Age Predictor is an online tool to estimate methylation age of the sample by three machine learning methods (SVM, Random Forest and Elastic Net).

Home
User Manual
Category
RNA Expression
Bulk RNA-seq Data AnalysisCancer Alternative Splicing AnalysisCCLHunterCIRI-deepCIRI3Cross-disease analysisDisease predictionEditing site annotationEditing site identificationEFDetectorFunGenGene-disease network constructionHELIXNCSelPredRNA-seq AnalysisSingle-cell RNA-seq Data AnalysisSPIRALTIVar diffTIVar predictVisualization of scRNA-seq Data Analysis Results
Virus
COVID-19 genome variation annotationCOVID-19 haplotype analysisDenovo AssemblyEvolutionary treeFastq-to-VariantsGenealogy and Evolutionary AnalysisGenome AnnotationGenome TracingMcANMonkeypox virus genome variation annotationMonkeypox virus genome variation identificationPangolin COVID-19 Lineage AssignerSeqQCVENASVISTA
Home
Epigenome analysis

BS-RNA
An efficient mapping and annotation tool for RNA bisulfite sequencing data
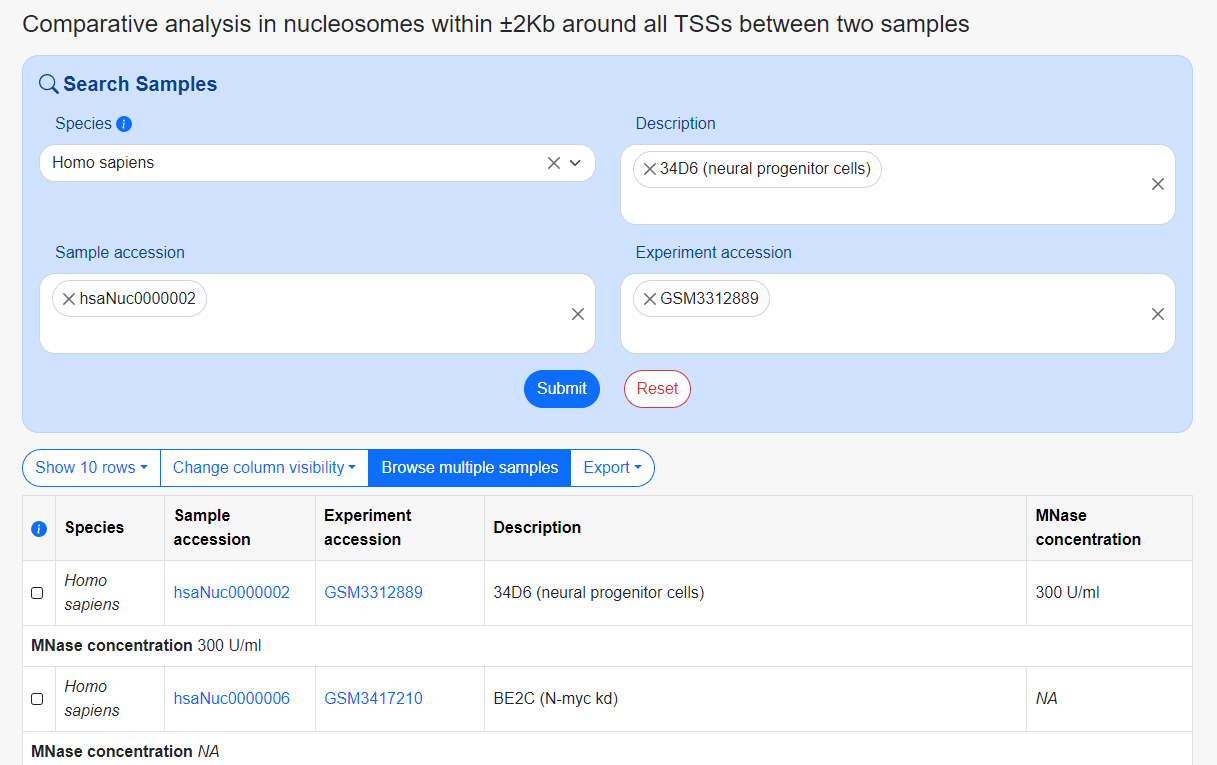
Comparative analysis in nucleosomes
Compare the displacement and occupancy differences of 2Kb upstream and downstream of the transcription start site between two samples

DMR Annotation
Differentially methylated regions (DMRs) enrichment and pathway analysis

DMR Toolkit
DMR toolkit is a pipline package for differentally methylated regions (DMRs) identification, annotation and enrichment for multiple species

Enrichment & Annotation
Visualization of chromosome position enrichment, functional enrichment, expression correlation, transcription factor enrichment, and other analyses related to methylation site sets

Enrichment analysis in nucleosome occupancy
Enrichment analysis of nucleosome occupancy in designated regions of the genome for multiple samples

EWAS Network Visualization
Visualization of epigenetic knowledge graph

GMQN
Standardization method for methylated chip data based on quantile of Gaussian mixture model

IDMP
Identification of differentially methylated promoters between two samples
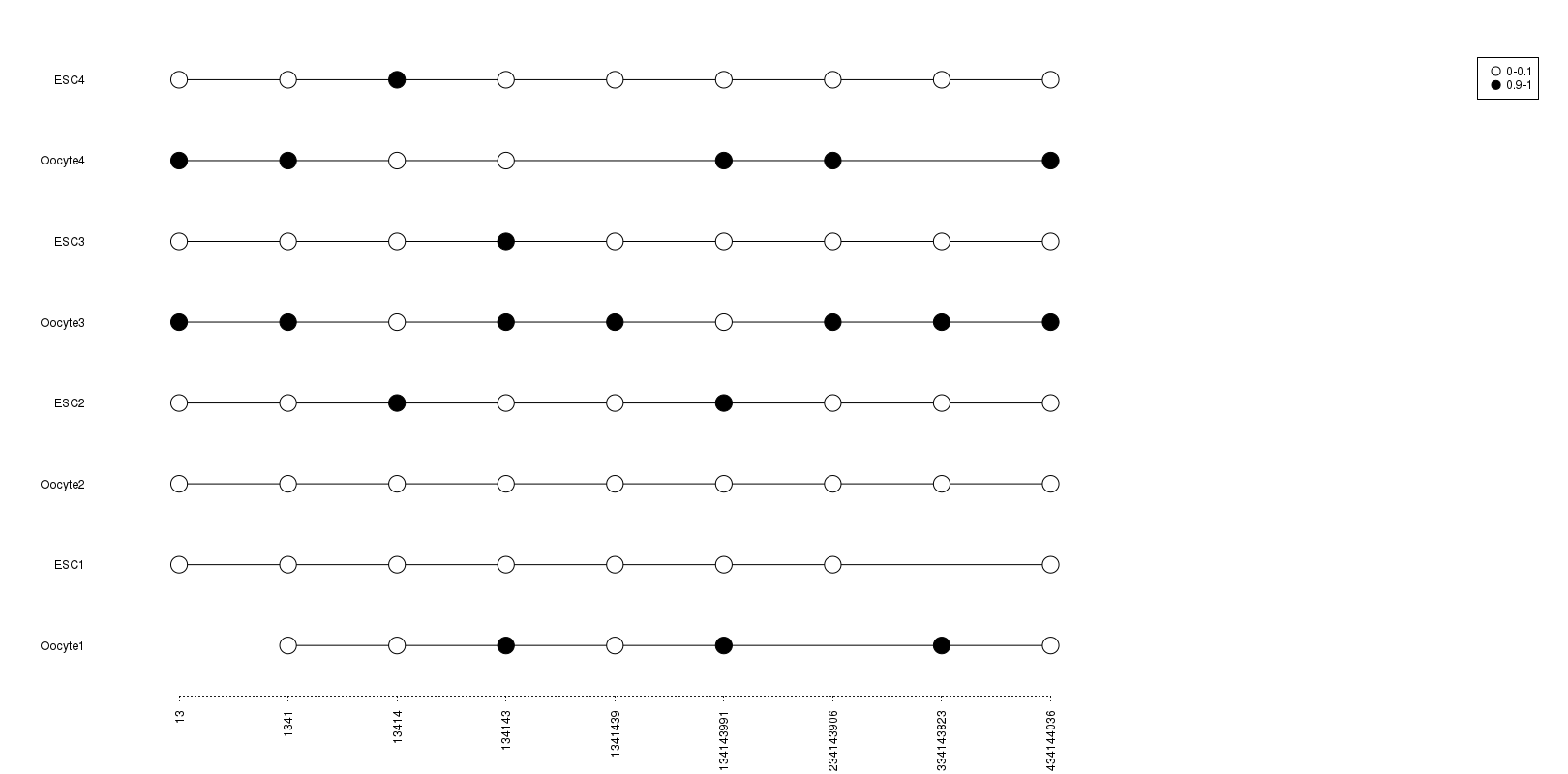
Lollipop Plotter
A web tool for Lollipop-style plot for single cell methylation patterns

MRAS
MRAS is designed to identify the critical splicing factors whose altered expression drives extensive splicing abnormalities.
