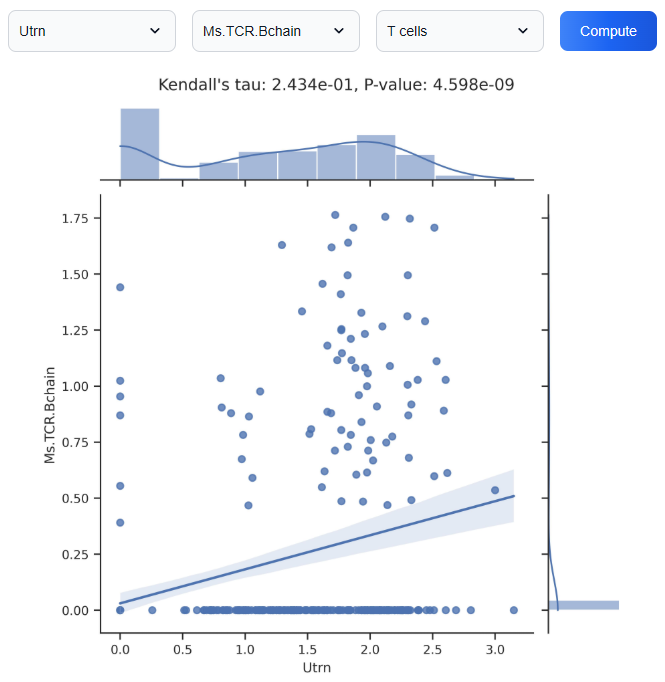
3.7. Cross-Modal Feature Regression Analysis
Use the cross-modal feature regression analysis to model how the expression of a target feature, such as a surface protein or marker gene is influenced by other features, including genes or chromatin accessibility peaks, across individual cells. This analytical tool enables the identification of potential regulatory features, supports the inference of directional interactions, and facilitates hypothesis generation regarding gene regulatory mechanisms within complex cellular systems.
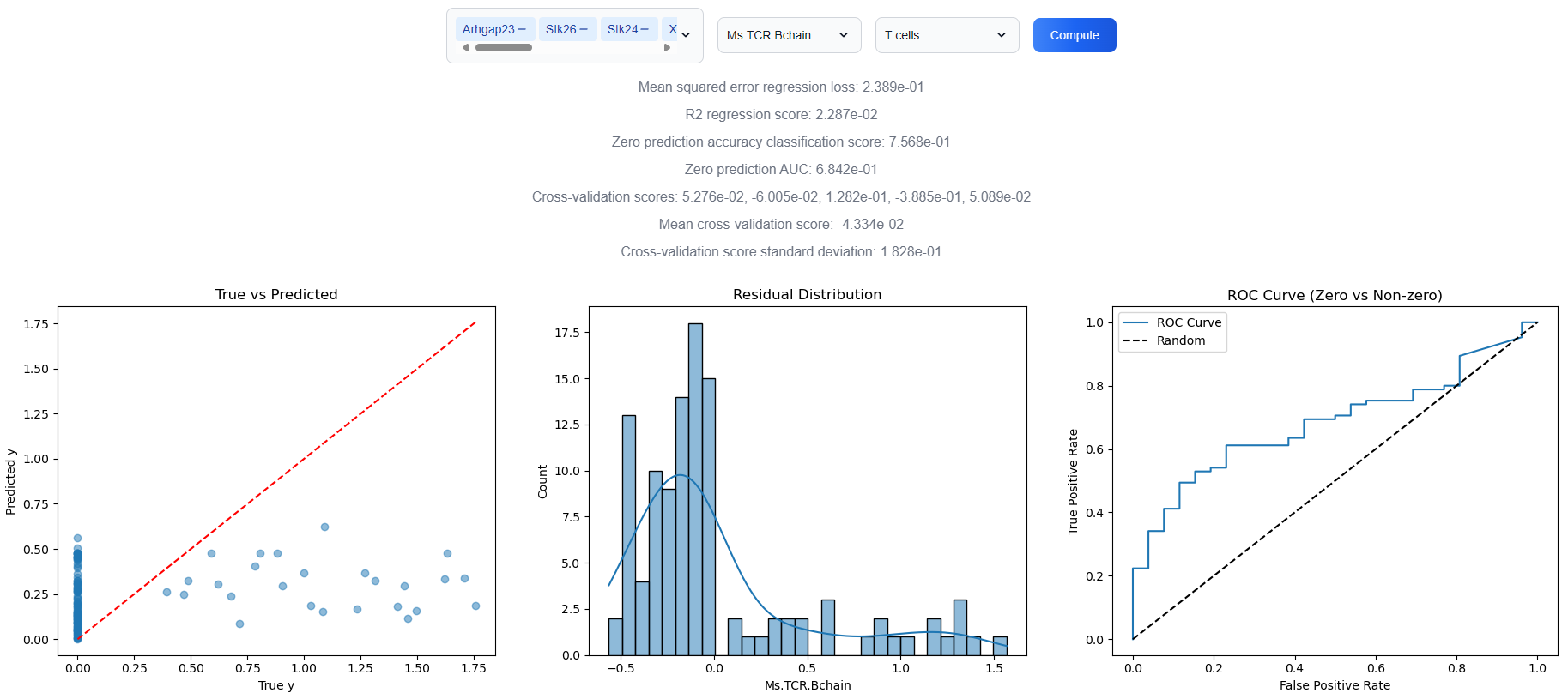
3.8. Cross-Modal Gene Regulatory Network Inference
Use the cross-modal gene regulatory network inference to predict and analyze transcription factor-regulatory element (TF-RE) binding scores, cis-regulatory scores and trans-regulatory scores at both the population level and within individual cell types. This approach integrates multimodal data to provide insights into the regulatory mechanisms governing gene expression in complex biological systems.

3.9. Cell Type Compositional Analysis
Use the cell type compositional analysis to identify cell types that exhibit statistically significant changes in relative abundance under different experimental or biological conditions. This analysis helps uncover condition-associated shifts in cellular composition and supports the interpretation of functional or pathological implications.

3.10. Cell-cell Communication Inference
Use the cell-cell communication inference to identify and analyze biologically significant cell-cell communication networks and signaling pathways. This analysis leverages expression data of ligands, receptors, and their interactions to infer potential communication mechanisms between different cell types. By uncovering these networks, researchers can gain insights into complex intercellular interactions and their roles in physiological and pathological processes.
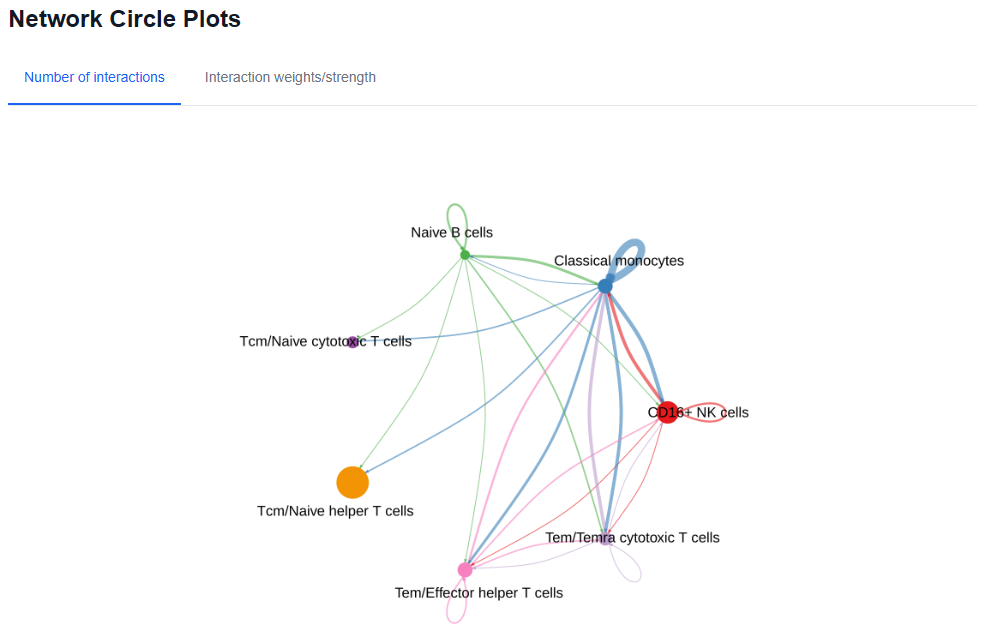
3.11. Downloads
On the Downloads page, users can access and download a variety of resources, including single-cell multimodal data in h5mu format, CellChat RDS files, and scMultiModalMap customized CellTypist models. These resources support downstream analyses, data reuse, and integration into custom pipelines.